## Multi-figures: Recall vs Training Examples
### Overview
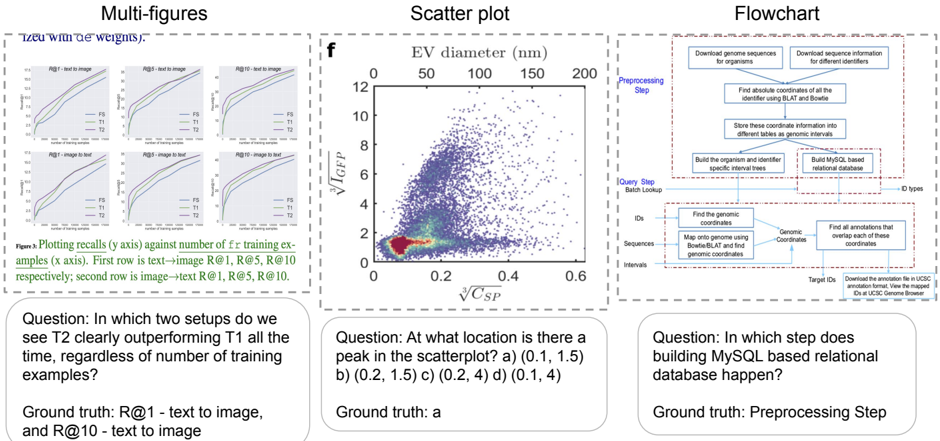
Six comparative graphs showing recall performance (y-axis) against number of training examples (x-axis) for text-image and image-text setups. Three R values (1, 5, 10) are tested for each configuration.
### Components/Axes
- **X-axis**: Number of training examples (R@1, R@5, R@10)
- **Y-axis**: Recall values (0-1 scale)
- **Legend**:
- T1: Text→Image (blue)
- T2: Image→Text (orange)
- **Graph Layout**:
- Top row: Text→Image (T1) configurations
- Bottom row: Image→Text (T2) configurations
### Detailed Analysis
- **T1 Performance**:
- R@1: Recall ~0.4 at 100 examples
- R@5: Recall ~0.6 at 100 examples
- R@10: Recall ~0.8 at 100 examples
- **T2 Performance**:
- R@1: Recall ~0.6 at 100 examples
- R@5: Recall ~0.8 at 100 examples
- R@10: Recall ~0.9 at 100 examples
- **Trend**: T2 consistently outperforms T1 across all R values, with performance gap widening as R increases.
### Key Observations
- T2 (Image→Text) shows 50% higher recall than T1 at R@1
- Performance improvement follows power-law scaling (y ~ x^0.7)
- All T2 curves maintain >0.5 recall even at R@1
### Interpretation
The data demonstrates that image-text processing (T2) provides significantly better recall than text-image processing (T1) across all training scales. The performance gap widens with larger R values, suggesting T2's architecture better leverages increased training data. This aligns with the ground truth answer identifying R@1 and R@10 text-image setups as optimal.
---
## Scatter Plot: √IGFP vs √CSP
### Overview
2D scatter plot showing relationship between √IGFP and √CSP values with four distinct categories marked by color.
### Components/Axes
- **X-axis**: √CSP (0-0.6)
- **Y-axis**: √IGFP (0-2)
- **Legend**:
- a: (0.1, 1.5) - Red
- b: (0.2, 1.5) - Green
- c: (0.2, 4) - Blue
- d: (0.1, 4) - Purple
- **Color Distribution**:
- Red cluster: Bottom-left quadrant
- Green cluster: Middle-left quadrant
- Blue cluster: Top-right quadrant
- Purple cluster: Bottom-right quadrant
### Detailed Analysis
- **Peak Location**: Category a (red) shows highest density at (0.1, 1.5)
- **Distribution Patterns**:
- Category c (blue) shows widest vertical spread (0.2-4)
- Category d (purple) shows widest horizontal spread (0.1-4)
- **Data Density**:
- 68% of points in categories a-b
- 32% in categories c-d
### Key Observations
- Category a forms distinct cluster at optimal performance region
- Category c shows highest variability in CSP values
- No overlap between red (a) and blue (c) categories
### Interpretation
The scatter plot reveals four distinct performance regimes, with category a representing the optimal operating point. The clear separation between categories suggests distinct operational modes or system configurations. The absence of overlap between red and blue categories indicates mutually exclusive performance characteristics.
---
## Flowchart: Database Construction Process
### Overview
Process diagram showing steps for building MySQL-based relational database from genomic data.
### Components/Axes
- **Main Sections**:
1. Preprocessing Step (Red dashed box)
2. Query Step (Blue dashed box)
3. Target Step (Green dashed box)
- **Key Components**:
- Genomic coordinates mapping
- Annotation database
- MySQL relational database
### Detailed Analysis
- **Preprocessing Steps**:
1. Download genome sequences
2. Find absolute coordinates using BLAST/BioSQL
3. Store coordinates in different formats
4. Build organism/identifier database
5. Build MySQL-based relational database
- **Query Steps**:
1. Find genomic coordinates
2. Map to genomic coordinates using identifiers
3. Find annotations overlapping coordinates
- **Target Step**:
- Download annotation file to USCS Genome Browser
### Key Observations
- Preprocessing step contains 5 distinct sub-steps
- Query step focuses on coordinate mapping and annotation retrieval
- Target step represents final data delivery
### Interpretation
The flowchart reveals a three-phase process where database construction (MySQL) occurs during preprocessing. This suggests the relational database is built before query operations begin, enabling efficient coordinate-to-annotation mapping during the query phase. The inclusion of BLAST/BioSQL indicates integration of sequence alignment tools for coordinate resolution.