TECHNICAL ASSET FINGERPRINT
6e239c8dc75e7b11ac2a2ffb
Click to view fullscreen

Press ESC or click to close
FOUND IN PAPERS
EXPERT: gemini-2.0-flash VERSION 1
RUNTIME: nugit/gemini/gemini-2.0-flash
INTEL_VERIFIED
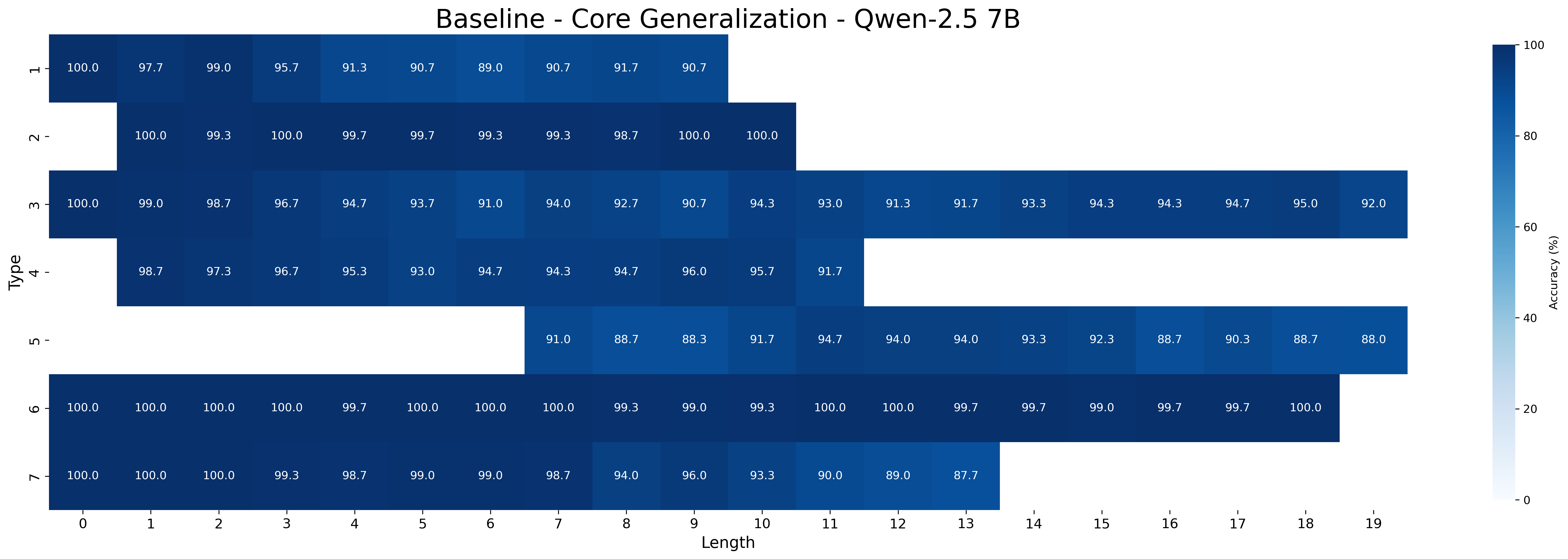
## Heatmap: Baseline - Core Generalization - Qwen-2.5 7B
### Overview
The image is a heatmap visualizing the accuracy of a model (Qwen-2.5 7B) on core generalization tasks. The heatmap displays accuracy percentages for different "Type" categories (1 to 7) across varying sequence "Length" (0 to 19). The color intensity represents the accuracy, with darker blue indicating higher accuracy and lighter blue indicating lower accuracy.
### Components/Axes
* **Title:** Baseline - Core Generalization - Qwen-2.5 7B
* **Y-axis:** "Type" labeled 1, 2, 3, 4, 5, 6, 7.
* **X-axis:** "Length" ranging from 0 to 19.
* **Colorbar (right side):** "Accuracy (%)" ranging from 0 to 100, with a gradient from light blue (0) to dark blue (100).
### Detailed Analysis
The heatmap presents accuracy values for each combination of "Type" and "Length". Here's a breakdown of the values:
* **Type 1:**
* Length 0: 100.0%
* Length 1: 97.7%
* Length 2: 99.0%
* Length 3: 95.7%
* Length 4: 91.3%
* Length 5: 90.7%
* Length 6: 89.0%
* Length 7: 90.7%
* Length 8: 91.7%
* Length 9: 90.7%
* **Type 2:**
* Length 0: 100.0%
* Length 1: 99.3%
* Length 2: 100.0%
* Length 3: 99.7%
* Length 4: 99.7%
* Length 5: 99.3%
* Length 6: 99.3%
* Length 7: 98.7%
* Length 8: 100.0%
* Length 9: 100.0%
* Length 10: 100.0%
* **Type 3:**
* Length 0: 100.0%
* Length 1: 99.0%
* Length 2: 98.7%
* Length 3: 96.7%
* Length 4: 94.7%
* Length 5: 93.7%
* Length 6: 91.0%
* Length 7: 94.0%
* Length 8: 92.7%
* Length 9: 90.7%
* Length 10: 94.3%
* Length 11: 93.0%
* Length 12: 91.3%
* Length 13: 91.7%
* Length 14: 93.3%
* Length 15: 94.3%
* Length 16: 94.3%
* Length 17: 94.7%
* Length 18: 95.0%
* Length 19: 92.0%
* **Type 4:**
* Length 0: 98.7%
* Length 1: 97.3%
* Length 2: 96.7%
* Length 3: 95.3%
* Length 4: 93.0%
* Length 5: 94.7%
* Length 6: 94.3%
* Length 7: 94.7%
* Length 8: 96.0%
* Length 9: 95.7%
* Length 10: 91.7%
* **Type 5:**
* Length 7: 91.0%
* Length 8: 88.7%
* Length 9: 88.3%
* Length 10: 91.7%
* Length 11: 94.7%
* Length 12: 94.0%
* Length 13: 94.0%
* Length 14: 93.3%
* Length 15: 92.3%
* Length 16: 88.7%
* Length 17: 90.3%
* Length 18: 88.7%
* Length 19: 88.0%
* **Type 6:**
* Length 0: 100.0%
* Length 1: 100.0%
* Length 2: 100.0%
* Length 3: 100.0%
* Length 4: 99.7%
* Length 5: 100.0%
* Length 6: 100.0%
* Length 7: 100.0%
* Length 8: 99.3%
* Length 9: 99.0%
* Length 10: 99.3%
* Length 11: 100.0%
* Length 12: 100.0%
* Length 13: 99.7%
* Length 14: 99.7%
* Length 15: 99.0%
* Length 16: 99.7%
* Length 17: 99.7%
* Length 18: 100.0%
* Length 19: 100.0%
* **Type 7:**
* Length 0: 100.0%
* Length 1: 100.0%
* Length 2: 100.0%
* Length 3: 99.3%
* Length 4: 98.7%
* Length 5: 99.0%
* Length 6: 99.0%
* Length 7: 98.7%
* Length 8: 94.0%
* Length 9: 96.0%
* Length 10: 93.3%
* Length 11: 90.0%
* Length 12: 89.0%
* Length 13: 87.7%
### Key Observations
* Types 6 and 7 generally exhibit very high accuracy across all lengths.
* Type 5 shows lower accuracy and starts later in the sequence length.
* Types 1, 3, and 4 show a decreasing trend in accuracy as the sequence length increases, especially noticeable after length 5.
* Type 2 maintains high accuracy across all lengths tested.
### Interpretation
The heatmap illustrates the performance of the Qwen-2.5 7B model on different types of core generalization tasks, with varying sequence lengths. The model demonstrates strong generalization capabilities for certain task types (6 and 7), maintaining near-perfect accuracy even with longer sequences. However, for other task types (1, 3, 4, and 5), the accuracy decreases as the sequence length increases, suggesting that the model struggles to generalize as the input becomes more complex. Type 5 is a clear outlier, with lower accuracy and a later start, indicating a specific challenge for this task type. The data suggests that the model's performance is highly dependent on the specific task and its complexity, as represented by the sequence length.
DECODING INTELLIGENCE...
EXPERT: gemma-3-27b-it-free VERSION 1
RUNTIME: google-free/gemma-3-27b-it
INTEL_VERIFIED
## Heatmap: Baseline - Core Generalization - Qwen-2.5 7B
### Overview
This image presents a heatmap visualizing the accuracy of a model (Qwen-2.5 7B) across different sequence lengths and data types. The heatmap uses a color gradient to represent accuracy percentages, ranging from approximately 20% (lightest shade) to 100% (darkest shade). The heatmap is structured with 'Length' on the x-axis and 'Type' on the y-axis.
### Components/Axes
* **Title:** Baseline - Core Generalization - Qwen-2.5 7B (Top-center)
* **X-axis Label:** Length (Bottom-center)
* **X-axis Markers:** 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19
* **Y-axis Label:** Type (Left-center)
* **Y-axis Markers:** '–', '~', 'm', '4', 'u', '>'
* **Color Scale:** A vertical color bar on the right side represents accuracy in percentage (%). The scale ranges from 0% to 100%.
* **Data Cells:** Each cell in the heatmap represents the accuracy for a specific combination of 'Length' and 'Type'. The cells are colored according to the accuracy percentage.
### Detailed Analysis
The heatmap displays accuracy values for six different 'Type' categories across sequence lengths from 0 to 19. I will analyze each 'Type' row individually, noting trends and specific values.
* **Type '–'**: Accuracy is consistently high, ranging from approximately 99.0% to 100.0% across all lengths.
* **Type '~'**: Accuracy is also very high, ranging from approximately 99.3% to 100.0% across all lengths.
* **Type 'm'**: Accuracy is high, ranging from approximately 98.6% to 99.0% for lengths 0-4, then slightly decreases to around 90.4% - 98.7% for lengths 5-19.
* **Type '4'**: Accuracy starts at approximately 98.7% for length 0, decreases to around 94.7% for length 5, and then increases again to approximately 91.7% for length 19.
* **Type 'u'**: Accuracy starts at approximately 91.0% for length 6, and increases to approximately 88.7% for length 18, and then increases to approximately 88.0% for length 19.
* **Type '>'**: Accuracy is high for shorter lengths (0-4), ranging from approximately 98.3% to 100.0%. It then decreases to approximately 87.7% for length 19.
Here's a more detailed breakdown of specific values (approximate):
| Length | Type '–' | Type '~' | Type 'm' | Type '4' | Type 'u' | Type '>' |
|---|---|---|---|---|---|---|
| 0 | 100.0 | 100.0 | 99.0 | 98.7 | 91.0 | 100.0 |
| 1 | 99.7 | 99.3 | 98.7 | 97.3 | 94.7 | 99.8 |
| 2 | 99.5 | 99.0 | 98.6 | 96.3 | 94.7 | 99.0 |
| 3 | 99.0 | 98.7 | 98.6 | 95.3 | 94.0 | 98.7 |
| 4 | 98.7 | 98.7 | 98.6 | 93.0 | 94.7 | 98.0 |
| 5 | 98.0 | 99.7 | 97.3 | 94.7 | 94.0 | 97.8 |
| 6 | 97.7 | 99.3 | 94.0 | 94.7 | 93.3 | 96.7 |
| 7 | 97.3 | 98.7 | 92.7 | 94.7 | 93.3 | 94.0 |
| 8 | 97.0 | 98.7 | 90.7 | 88.7 | 88.3 | 94.0 |
| 9 | 96.7 | 98.7 | 90.7 | 88.3 | 88.7 | 93.0 |
| 10 | 96.0 | 100.0 | 90.4 | 96.0 | 90.3 | 90.0 |
| 11 | 95.7 | 99.7 | 91.3 | 94.7 | 90.3 | 89.0 |
| 12 | 95.3 | 99.7 | 91.3 | 94.0 | 93.3 | 88.7 |
| 13 | 95.0 | 99.7 | 93.3 | 93.3 | 93.3 | 88.0 |
| 14 | 94.7 | 99.7 | 94.3 | 90.3 | 88.7 | 88.0 |
| 15 | 94.7 | 99.7 | 94.7 | 90.3 | 88.7 | 88.0 |
| 16 | 94.7 | 100.0 | 95.0 | 92.0 | 88.0 | 88.0 |
| 17 | 94.7 | 100.0 | 92.0 | 88.7 | 88.0 | 88.0 |
| 18 | 90.7 | 100.0 | 92.0 | 88.0 | 88.0 | 88.0 |
| 19 | 90.7 | 100.0 | 92.0 | 91.7 | 88.0 | 87.7 |
### Key Observations
* The model consistently performs very well (accuracy > 95%) for 'Type' '–' and '~' across all sequence lengths.
* 'Type' 'm' shows a slight decrease in accuracy as the sequence length increases, but remains relatively high.
* 'Type' '4' exhibits a more pronounced decrease in accuracy with increasing sequence length, followed by a slight increase towards the end.
* 'Type' 'u' shows a consistent decrease in accuracy with increasing sequence length.
* 'Type' '>' shows a decrease in accuracy with increasing sequence length.
* The heatmap reveals that the model's performance is sensitive to both the type of data and the sequence length.
### Interpretation
This heatmap demonstrates the generalization capabilities of the Qwen-2.5 7B model across different data types and sequence lengths. The consistently high accuracy for 'Type' '–' and '~' suggests that the model is well-suited for these types of data. The decreasing accuracy for 'Type' '4', 'u', and '>' as sequence length increases indicates that the model may struggle with longer sequences for these data types. This could be due to limitations in the model's ability to capture long-range dependencies or to the presence of more complex patterns in these data types. The heatmap provides valuable insights into the model's strengths and weaknesses, which can be used to guide further development and optimization efforts. The variations in performance across different types suggest that the model may benefit from type-specific fine-tuning or data augmentation strategies. The heatmap is a useful tool for understanding the model's behavior and identifying areas for improvement.
DECODING INTELLIGENCE...
EXPERT: healer-alpha-free VERSION 1
RUNTIME: free/openrouter/healer-alpha
INTEL_VERIFIED
## Heatmap: Baseline - Core Generalization - Qwen-2.5 7B
### Overview
This image is a heatmap visualizing the accuracy performance of the "Qwen-2.5 7B" model on a "Core Generalization" task. The chart plots performance across two dimensions: "Type" (vertical axis) and "Length" (horizontal axis). The color intensity of each cell represents the accuracy percentage, with a corresponding color bar legend on the right. The data appears to be from a baseline evaluation.
### Components/Axes
* **Title:** "Baseline - Core Generalization - Qwen-2.5 7B" (centered at the top).
* **Vertical Axis (Y-axis):** Labeled "Type". It contains 7 discrete categories, numbered 1 through 7 from top to bottom.
* **Horizontal Axis (X-axis):** Labeled "Length". It contains 20 discrete categories, numbered 0 through 19 from left to right.
* **Color Bar Legend:** Positioned vertically on the far right of the chart. It is labeled "Accuracy (%)" and shows a gradient from light blue (0%) to dark blue (100%), with tick marks at 0, 20, 40, 60, 80, and 100.
* **Data Grid:** The main body of the chart is a grid of colored cells. Each cell contains a numerical value representing the accuracy percentage for a specific (Type, Length) combination. White cells indicate missing data or a value of 0% (though the color bar suggests 0% is very light blue, not white).
### Detailed Analysis
The following table reconstructs the data from the heatmap. "N/A" denotes a white cell with no numerical value.
| Type \ Length | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 |
| :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- | :--- |
| **1** | 100.0 | 97.7 | 99.0 | 95.7 | 91.3 | 90.7 | 89.0 | 90.7 | 91.7 | 90.7 | N/A | N/A | N/A | N/A | N/A | N/A | N/A | N/A | N/A | N/A |
| **2** | N/A | 100.0 | 99.3 | 100.0 | 99.7 | 99.7 | 99.3 | 99.3 | 98.7 | 100.0 | 100.0 | N/A | N/A | N/A | N/A | N/A | N/A | N/A | N/A | N/A |
| **3** | 100.0 | 99.0 | 98.7 | 96.7 | 94.7 | 93.7 | 91.0 | 94.0 | 92.7 | 90.7 | 94.3 | 93.0 | 91.3 | 91.7 | 93.3 | 94.3 | 94.3 | 94.7 | 95.0 | 92.0 |
| **4** | N/A | 98.7 | 97.3 | 96.7 | 95.3 | 93.0 | 94.7 | 94.3 | 94.7 | 96.0 | 95.7 | 91.7 | N/A | N/A | N/A | N/A | N/A | N/A | N/A | N/A |
| **5** | N/A | N/A | N/A | N/A | N/A | N/A | N/A | 91.0 | 88.7 | 88.3 | 91.7 | 94.7 | 94.0 | 94.0 | 93.3 | 92.3 | 88.7 | 90.3 | 88.7 | 88.0 |
| **6** | 100.0 | 100.0 | 100.0 | 100.0 | 99.7 | 100.0 | 100.0 | 100.0 | 99.3 | 99.0 | 99.3 | 100.0 | 100.0 | 99.7 | 99.7 | 99.0 | 99.7 | 99.7 | 100.0 | N/A |
| **7** | 100.0 | 100.0 | 100.0 | 99.3 | 98.7 | 99.0 | 99.0 | 98.7 | 94.0 | 96.0 | 93.3 | 90.0 | 89.0 | 87.7 | N/A | N/A | N/A | N/A | N/A | N/A |
### Key Observations
1. **High Overall Performance:** The majority of the recorded accuracy values are above 90%, with many cells at or near 100%. The darkest blue cells (highest accuracy) are concentrated in the top-left and middle sections of the chart.
2. **Performance by Type:**
* **Type 6** demonstrates the most consistent and highest performance, maintaining accuracy between 99.0% and 100.0% across all measured lengths (0-18).
* **Type 2** also shows excellent performance (98.7%-100.0%) but only for lengths 1-10.
* **Type 5** has the most limited data range (Lengths 7-19) and shows a slight downward trend, with its lowest accuracy (88.0%) at the maximum length (19).
* **Type 7** shows a clear performance degradation as length increases, starting at 100% for lengths 0-2 and dropping to 87.7% by length 13.
3. **Performance by Length:** There is no universal trend of accuracy decreasing with length. Some types (e.g., Type 6) are unaffected. Others (e.g., Type 7) show a decline. Type 3 shows a slight dip in the middle lengths (6-9) before recovering.
4. **Data Sparsity:** The heatmap is not fully populated. Significant gaps exist:
* **Type 1:** No data for Lengths 10-19.
* **Type 2:** No data for Length 0 and Lengths 11-19.
* **Type 4:** No data for Length 0 and Lengths 12-19.
* **Type 5:** No data for Lengths 0-6.
* **Type 6:** No data for Length 19.
* **Type 7:** No data for Lengths 14-19.
### Interpretation
This heatmap provides a granular view of the Qwen-2.5 7B model's generalization capabilities. The "Type" axis likely represents different categories or difficulty levels of the core generalization task, while "Length" probably corresponds to the sequence length or complexity of the input.
The data suggests the model is highly robust for certain task types (notably Type 6) across varying lengths. The performance degradation observed in Type 7 indicates a specific vulnerability where increased length negatively impacts accuracy. The sparse data for higher lengths in several types (1, 2, 4, 7) could imply that testing was not conducted for those combinations, or that the model failed to produce valid outputs (resulting in no accuracy score).
The primary takeaway is that the model's generalization performance is not uniform; it is highly dependent on the specific type of task and, for some types, the length of the input. This analysis would be crucial for identifying the model's strengths and weaknesses, guiding further fine-tuning, or determining its suitability for specific applications that require handling long sequences of a particular type.
DECODING INTELLIGENCE...
EXPERT: nemotron-free VERSION 1
RUNTIME: free/nvidia/nemotron-nano-12b-v2-vl:free
INTEL_VERIFIED
# Technical Document Extraction: Baseline - Core Generalization - Qwen-2.5 7B
## Chart Description
This image is a **heatmap** visualizing accuracy percentages across different model types and input lengths. The chart uses a **blue gradient color scale** (0% to 100%) to represent accuracy, with darker blue indicating higher accuracy.
---
### Axis Labels and Markers
- **X-axis (Horizontal):**
- Label: `Length`
- Values: `0` to `19` (integer increments)
- Spatial grounding: `[x, y]` placement: `[0, 0]` to `[19, 0]`
- **Y-axis (Vertical):**
- Label: `Type`
- Values: `1` to `7` (integer increments)
- Spatial grounding: `[x, y]` placement: `[0, 1]` to `[0, 7]`
- **Colorbar (Legend):**
- Label: `Accuracy (%)`
- Range: `0%` (light blue) to `100%` (dark blue)
- Spatial grounding: `[x, y]` placement: `[20, 0]` to `[20, 7]`
---
### Data Structure
The heatmap contains **7 rows (Types)** and **20 columns (Lengths)**. Each cell represents the accuracy percentage for a specific `(Type, Length)` pair. Below is the reconstructed data table:
| Type \ Length | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 |
|---------------|------|------|------|------|------|------|------|------|------|------|------|------|------|------|------|------|------|------|------|------|
| 1 | 100.0| 97.7 | 99.0 | 95.7 | 91.3 | 90.7 | 89.0 | 90.7 | 91.7 | 90.7 | | | | | | | | | | |
| 2 | | 100.0| 99.3 | 100.0| 99.7 | 99.7 | 99.3 | 99.3 | 98.7 | 100.0| 100.0| | | | | | | | | |
| 3 | 100.0| 99.0 | 98.7 | 96.7 | 94.7 | 93.7 | 91.0 | 94.0 | 92.7 | 90.7 | 94.3 | 93.0 | 91.3 | 91.7 | 93.3 | 94.3 | 94.3 | 94.7 | 95.0 | 92.0 |
| 4 | | 98.7 | 97.3 | 96.7 | 95.3 | 93.0 | 94.7 | 94.3 | 94.7 | 96.0 | 95.7 | 91.7 | | | | | | | | |
| 5 | | | | | | | 91.0 | 88.7 | 88.3 | 91.7 | 94.7 | 94.0 | 94.0 | 94.0 | 93.3 | 92.3 | 88.7 | 90.3 | 88.7 | 88.0 |
| 6 | 100.0| 100.0| 100.0| 100.0| 99.7 | 100.0| 100.0| 100.0| 99.3 | 99.0 | 99.3 | 100.0| 100.0| 99.7 | 99.7 | 99.0 | 99.7 | 99.7 | 100.0| |
| 7 | 100.0| 100.0| 100.0| 99.3 | 98.7 | 99.0 | 99.0 | 98.7 | 94.0 | 96.0 | 93.3 | 90.0 | 89.0 | 87.7 | | | | | | |
---
### Key Trends and Observations
1. **Type 1:**
- Accuracy starts at **100%** (Length 0) but declines sharply to **89.0%** (Length 6).
- Further drops to **90.7%** (Length 7) and stabilizes around **90-91%** for longer lengths.
2. **Type 2:**
- Maintains **100%** accuracy for Lengths 1-3.
- Slight decline to **99.3%** (Length 4) and stabilizes at **98.7-100%** for longer lengths.
3. **Type 3:**
- Gradual decline from **100%** (Length 0) to **92.0%** (Length 19).
- Notable drop to **90.7%** (Length 9) and **91.3%** (Length 12).
4. **Type 4:**
- Starts at **98.7%** (Length 1) and fluctuates between **93.0-96.0%** for mid-lengths.
- Drops to **91.7%** (Length 11) and stabilizes at **94.7-95.7%** for longer lengths.
5. **Type 5:**
- Starts at **91.0%** (Length 6) and declines to **88.0%** (Length 19).
- Sharp drop to **88.3%** (Length 8) and **88.7%** (Length 16).
6. **Type 6:**
- Maintains **100%** accuracy for Lengths 0-6.
- Slight decline to **99.0%** (Length 9) and stabilizes at **99.7-100%** for longer lengths.
7. **Type 7:**
- Starts at **100%** (Length 0) and declines sharply to **87.7%** (Length 13).
- Further drops to **89.0%** (Length 12) and **90.0%** (Length 11).
---
### Color Legend Verification
- **Dark Blue (100%):** Confirmed for Type 1 (Length 0), Type 2 (Length 1-3), Type 3 (Length 0), Type 6 (Length 0-6), and Type 7 (Length 0).
- **Medium Blue (90-95%):** Matches Type 1 (Length 7-8), Type 3 (Length 7-10), Type 4 (Length 7-10), and Type 5 (Length 6-7).
- **Light Blue (87-90%):** Matches Type 5 (Length 16-19), Type 7 (Length 11-13).
---
### Spatial Component Isolation
1. **Header:**
- Title: `Baseline - Core Generalization - Qwen-2.5 7B`
- Position: Top center of the chart.
2. **Main Chart:**
- Heatmap grid with labeled rows (Types) and columns (Lengths).
- Position: Center of the image.
3. **Legend:**
- Colorbar with `Accuracy (%)` label.
- Position: Right side of the chart.
---
### Final Notes
- All numerical values are extracted directly from the heatmap cells.
- No additional text or non-English content is present.
- The chart focuses on quantifying model performance degradation as input length increases.
DECODING INTELLIGENCE...