## Complexity of Networks (reprise)
## Russell K. Standish
Mathematics and Statistics, University of New South Wales hpcoder@hpcoders.com.au
November 3, 2018
## Abstract
Network or graph structures are ubiquitous in the study of complex systems. Often, we are interested in complexity trends of these system as it evolves under some dynamic. An example might be looking at the complexity of a food web as species enter an ecosystem via migration or speciation, and leave via extinction.
In a previous paper, a complexity measure of networks was proposed based on the complexity is information content paradigm. To apply this paradigm to any object, one must fix two things: a representation language, in which strings of symbols from some alphabet describe, or stand for the objects being considered; and a means of determining when two such descriptions refer to the same object. With these two things set, the information content of an object can be computed in principle from the number of equivalent descriptions describing a particular object.
The previously proposed representation language had the deficiency that the fully connected and empty networks were the most complex for a given number of nodes. A variation of this measure, called zcomplexity, applied a compression algorithm to the resulting bitstring representation, to solve this problem. Unfortunately, zcomplexity proved too computationally expensive to be practical.
In this paper, I propose a new representation language that encodes the number of links along with the number of nodes and a representation of the linklist. This, like zcomplexity, exhibits minimal complexity for fully connected and empty networks, but is as tractable as the original measure.
This measure is extended to directed and weighted links, and several real world networks have their network complexities compared with randomly generated model networks with matched node and link counts, and matched link weight distributions. Compared with the random networks, the real world networks have significantly higher complexity, as do artificially generated food webs created via an evolutionary process, in several well known ALife models.
Keywords: complexity; information; networks; foodwebs
## 1 Introduction
This work situates itself firmly within the complexity is information content paradigm, a topic that dates back to the 1960s with the work of Kolmogorov, Chaitin and Solomonoff. Prokopenko et al. [28] provide a recent review of the area, and argue for the importance of information theory to the notion of complexity measure. In [32], I argue that information content provides an overarching complexity measure that connects the many and various complexity measures proposed (see [14] for a review) up to that time. In some ways, information-based complexity measures are priveleged, motivating the present work to develop a satisfactory information-based complexity measure of networks.
The idea is fairly simple. In most cases, there is an obvious prefix-free representation language within which descriptions of the objects of interest can be encoded. There is also a classifier of descriptions that can determine if two descriptions correspond to the same object. This classifier is commonly called the observer , denoted O ( x ).
To compute the complexity of some object x , count the number of equivalent descriptions ω ( /lscript,x ) of length /lscript that map to the object x under the agreed classifier. Then the complexity of x is given in the limit as /lscript →∞ :
<!-- formula-not-decoded -->
where N is the size of the alphabet used for the representation language.
Because the representation language is prefix-free, every description y in that language has a unique prefix of length s ( y ). The classifier does not care what symbols appear after this unique prefix. Hence ω ( /lscript,O ( y )) ≥ N /lscript -s ( y ) . As /lscript increases, ω must increase as fast, if not faster than N /lscript , and do so monotonically. Therefore C ( O ( y )) decreases monotonically with /lscript , but is bounded below by 0. So equation (1) converges.
The relationship of this algorithmic complexity measure to more familiar measures such as Kolmogorov (KCS) complexity, is given by the coding theorem[22, Thm 4.3.3]. In this case, the descriptions are halting programs of some given universal Turing machine U , which is also the classifier. Equation (1) then corresponds to the logarithm of the universal a priori probability , which is a kind of formalised Occam's razor that gives higher weight to simpler (in the KCS sense) computable theories for generating priors in Bayesian reasoning. The difference between this version of C and KCS complexity is bounded by a constant independent of the complexity of x , so these measures become equivalent in the limit as message size goes to infinity.
Many measures of network properties have been proposed, starting with node count and connectivity (no. of links), and passing in no particular order through cyclomatic number (no. of independent loops), spanning height (or width), no. of spanning trees, distribution of links per node and so on. Graphs tend to be classified using these measures small world graphs tend to have small spanning height relative to the number of nodes and scale free networks exhibit a power law distribution of node link count.
Some of these measures are related to graph complexity, for example node count and connectivity can be argued to be lower and upper bounds of the network complexity respectively. More recent attempts include offdiagonal complexity [9], which was compared with an earlier version of this proposal in [36], and medium articulation [42, 19].
However, none of the proposed measures gives a theoretically satisfactory complexity measure, which in any case is context dependent (ie dependent on the observer O , and the representation language).
In setting the classifier function, we assume that only the graph's topology counts - positions, and labels of nodes and links are not considered important. Links may be directed or undirected. We consider the important extension of weighted links in a subsequent section.
A plausible variant classifier problem might occur when the network describes a dynamical system (such as interaction matrix of the EcoLab model, which will be introduced later). In such a case, one might be interested in classifying the network according to numbers and types of dynamical attractors, which will be strongly influenced by the cyclomatic count of the core network, but only weakly influenced by the periphery. However, this problem lies beyond the scope of this paper.
The issue of representation language, however is far more problematic. In some cases, eg with genetic regulatory networks, there may be a clear representation language, but for many cases there is no uniquely identifiable language. However, the invariance theorem [22, Thm 2.1.1] states that the difference in complexity determined by two different Turing complete representation languages (each of which is determined by a universal Turing machine) is at most a constant, independent of the objects being measured. Thus, in some sense it does not matter what representation one picks - one is free to pick a representation that is convenient, however one must take care with non-Turing complete representations.
In the next section, I will present a concrete graph description language that can be represented as binary strings, and is amenable to analysis. The quantity ω in eq (1) can be simply computed from the size of the automorphism group, for which computationally feasible algorithms exist[23].
The notion of complexity presented in this paper naturally marries with thermodynamic entropy S [21]:
<!-- formula-not-decoded -->
where S max is called potential entropy , ie the largest possible value that entropy can assume under the specified conditions. The interest here is that a dynamical process updating network links can be viewed as a dissipative system, with links being made and broken corresponding to a thermodynamic flux. It would be interesting to see if such processes behave according the maximum entropy production principle[10] or the minimum entropy production principle[27].
In artificial life, the issue of complexity trends in evolution is extremely important[5]. I have explored the complexity of individual Tierran organisms[33,
35], which, if anything, shows a trend to simpler organisms. However, it is entirely plausible that complexity growth takes place in the network of ecological interactions between individuals. For example, in the evolution of the eukaryotic cell, mitochondria are simpler entities than the free-living bacteria they were supposedly descended from. A computationally feasible measure of network complexity is an important prerequisite for further studies of evolutionary complexity trends.
In the results section, several well-known food web datasets are analysed, and compared with randomly-shuffled 'neutral models'. An intriguing 'complexity surplus' is observed, which is also observed in the food webs of several different ALife systems that have undergone evolution.
## 2 Representation Language
One very simple implementation language for undirected graphs is to label the nodes 1 ..n , and the links by the pair ( i, j ) , i < j of nodes that the links connect. The linklist can be represented simply by an L = n ( n -1) / 2 length bitstring, where the 1 2 j ( j -1) + i th position is 1 if link ( i, j ) is present, and 0 otherwise.
The directed case requires doubling the size of the linklist, ie or L = n ( n -1). We also need to prepend the string with the value of N in order to make it prefixfree - the simplest approach is to interpret the number of leading 1s as the number n , which adds a term n +1 to the measured complexity.
This proposal was analysed in [36], and has the unsatisfactory property that the fully connected or empty networks are maximally complex for a given node count. An alternative scheme is to also include the link count as part of the prefix, and to use binary coding for both the node and link counts. The sequence will start with /ceilingleft log 2 n /ceilingright 1's, followed by a zero stop bit, so the prefix will be 2 /ceilingleft log 2 n /ceilingright + /ceilingleft log 2 L /ceilingright +1 bits.
This scheme entails that some of bitstrings are not valid networks, namely ones where the link count does not match the number of 1s in the linklist. We can, however, use rank encoding[25] of the linklist to represent the link pattern. The number of possible linklists corresponding to a given node/link specification is given by
<!-- formula-not-decoded -->
This will have a minimum value of 1 at l = 0 (empty network) and l = L , the fully connected network.
Finally, we need to compute ω of the linklist, which is just the total number of possible renumberings of the nodes ( n !), divided by the size of the graph automorphism group, which can be practically computed by Nauty[23], or a new algorithm I developed called SuperNOVA[38] which exhibits better performance on sparsely linked networks.
A network A that has a link wherever B doesn't, and vice-versa might be called a complement of B . A bitstring for A can be found by inverting the 1s and 0s in the linklist part of the network description. Obviously, ω ( A,L ) = ω ( B,L ),
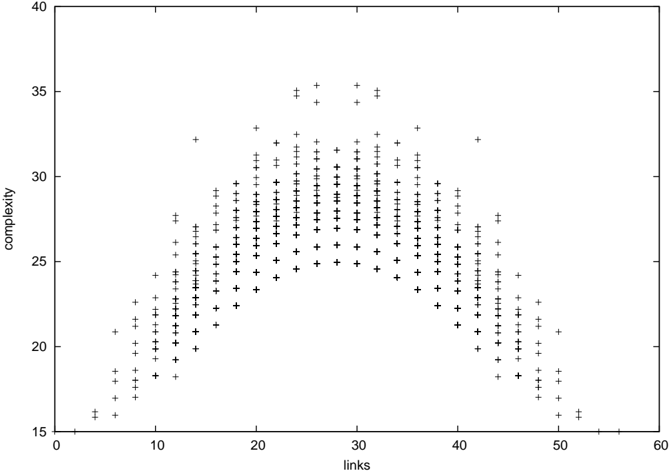
Figure 1: The new complexity measure as a function of link count for all networks with 8 nodes. This shows the strong dependence of complexity on link count, and the symmetry between networks and their complements.
<details>
<summary>Image 1 Details</summary>
### Visual Description
## Scatter Plot: Complexity vs. Links
### Overview
The image is a scatter plot showing the relationship between "complexity" (y-axis) and "links" (x-axis). The data points are represented by '+' symbols. The plot shows an inverted U-shaped distribution, where complexity increases with links up to a certain point, then decreases as links increase further.
### Components/Axes
* **X-axis:** "links", ranging from 0 to 60, with tick marks at intervals of 10 (0, 10, 20, 30, 40, 50, 60).
* **Y-axis:** "complexity", ranging from 15 to 40, with tick marks at intervals of 5 (15, 20, 25, 30, 35, 40).
* **Data Points:** Represented by '+' symbols.
### Detailed Analysis
* **Trend:** The data points form an inverted U-shape.
* From links = 0 to approximately links = 30, the complexity generally increases.
* From links = 30 to links = 60, the complexity generally decreases.
* **Specific Values (Approximate):**
* At links = 5, complexity ranges from approximately 16 to 22.
* At links = 10, complexity ranges from approximately 17 to 24.
* At links = 15, complexity ranges from approximately 20 to 28.
* At links = 20, complexity ranges from approximately 23 to 31.
* At links = 25, complexity ranges from approximately 25 to 35.
* At links = 30, complexity ranges from approximately 25 to 36.
* At links = 35, complexity ranges from approximately 25 to 33.
* At links = 40, complexity ranges from approximately 22 to 30.
* At links = 45, complexity ranges from approximately 19 to 27.
* At links = 50, complexity ranges from approximately 17 to 23.
* At links = 55, complexity ranges from approximately 16 to 20.
### Key Observations
* The highest complexity values are observed around links = 30.
* The spread of complexity values is wider in the middle range of links (approximately 15 to 45) compared to the extreme ends.
* There appears to be a degree of symmetry in the distribution around links = 30.
### Interpretation
The scatter plot suggests that there is a non-linear relationship between the number of links and complexity. Complexity increases as the number of links increases, up to a certain point (around 30 links), after which complexity decreases as the number of links continues to increase. This could indicate that there is an optimal number of links for maximizing complexity, and that beyond this point, adding more links may actually reduce complexity. The spread of data points suggests that there is variability in complexity for any given number of links.
</details>
so the complexity of a network is equal to that of its complement, as can be seen in Figure 1.
A connection can be drawn from this complexity measure, to that proposed in [11]. In their proposal, nodes are labelled, so the network is uniquely specified by the node and link counts, along with a rank-encoded linklist. Also, nodes may link to themselves, so L = n 2 . Consequently, their complexity measure will be
<!-- formula-not-decoded -->
This is, to within a term of order log 2 n corresponding to the lead-in prefix specifying the node count, equal to the unnumbered formula given on page 326 of [11].
## 3 New complexity measure compared with the previous proposals
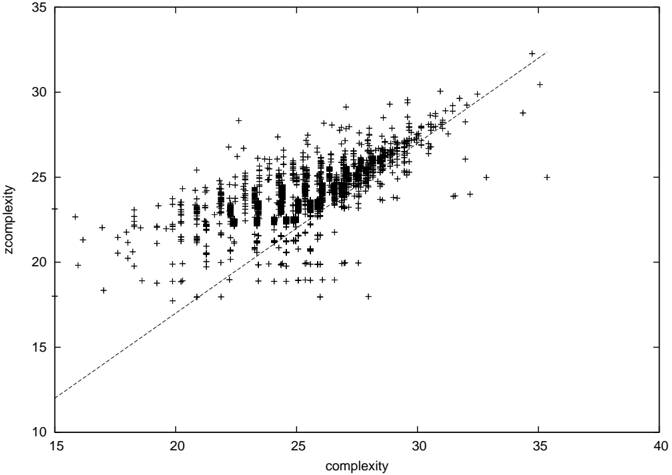
Figures 2 and 3 [36, Fig. 1] shows zcomplexity plotted against the new complexity and the original complexity proposal for all networks of order 8 respectively.
Figure 2: C z plotted against C for all networks of order 8. The diagonal line corresponds to C = C z +3.
<details>
<summary>Image 2 Details</summary>
### Visual Description
## Scatter Plot: Complexity vs. zcomplexity
### Overview
The image is a scatter plot showing the relationship between two variables: "complexity" on the x-axis and "zcomplexity" on the y-axis. The plot contains numerous data points, represented by plus signs, and a dashed line that appears to indicate a trend or a reference line.
### Components/Axes
* **X-axis:** "complexity" with a scale from 15 to 40, marked at intervals of 5 (15, 20, 25, 30, 35, 40).
* **Y-axis:** "zcomplexity" with a scale from 10 to 35, marked at intervals of 5 (10, 15, 20, 25, 30, 35).
* **Data Points:** Represented by '+' symbols.
* **Trend Line:** A dashed line extending from approximately (16, 12) to (36, 32).
### Detailed Analysis
The data points are clustered, showing a positive correlation between "complexity" and "zcomplexity." The density of points is higher in the region where both complexity and zcomplexity are between 20 and 30.
* **Trend Line:** The dashed line starts at approximately x=16, y=12 and extends to x=36, y=32. It suggests a linear relationship where zcomplexity increases with complexity.
* **Data Point Distribution:**
* For complexity values between 20 and 25, zcomplexity values range from approximately 18 to 28.
* For complexity values between 25 and 30, zcomplexity values range from approximately 22 to 30.
* For complexity values between 30 and 35, zcomplexity values range from approximately 25 to 32.
### Key Observations
* There is a positive correlation between "complexity" and "zcomplexity."
* The data points are more densely clustered in the middle range of both axes (20-30).
* There are fewer data points at the extreme ends of the axes.
* The dashed line suggests a linear relationship, but the scatter of the points indicates that the relationship is not perfectly linear.
### Interpretation
The scatter plot suggests that as "complexity" increases, "zcomplexity" also tends to increase. The dashed line provides a visual guide to this trend. However, the spread of the data points around the line indicates that there are other factors influencing "zcomplexity" besides "complexity." The clustering of points in the middle range suggests that the majority of the data falls within a certain range of complexity and zcomplexity values. The plot could be used to identify outliers or to develop a model to predict "zcomplexity" based on "complexity," although the model would need to account for the variability in the data.
</details>
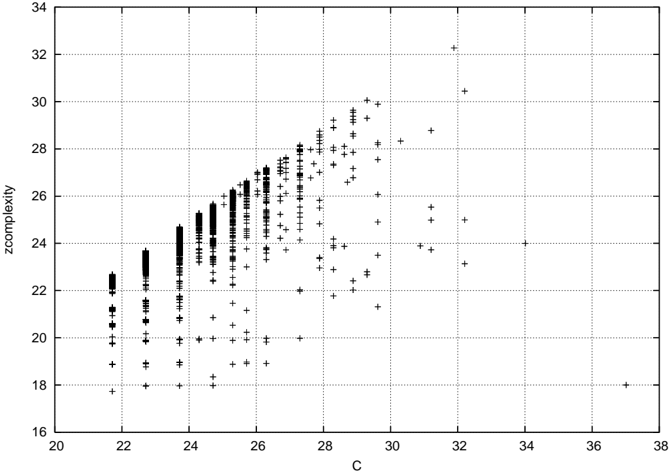
Figure 3: C z plotted against the original C from [36] for all networks of order 8, reproduced from that paper.
<details>
<summary>Image 3 Details</summary>
### Visual Description
## Scatter Plot: Zcomplexity vs. C
### Overview
The image is a scatter plot showing the relationship between two variables, "C" on the x-axis and "zcomplexity" on the y-axis. The data points are represented by plus signs (+). The plot shows a general upward trend, indicating a positive correlation between C and zcomplexity. The data points are more densely clustered at lower values of C and zcomplexity, and become more scattered as both values increase.
### Components/Axes
* **X-axis:** Labeled "C", with a scale from 20 to 38 in increments of 2.
* **Y-axis:** Labeled "zcomplexity", with a scale from 16 to 34 in increments of 2.
* **Data Points:** Represented by "+" symbols.
* **Grid:** The plot has a grid of dotted lines to aid in reading values.
### Detailed Analysis
The data points show a general upward trend.
* **For C = 22:** zcomplexity ranges approximately from 19 to 24.
* **For C = 24:** zcomplexity ranges approximately from 20 to 26.
* **For C = 26:** zcomplexity ranges approximately from 22 to 28.
* **For C = 28:** zcomplexity ranges approximately from 23 to 30.
* **For C = 30:** zcomplexity ranges approximately from 24 to 32.
* **For C = 32:** zcomplexity ranges approximately from 25 to 33.
* **For C = 34:** zcomplexity ranges approximately from 26 to 33.
* **For C = 36:** zcomplexity ranges approximately from 27 to 33.
* **For C = 38:** zcomplexity ranges approximately from 28 to 33.
The density of points decreases as C increases.
### Key Observations
* There is a positive correlation between C and zcomplexity. As C increases, zcomplexity tends to increase as well.
* The spread of zcomplexity values for a given C increases as C increases. This suggests that the relationship between C and zcomplexity becomes less predictable at higher values of C.
* There are no obvious outliers, but the data points are more scattered at higher values of C.
### Interpretation
The scatter plot suggests that there is a relationship between the variables C and zcomplexity. The positive correlation indicates that as C increases, zcomplexity also tends to increase. However, the increasing spread of data points at higher values of C suggests that other factors may also be influencing zcomplexity. The data suggests that C is a contributing factor to zcomplexity, but not the sole determinant. Further investigation would be needed to determine the nature of the relationship and the influence of other factors.
</details>
The new complexity proposal is quite well correlated with zcomplexity, and is in this example about 3 bits higher than zcomplexity. This is just the difference in prefixes between the two schemes (the original proposal used 8 1 bits followed by a stop bit = 9 bits overall to represent the node count, and the new scheme uses 3 1 bits and a stop bit to represent the field width of the node count, 3 bits for the node field and 5 bits for the link count field = 12 bits overall). Data points appearing above the line correspond to networks whose linkfields compress better with the new algorithm than the scheme used with zcomplexity. Conversely, data points below the line are better compressed with the old algorithm.
We can conclude that the new scheme usually compresses the link list field better than the run length encoding scheme employed in [36], and so is a better measure of complexity than zcomplexity, as well as being far more tractable. The slightly more complex prefix of the new scheme grows logarithmically with node count, so will ultimately be more compressed than the prefix of the old scheme, which grows linearly.
## 4 Comparison with medium articulation
In the last few years Wilhelm[42, 19] introduced a new complexity like measure that addresses the intuition that complexity should be minimal for the empty and full networks, and peak for intermediate values (like figure 1). It is obtained by multiplying an information quantity that increases with link count by a different information that falls. The resulting measure is therefore in square bits, so one should perhaps consider the square root as the complexity measure.
Precisely, medium articulation is given by
<!-- formula-not-decoded -->
where w ij is the normalised weight ( ∑ ij w ij = 1) of the link from node i to node j .
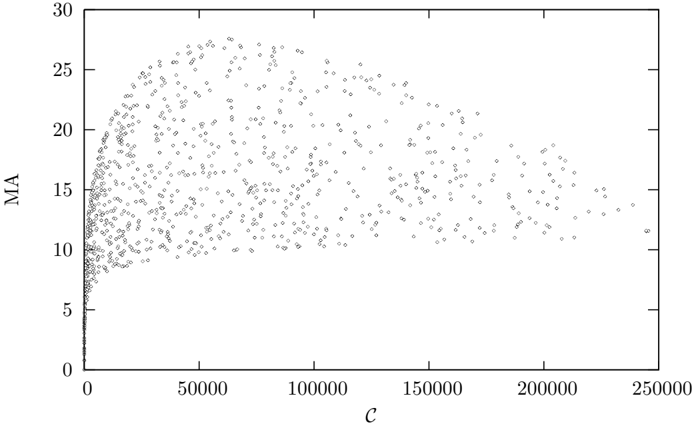
Figure 4 shows medium articulation plotted against C for a sample of 1000 Erd¨ os-R´ enyi networks up to order 500. There is no clear relationship between medium articulation and complexity for the average network. Medium articulation does not appear to discriminate between complex networks. however if we restrict our attention to simple networks (Figures 5 and 6) medium articulation is strongly correlated with complexity, and so could be used as a proxy for complexity for these cases.
## 5 Weighted links
Whilst the information contained in link weights might be significant in some circumstances (for instance the weights of a neural network can only be varied in a limited range without changing the overall qualitative behaviour of the network), of particular theoretical interest is to consider the weights as continuous
Figure 4: Medium Articulation plotted against complexity for 1000 randomly sampled Erd¨ os-R´ enyi graphs up to order 500.
<details>
<summary>Image 4 Details</summary>
### Visual Description
## Scatter Plot: MA vs C
### Overview
The image is a scatter plot showing the relationship between two variables, MA (on the y-axis) and C (on the x-axis). The plot consists of numerous data points scattered across the graph. The data points are concentrated at lower values of C and spread out as C increases.
### Components/Axes
* **X-axis:** Labeled "C". The scale ranges from 0 to 250000, with tick marks at 0, 50000, 100000, 150000, 200000, and 250000.
* **Y-axis:** Labeled "MA". The scale ranges from 0 to 30, with tick marks at 0, 5, 10, 15, 20, 25, and 30.
* **Data Points:** Each data point is represented by a small, unfilled diamond shape.
### Detailed Analysis
* **Trend:** The data points show a rapid increase in MA as C increases from 0 to approximately 20000. Beyond this point, the increase in MA slows down significantly, and the data points become more scattered.
* **Distribution:** At low values of C (0-20000), the data points are densely packed, indicating a strong correlation between C and MA. As C increases, the data points spread out vertically, suggesting a weaker correlation and a wider range of MA values for a given C value.
* **Specific Values:**
* At C = 0, MA ranges from approximately 0 to 10.
* At C = 50000, MA ranges from approximately 8 to 26.
* At C = 100000, MA ranges from approximately 10 to 28.
* At C = 150000, MA ranges from approximately 10 to 26.
* At C = 200000, MA ranges from approximately 10 to 24.
* At C = 250000, MA ranges from approximately 12 to 16.
### Key Observations
* The relationship between C and MA is non-linear.
* There is a saturation effect, where increasing C beyond a certain point does not lead to a significant increase in MA.
* The variability in MA increases with C.
### Interpretation
The scatter plot suggests that the variable C has a strong influence on MA at lower values, but its effect diminishes as C increases. The saturation effect indicates that there may be other factors limiting the value of MA at higher values of C. The increasing variability in MA with C suggests that other variables may be influencing MA, and their effects become more pronounced as C increases. The plot could represent a physical or computational system where C is an input parameter and MA is an output, and the system's response to C changes as C increases.
</details>
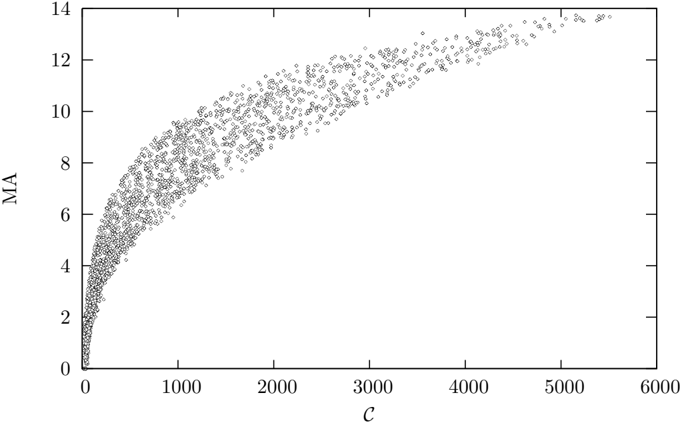
Figure 5: Medium Articulation plotted against complexity for 1000 randomly sampled Erd¨ os-R´ enyi graphs up to order 500 with no more than 2 n links.
<details>
<summary>Image 5 Details</summary>
### Visual Description
## Scatter Plot: MA vs. C
### Overview
The image is a scatter plot showing the relationship between two variables, labeled "MA" on the y-axis and "C" on the x-axis. The plot consists of numerous data points, and the overall trend suggests a positive correlation between C and MA, with the rate of increase in MA decreasing as C increases.
### Components/Axes
* **X-axis:** Labeled "C". The scale ranges from 0 to 6000, with tick marks at intervals of 1000 (0, 1000, 2000, 3000, 4000, 5000, 6000).
* **Y-axis:** Labeled "MA". The scale ranges from 0 to 14, with tick marks at intervals of 2 (0, 2, 4, 6, 8, 10, 12, 14).
* **Data Points:** Each data point is represented by a small, filled circle.
### Detailed Analysis
The data points are densely clustered, especially at lower values of C. As C increases, the density of points decreases, and the spread of MA values for a given C also increases.
* **Trend:** The general trend is that as C increases, MA also increases, but the rate of increase diminishes. The curve appears to be approaching a horizontal asymptote.
* **Specific Values (Approximate):**
* At C = 0, MA ranges from approximately 0 to 4.
* At C = 1000, MA ranges from approximately 3 to 8.
* At C = 2000, MA ranges from approximately 5 to 11.
* At C = 3000, MA ranges from approximately 7 to 12.
* At C = 4000, MA ranges from approximately 8 to 13.
* At C = 5000, MA ranges from approximately 9 to 13.5.
* At C = 6000, MA ranges from approximately 9.5 to 14.
### Key Observations
* The data points are not uniformly distributed. They are more concentrated at lower values of C.
* There is a noticeable spread in MA values for any given value of C, indicating variability in the relationship.
* The rate of increase in MA decreases as C increases, suggesting a diminishing returns effect.
### Interpretation
The scatter plot suggests a positive but non-linear relationship between C and MA. The diminishing rate of increase in MA as C increases indicates that there may be a saturation effect or a limiting factor influencing the relationship. The spread of MA values for a given C suggests that other factors besides C also influence MA. The data suggests that increasing C will lead to an increase in MA, but the magnitude of the increase will be smaller at higher values of C. The relationship could be modeled by a logarithmic or exponential function.
</details>
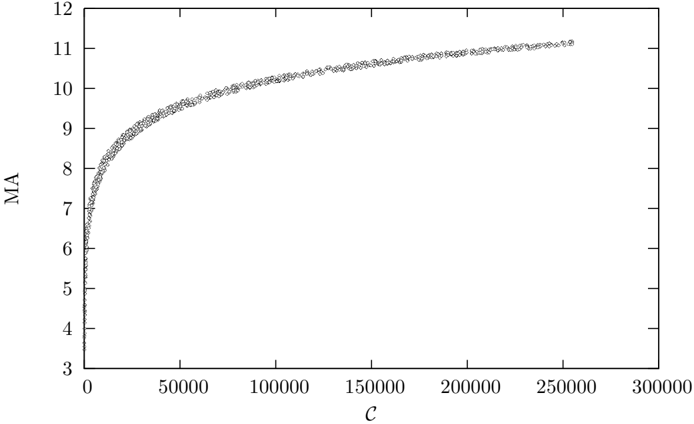
Figure 6: Medium Articulation plotted against complexity for 1000 randomly sampled Erd¨ os-R´ enyi graphs up to order 500 with more than n ( n -5) / 2 links.
<details>
<summary>Image 6 Details</summary>
### Visual Description
## Scatter Plot: MA vs C
### Overview
The image is a scatter plot showing the relationship between two variables, MA (on the y-axis) and C (on the x-axis). The plot shows a non-linear relationship where MA increases rapidly with C at lower values of C, and then the rate of increase slows down as C increases.
### Components/Axes
* **X-axis:** Labeled "C". The scale ranges from 0 to 300000, with tick marks at 0, 50000, 100000, 150000, 200000, 250000, and 300000.
* **Y-axis:** Labeled "MA". The scale ranges from 3 to 12, with tick marks at each integer value.
* **Data Points:** The data points are represented by small circles.
### Detailed Analysis
The data points form a curve that starts at approximately (0, 4) and rises sharply.
* **Initial Rise:** From C = 0 to C = 50000, MA increases rapidly from approximately 4 to 9.
* **Slowing Increase:** From C = 50000 to C = 150000, the rate of increase of MA slows down. MA increases from approximately 9 to 10.
* **Plateau:** From C = 150000 to C = 250000, the curve begins to flatten out, with MA increasing only slightly.
* **Final Value:** At C = 250000, MA is approximately 11.
### Key Observations
* The relationship between MA and C is non-linear.
* The rate of increase of MA decreases as C increases.
* The curve appears to be approaching a horizontal asymptote.
### Interpretation
The scatter plot suggests that there is a diminishing return on MA as C increases. Initially, a small increase in C results in a large increase in MA. However, as C continues to increase, the corresponding increase in MA becomes smaller and smaller. This could indicate that there is a saturation point for MA, beyond which further increases in C have little effect. The data suggests that the relationship between MA and C is not linear, and that there may be an upper limit to the value of MA.
</details>
parameters connecting one network structure with another. For instance if a network X has the same network structure as the unweighted graph A, with b links of weight 1 describing the graph B and the remaining a -b links of weight w , then we would like the network complexity of X to vary smoothly between that of A and B as w varies from 1 to 0. [16] introduced a similar measure.
The most obvious way of defining this continuous complexity measure is to start with normalised weights ∑ i w i = 1. Then arrange the links in weight order, and compute the complexity of networks with just those links of weights less than w . The final complexity value is obtained by integrating:
<!-- formula-not-decoded -->
Obviously, since the integrand is a stepped function, this is computed in practice by a sum of complexities of partial networks.
## 6 Comparing network complexity with the Erd¨ osR´ enyi random model
I applied the weighted network complexity measure (6) to several well-known real network datasets, obtained from Mark Newman's website[41, 20, 26], the Pajek website[7, 39, 40, 17, 2, 1, 3] and Duncan Watt's website[13, 18], with the
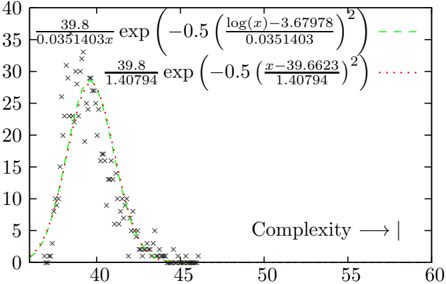
Figure 7: The distribution of complexities of the shuffled Narragansett Bay food web[24]. Both a normal and a log-normal distribution have been fitted to the data - the log-normal is a slightly better fit than the normal distribution. In the bottom right hand corner is marked the complexity value computed for the actual Narragansett food web (58.2 bits).
<details>
<summary>Image 7 Details</summary>
### Visual Description
## Chart: Complexity Distribution
### Overview
The image is a scatter plot overlaid with two curves, representing a distribution of data points related to "Complexity". The x-axis represents complexity, ranging from approximately 35 to 60. The y-axis represents frequency or count, ranging from 0 to 35. The scatter plot shows the distribution of data points, while the curves represent two different mathematical models fitted to the data.
### Components/Axes
* **X-axis:** Labeled "Complexity" with an arrow pointing to the right, indicating increasing complexity. The axis ranges from approximately 35 to 60, with tick marks at intervals of 5.
* **Y-axis:** Ranges from 0 to 35, with tick marks at intervals of 5.
* **Scatter Plot:** A collection of 'x' marks representing individual data points.
* **Curve 1 (Green, Dashed):** Represented by the equation: `(39.8 / 0.0351403) * exp(-0.5 * ((log(x) - 3.67978) / 0.0351403)^2)`
* **Curve 2 (Red, Dotted):** Represented by the equation: `(39.8 / 1.40794) * exp(-0.5 * ((x - 39.6623) / 1.40794)^2)`
### Detailed Analysis
* **Scatter Plot Distribution:** The data points are concentrated between x = 37 and x = 45, with a peak around x = 39. Beyond x = 45, the data points become sparse.
* **Green Dashed Curve:** This curve starts near 0 at x=35, rises sharply to a peak around x=39, and then decreases rapidly, approaching 0 around x=45.
* The equation for the green curve is: `(39.8 / 0.0351403) * exp(-0.5 * ((log(x) - 3.67978) / 0.0351403)^2)`
* **Red Dotted Curve:** This curve also starts near 0 at x=35, rises sharply to a peak around x=39, and then decreases rapidly, approaching 0 around x=45. It closely follows the green curve.
* The equation for the red curve is: `(39.8 / 1.40794) * exp(-0.5 * ((x - 39.6623) / 1.40794)^2)`
### Key Observations
* The data points are clustered around a complexity value of approximately 39.
* Both curves provide a model for the distribution of the data, with peaks around x = 39.
* The green dashed curve and the red dotted curve are very similar in shape and position.
### Interpretation
The chart illustrates the distribution of complexity values, with a clear concentration around a specific value (approximately 39). The two curves represent different attempts to model this distribution mathematically. The similarity between the curves suggests that both models capture the underlying trend in the data. The scatter plot provides the raw data, while the curves offer a smoothed representation of the distribution. The equations provided for the curves indicate that they are Gaussian-like functions, suggesting that the complexity values are normally distributed around a mean. The data suggests that systems or processes being measured tend to have a complexity value close to 39, with fewer instances of very low or very high complexity.
</details>
results shown in Table 1. The number of nodes and links of the networks varied greatly, so the raw complexity values are not particularly meaningful, as the computed value is highly dependent on these two network parameters. What is needed is some neutral model for each network to compare the results to.
At first, one might want to compare the values to an Erd¨ os-R´ enyi random network with the same number of nodes and links. However, in practice, the real network complexities are much less than that of an ER network with the same number of nodes and links. This is because in our scheme, a network with weighted link weights looks somewhat like a simpler network formed by removing some of the weakest links from the original. An obvious neutral network model with weighted network links has the weights drawn from a normal distribution with mean 0. The sign of the weight can be interpreted as the link direction. Because the weights in equation (6) are normalised, the complexity value is independent of the standard deviation of the normal distribution. However, such networks are still much more complex than the real networks, as the link weight distribution doesn't match that found in the real network.
Instead, a simple way of generating a neutral model is to break and reattach the network links to random nodes, without merging links, leaving the original link weights unaltered. In the following experiment, we generate 1000 of these shuffled networks. The distribution of complexities can be fitted to a lognormal distribution, which gives a better likelihood than a normal distribution for all networks studied here[8], although the difference between a log-normal and a normal fit becomes less pronounced for more complex networks. Figure 7 shows the distribution of complexity values computed by shuffling the Narragansett food web, and the best fit normal and lognormal distributions.
In what follows we compute the average 〈 ln C ER 〉 and standard deviation
σ ER of the logarithm of the neutral model complexities. We can then compare the network complexity with the ensemble of neutral models to obtain a p -value, the probability that the observed complexity is that of a random network. The p -values are so small that is better to represent the value as a number of standard deviations ('sigmas') that the logarithm of the measured complexity value is from the mean logarithm of the shuffled networks. A value of 6 sigmas corresponds to a p -value less than 1 . 3 × 10 -4 , although this must be taken with a certain grain of salt, as the distribution of shuffled complexity values has a fatter tail than the fitted log-normal distribution. In none of these samples did the shuffled network complexity exceed the original network's complexity, meaning the p -value is less than 10 -3 . The difference C exp( 〈 ln C ER 〉 ) is the amount of information contained in the specific arrangement of links.
A code implementing this algorithm is implemented as a C++ library, and is available from version 4.D36 onwards as part of the Eco Lab system, an open source modelling framework hosted at http://ecolab.sourceforge.net.
## 7 ALife models
## 7.1 Tierra
Tierra [29] is a well known artificial life system in which self reproducing computer programs written in an assembly-like language are allowed to evolve. The programs, or digital organisms can interact with each other via template matching operations, modelled loosely on the way proteins interact in real biological systems. A number of distinct strategies evolve, including parasitism, where organisms make use of another organism's code and hyper-parasitism where an organism sets traps for parasites in order to steal their CPU resources. At any point in time in a Tierra run, there is an interaction network between the species present, which is the closest thing in the Tierra world to a foodweb.
Tierra is an aging platform, with the last release (v6.02) having been released more than six years ago. For this work, I used an even older release (5.0), for which I have had some experience in working with. Tierra was originally written in C for an environment where ints were 16 bits and long ints 32 bits. This posed a problem for using it on the current generation of 64 bit computers, where the word sizes are doubled. Some effort was needed to get the code 64 bit clean. Secondly a means of extracting the interaction network was needed. Whilst Tierra provided the concept of 'watch bits', which recorded whether a digital organism had accessed another's genome or vice versa, it did not record which other genome was accessed. So I modified the template matching code to log the pair of genome labels that performed the template match to a file.
Having a record of interactions by genotype label, it is necessary to map the genotype to phenotype. In Tierra, the phenotype is the behaviour of the digital organism, and can be judged by running the organisms pairwise in a tournament, to see what effect each has on the other. The precise details for how this can be done is described in [33].
Having a record of interactions between phenotypes, and discarding self-self interactions, there are a number of ways of turning that record into a foodweb. The simplest way, which I adopted, was sum the interactions between each pair of phenotypes over a sliding window of 20 million executed instructions, and doing this every 100 million executed instructions. This lead to time series of around 1000 foodwebs for each Tierra run.
In Tierra, parsimony pressure is controlled by the parameter SlicePow. CPU time is allocated proportional to genome size raised to SlicePow. If SlicePow is close to 0, then there is great evolutionary pressure for the organisms to get as small as possible to increase their replication rate. When it is one, this pressure is eliminated. In [35], I found that a SlicePow of around 0.95 was optimal. If it were much higher, the organisms grow so large and so rapidly that they eventually occupy more than 50% of the soup. At which point they kill the soup at their next Mal (memory allocation) operation. In this work, I altered the implementation of Mal to fail if the request was more than than the soup size divided by minimum population save threshold (usually around 10). Organisms any larger than this will never appear in the Genebanker (Tierra's equivalent of the fossil record), as their population can never exceed the save threshold. This modification allows SlicePow = 1 runs to run for an extensive period of time without the soup dying.
## 7.2 EcoLab
EcoLab was introduced by the author as a simple model of an evolving ecosystem [31]. The ecological dynamics is described by an n -dimensional generalised Lotka-Volterra equation:
<!-- formula-not-decoded -->
where n i is the population density of species i , r i its growth rate and β ij the interaction matrix. Extinction is handled via a novel stochastic truncation algorithm, rather than the more usual threshold method. Speciation occurs by randomly mutating the ecological parameters ( r i and β ij ) of the parents, subject to the constraint that the system remain bounded [30].
The interaction matrix is a candidate foodweb, but has too much information. Its offdiagonal terms may be negative as well as positive, whereas for the complexity definition (6), we need the link weights to be positive. There are a number of ways of resolving this issue, such as ignoring the sign of the off-diagonal term (ie taking its absolute value), or antisymmetrising the matrix by subtracting its transpose, then using the sign of the offdiagonal term to determine the link direction.
For the purposes of this study, I chose to subtract just the negative β ij terms from themselves and their transpose terms β ji . This effects a maximal encoding of the interaction matrix information in the network structure, with link direction and weight encoding the direction and size of resource flow. The effect is as follows:
- Both β ij and β ji are positive (the mutualist case). Neither offdiagonal term changes, and the two nodes have links pointing in both directions, with weights given by the two offdiagonal terms.
- Both β ij and β ji are negative (the competitive case). The terms are swapped, and the signs changed to be positive. Again the two nodes have links pointing in both directions, but the link direction reflects the direction of resource flow.
- Both β ij and β ji are of opposite sign (the predator-prey or parasitic case). Only a single link exists between species i and j , whose weight is the summed absolute values of the offdiagonal terms, and whose link direction reflects the direction of resource flow.
## 7.3 Webworld
Webworld is another evolving ecology model, similar in some respects to EcoLab, introduced by [6], with some modifications described in [12]. It features more realistic ecological interactions than does EcoLab, in that it tracks biomass resources. It too has an interaction matrix called a functional response in that model that could serve as a foodweb, which is converted to a directed weighted graph in the same way as the EcoLab interaction matrix. I used the Webworld implementation distributed with the Eco Lab simulation platform [34]. The parameters were chosen as R = 10 5 , b = 5 × 10 -3 , c = 0 . 8 and λ = 0 . 1.
## 8 Methods and materials
Tierra was run on a 512KB soup, with SlicePow set to 1, until the soup died, typically after some 5 × 10 10 instructions have executed. Some variant runs were performed with SlicePow=0.95, and with different random number generators, but no difference in the outcome was observed.
The source code of Tierra 5.0 was modified in a few places, as described in the Tierra section of this paper. The final source code is available as tierra.5.0.D7.tar.gz from the Eco Lab website hosted on SourceForge (http://ecolab.sf.net).
The genebanker output was processed by the eco-tierra.3.D13 code, also available from the Eco Lab website, to produce a list of phenotype equivalents for each genotype. A function for processing the interaction log file generated by Tierra and producing a timeseries of foodweb graphs was added to Eco-tierra. The script for running this postprocessing step is process ecollog.tcl.
The EcoLab model was adapted to convert the interaction matrix into a foodweb and log the foodweb to disk every 1000 time steps for later processing. The Webworld model was adapted similarly. The model parameters were as documented in the included ecolab.tcl and webworld.tcl experiment files of the ecolab.5.D1 distribution, which is also available from the Eco Lab website.
Finally, each foodweb, whether real world, or ALife generated, and 100 linkshuffled control versions were run through the network complexity algorithm
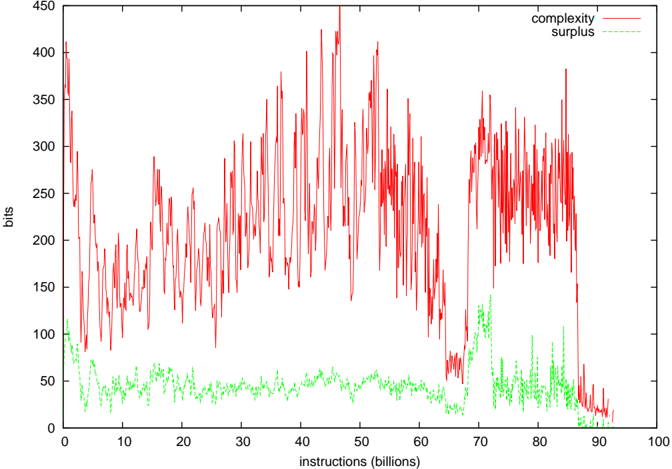
Figure 8: Complexity of the Tierran interaction network for SlicePow=0.95, and the associated complexity surplus. The surplus was typically 15-30 times the standard deviation of the ER neutral network complexities.
<details>
<summary>Image 8 Details</summary>
### Visual Description
## Line Chart: Complexity and Surplus vs. Instructions
### Overview
The image is a line chart comparing "complexity" and "surplus" over a range of instructions (in billions). The x-axis represents the number of instructions (in billions), and the y-axis represents the value in bits. The "complexity" is represented by a solid red line, and the "surplus" is represented by a dashed green line.
### Components/Axes
* **X-axis:** "instructions (billions)" with tick marks every 10 billion instructions from 0 to 100.
* **Y-axis:** "bits" with tick marks every 50 bits from 0 to 450.
* **Legend (top-right):**
* "complexity" - solid red line
* "surplus" - dashed green line
### Detailed Analysis
* **Complexity (Red Line):**
* **Trend:** The complexity line starts high, fluctuates significantly, then decreases sharply after approximately 70 billion instructions.
* **Data Points:**
* Starts at approximately 400 bits at 0 billion instructions.
* Fluctuates between approximately 100 and 450 bits between 0 and 70 billion instructions.
* Drops to approximately 200 bits at 70 billion instructions.
* Decreases to approximately 0 bits by 90 billion instructions.
* **Surplus (Green Dashed Line):**
* **Trend:** The surplus line starts relatively high, fluctuates, then increases around 70 billion instructions before decreasing to zero.
* **Data Points:**
* Starts at approximately 75 bits at 0 billion instructions.
* Fluctuates between approximately 30 and 60 bits between 0 and 70 billion instructions.
* Increases to approximately 125 bits around 70 billion instructions.
* Decreases to approximately 0 bits by 90 billion instructions.
### Key Observations
* The complexity exhibits high volatility throughout the initial range of instructions.
* Both complexity and surplus decrease to zero after 70 billion instructions.
* There is an inverse relationship between complexity and surplus around 70 billion instructions, where complexity decreases and surplus increases.
### Interpretation
The chart illustrates the relationship between complexity and surplus as a function of the number of instructions processed. The initial high complexity suggests a period of intense processing or high computational load. The subsequent decrease in complexity, coupled with an increase in surplus, may indicate a shift towards more efficient processing or resource optimization. The eventual convergence of both complexity and surplus to zero suggests a completion of the task or a state of equilibrium. The data suggests that the system undergoes a significant change in behavior around 70 billion instructions, possibly due to an optimization or a change in the nature of the instructions being processed.
</details>
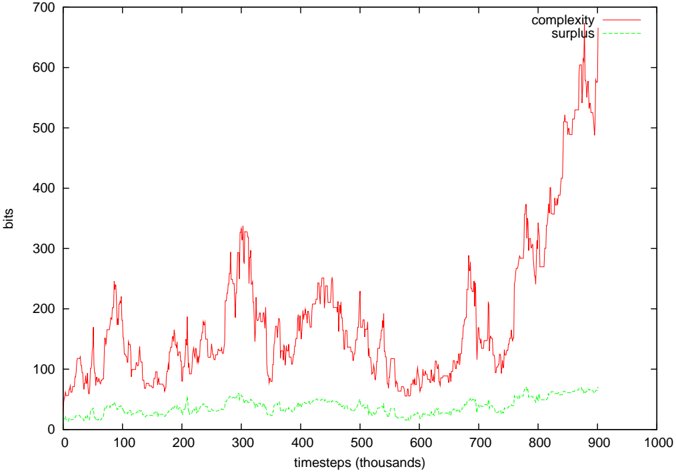
(6). This is documented in the cmpERmodel.tcl script of ecolab.5.D1. The average and standard deviation of ln C was calculated, rather than C directly, as the shuffled complexity values fitted a log-normal distribution better than a standard normal distribution. The difference between the measured complexity and exp 〈 ln C〉 (ie the geometric mean of the control network complexities) is what is reported as the surplus in Figures 8-10.
## 9 ALife results
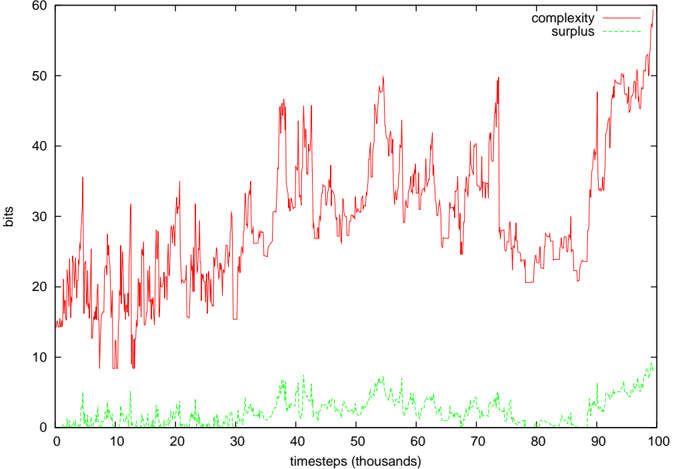
Figures 8-10 show the computed complexity values and the surplus of the complexity over the average randomly shuffled networks. In both the EcoLab and Tierra cases, significant amounts of surplus complexity can be observed, but not in the case of WebWorld. In WebWorld's case, this is probably because the system complexity was not very high in the first place - the diversity typically ranged between 5 and 10 species throughout the WebWorld run. It is possible that the competition parameter c was too high for this run to generate any appreciable foodweb complexity.
An earlier report of these results [37] did not show significant complexity,
Figure 9: Complexity of EcoLab's foodweb, and the associated complexity surplus. For most of the run, the surplus was 10-40 times the standard deviation of the ER neutral network complexities, but in the final complexity growth phase, it was several 100.
<details>
<summary>Image 9 Details</summary>
### Visual Description
## Line Chart: Complexity and Surplus over Time
### Overview
The image is a line chart that plots "complexity" and "surplus" over time. The x-axis represents time in thousands of steps, ranging from 0 to 1000. The y-axis represents the value in bits, ranging from 0 to 700. The chart displays two data series: "complexity" shown in red, and "surplus" shown in green.
### Components/Axes
* **X-axis:**
* Label: "timesteps (thousands)"
* Scale: 0 to 1000, with markers at 0, 100, 200, 300, 400, 500, 600, 700, 800, 900, and 1000.
* **Y-axis:**
* Label: "bits"
* Scale: 0 to 700, with markers at 0, 100, 200, 300, 400, 500, 600, and 700.
* **Legend:** Located in the top-right corner.
* "complexity" - represented by a solid red line.
* "surplus" - represented by a dashed green line.
### Detailed Analysis
* **Complexity (Red Line):**
* Trend: The complexity fluctuates significantly over time. Initially, it oscillates between approximately 50 and 250 bits for the first 750,000 timesteps. After 750,000 timesteps, the complexity exhibits a sharp upward trend, reaching a peak of approximately 650 bits near the end of the observed period.
* Data Points:
* At timestep 0: ~50 bits
* At timestep 300,000: ~320 bits
* At timestep 500,000: ~150 bits
* At timestep 750,000: ~100 bits
* At timestep 900,000: ~650 bits
* **Surplus (Green Dashed Line):**
* Trend: The surplus remains relatively stable over time, with minor fluctuations. It generally stays between 20 and 60 bits throughout the entire period.
* Data Points:
* At timestep 0: ~30 bits
* At timestep 500,000: ~40 bits
* At timestep 900,000: ~60 bits
### Key Observations
* The complexity exhibits significant volatility, with several peaks and troughs before timestep 750,000.
* The surplus remains relatively constant compared to the complexity.
* There is a dramatic increase in complexity after timestep 750,000, suggesting a potential shift in the underlying system's behavior.
### Interpretation
The chart illustrates the relationship between complexity and surplus over time. The data suggests that the system initially operates with fluctuating complexity and a relatively stable surplus. However, after a certain point (750,000 timesteps), the complexity undergoes a significant increase, potentially indicating a phase transition or a change in the system's dynamics. The surplus, on the other hand, does not exhibit a similar increase, suggesting that it may not be directly correlated with the observed increase in complexity. The sudden jump in complexity could be due to external factors or internal dynamics that trigger a more complex state. Further investigation would be needed to understand the underlying mechanisms driving these changes.
</details>
Figure 10: Complexity of Webworld's foodweb, and the associated complexity surplus. The surplus was typically 1-5 times the standard deviation of the ER neutral network complexities.
<details>
<summary>Image 10 Details</summary>
### Visual Description
## Line Chart: Complexity and Surplus over Time
### Overview
The image is a line chart that plots "complexity" and "surplus" over time. The x-axis represents time in thousands of steps, ranging from 0 to 100. The y-axis represents the value in "bits", ranging from 0 to 60. The "complexity" is represented by a solid red line, and the "surplus" is represented by a dashed green line.
### Components/Axes
* **X-axis:**
* Label: "timesteps (thousands)"
* Scale: 0 to 100, with tick marks at intervals of 10.
* **Y-axis:**
* Label: "bits"
* Scale: 0 to 60, with tick marks at intervals of 10.
* **Legend (top-right):**
* "complexity" - solid red line
* "surplus" - dashed green line
### Detailed Analysis
* **Complexity (Red Line):**
* Trend: Initially fluctuates between approximately 15 and 25 bits for the first 30,000 timesteps. Between 30,000 and 80,000 timesteps, the complexity fluctuates between approximately 25 and 45 bits. After 80,000 timesteps, the complexity increases sharply, reaching approximately 55 bits at 100,000 timesteps.
* Approximate Values:
* At timestep 0: ~15 bits
* At timestep 30,000: ~30 bits
* At timestep 50,000: ~35 bits
* At timestep 80,000: ~30 bits
* At timestep 100,000: ~55 bits
* **Surplus (Green Dashed Line):**
* Trend: Remains relatively low and stable between 0 and 5 bits for the first 80,000 timesteps. After 80,000 timesteps, the surplus begins to increase, reaching approximately 10 bits at 100,000 timesteps.
* Approximate Values:
* At timestep 0: ~2 bits
* At timestep 80,000: ~2 bits
* At timestep 100,000: ~10 bits
### Key Observations
* The complexity exhibits significant fluctuations throughout the entire period.
* The surplus remains low for a long period before increasing towards the end.
* Both complexity and surplus increase significantly after 80,000 timesteps.
### Interpretation
The chart suggests that the system's complexity and surplus are initially relatively stable. However, after a certain point (around 80,000 timesteps), both measures experience a notable increase. This could indicate a phase transition or a significant change in the system's dynamics. The initial stability in surplus, followed by a later increase, might suggest a build-up of resources or potential before a period of growth or change. The fluctuations in complexity could represent ongoing adaptation or exploration within the system. The correlation between the increase in complexity and surplus after 80,000 timesteps suggests that they are related, possibly with increased complexity leading to a higher surplus.
</details>
unlike what is shown here. It was discovered that the process of writing the foodweb data to an intermediate file stripped the weight data from the network links, preserving only the graph's topological structure. It turns out that the complexity surplus phenomenon is only present in weighted networks - unweighted networks tend to be indistinguishable from randomly generated networks.
## 10 Discussion
In this paper, a modified version of a previously proposed simple representation language for n -node directed graphs is given. This modification leads to a more intuitive complexity measure that gives a minimum value for empty and full networks. When compared with the zcomplexity measure introduced in the previous paper, it is somewhat correlated with, and is at least as good a measure as zcomplexity, but has the advantage of being far more tractable.
When compared with random networks created by shuffling the links, all real world example networks exhibited higher complexity than the random network. The difference between the complexity of the real network and the mean of the complexities of the randomly shuffled network represents structural information contained in the specific arrangement of links making up the network. To test the hypothesis that networks generated by evolutionary selection will also have a complexity surplus representing the functional nature of the network, I ran the same analysis on foodwebs generated by various artificial life systems. These too, exhibited the same complexity surplus seen in the real world networks.
By contrast, I have applied this technique to some scale free networks generated by the Barab´ asi-Albert preferential attachment process [4], with varying numbers of attachment points per node and different weight distributions. On none of these graphs were significant complexity surpluses generated.
If the weights are stripped off the edges, then the effect disappears, which provides a clue as to the origin of the effect. In equation (6), the components dominating the sum will tend to be connected giant components of the overall network with very few cycles. The randomly shuffled versions of these, however, will tend to be composed of small disconnected components, with many of the same motifs represented over and over again. Consequently, the shuffled network has a great deal of symmetry, reducing the overall complexity. Even though naively one might think that random networks should be the most complex within the class of networks of a given node and link count, given the nature of Kolmogorov complexity to favour random strings, the weighted complexity formula (6) is sensitive to the structure within the network that has meaning to system behaviour that the network represents.
Reverting to an earlier posed question about the amount of complexity stored within the foodweb as compared with individual organism's phenotypic complexity, the results in Figure 8 gives a preliminary answer. Each foodweb in Figure 8 averaged around 100 species, the average phenotypic complexity of which is around 150-200 bits (750 bits at maximum)[33]. So the 250-400 bits of network complexity is but a drop in the bucket of phenotypic complexity.
## References
- [1] Almunia, J., Basterretxea, G., Aristegui, J., and Ulanowicz., R. (1999) Benthicpelagic switching in a coastal subtropical lagoon. Estuarine, Coastal and Shelf Science , 49 , 363-384.
- [2] Baird, D., Luczkovich, J., and Christian, R. R. (1998) Assessment of spatial and temporal variability in ecosystem attributes of the St Marks National Wildlife Refuge, Apalachee Bay, Florida. Estuarine, Coastal, and Shelf Science , 47 , 329-349.
- [3] Baird, D. and Ulanowicz, R. (1989) The seasonal dynamics of the Chesapeake Bay ecosystem. Ecological Monographs , 59 , 329-364.
- [4] Barab´ asi, A.-L. and Albert, R. (1999) Emergence of scaling in random networks. Science , 286 , 509-512.
- [5] Bedau, M. A., McCaskill, J. S., Packard, N. H., Rasmussen, S., Adami, C., Green, D. G., Ikegami, T., Kaneko, K., and Ray, T. S. (2000) Open problems in artificial life. Artificial Life , 6 , 363-376.
- [6] Caldarelli, G., Higgs, P. G., and McKane, A. J. (1998) Modelling coevolution in multispecies communities. J. Theor, Biol. , 193 , 345-358.
- [7] Christian, R. and Luczkovich, J. (1999) Organizing and understanding a winter's seagrass foodweb network through effective trophic levels. Ecological Modelling , 117 , 99-124.
- [8] Clauset, A., Shalizi, C. R., and Newman, M. E. J. (2009) Power-law distributions in empirical data. SIAM Review , 51 , 661-703, arXiv:0706.1062.
- [9] Claussen, J. C. (2007) Offdiagonal complexity: A computationally quick complexity measure for graphs and networks. Physica A , 375 , 365-373, arXiv:q-bio.MN/0410024.
- [10] Dewar, R. (2003) Informational theory explanation of the fluctuation theorem, maximum entropy production and self-organized criticality in nonequilibrium stationary states. J. Phys. A , 36 , 631-641.
- [11] Dorin, A. and Korb, K. B. (2010) Network measures of ecosystem complexity. Fellerman et al. [15], pp. 323-328.
- [12] Drossel, B., Higgs, P. G., and McKane, A. J. (2001) The influence of predator-prey population dynamics on the long-term evolution of food web structure. J. Theor. Biol. , 208 , 91-107.
- [13] Duch, J. and Arenas, A. (2005) Community identification using extremal optimization. Physical Review E , 72 , 027104.
- [14] Edmonds, B. (1999) Syntactic Measures of Complexity . Ph.D. thesis, University of Manchester, http://bruce.edmonds.name/thesis/, Complexity Bibliography: http://bruce.edmonds.name/compbib/.
- [15] Fellerman, H. et al. (eds.) (2010) Proceedings of Artificial Life XII , MIT Press.
- [16] G¨ ornerup, O. and Crutchfield, J. P. (2008) Hierarchical self-organization in the finitary process soup. Artificial Life , 14 , 245-254.
- [17] Hagy, J. (2002) Eutrophication, hypoxia and trophic transfer efficiency in Chesapeake Bay . Ph.D. thesis, University of Maryland at College Park (USA).
- [18] Jeong, H., Mason, S., Barab´ asi, A.-L., and Oltvai, Z. N. (2001) Centrality and lethality of protein networks. Nature , 411 , 41.
- [19] Kim, J. and Wilhelm, T. (2008) What is a complex graph. Physica A , 387 , 2637-2652.
- [20] Knuth, D. E. (1993) The Stanford GraphBase: A Platform for Combinatorial Computing . Addison-Wesley.
- [21] Layzer, D. (1988) Growth and order in the universe. Weber, B. H., Depew, D. J., and Smith, J. D. (eds.), Entropy, Information and Evolution , pp. 23-39, MIT Press.
- [22] Li, M. and Vit´ anyi, P. (1997) An Introduction to Kolmogorov Complexity and its Applications . Springer, 2nd edn.
- [23] McKay, B. D. (1981) Practical graph isomorphism. Congressus Numerantium , 30 , 45-87.
- [24] Monaco, M. and Ulanowicz, R. (1997) Comparative ecosystem trophic structure of three U.S. Mid-Atlantic estuaries. Mar. Ecol. Prog. Ser. , 161 , 239-254.
- [25] Myrvold, W. and Ruskey, F. (2001) Ranking and unranking permutations in linear time. Information Processing Letters , 79 , 281-284.
- [26] Newman, M. E. J. (2006) Finding community structure in networks using the eigenvectors of matrices. Phys. Rev. E , 74 , 036104.
- [27] Prigogine, I. (1980) From Being into Becoming . W.H. Freeman.
- [28] Prokopenko, M., Boschetti, F., and Ryan, A. J. (2009) An informationtheoretic primer on complexity, self-organization, and emergence. Complexity , 15 , 11-28.
- [29] Ray, T. (1991) An approach to the synthesis of life. Langton, C. G., Taylor, C., Farmer, J. D., and Rasmussen, S. (eds.), Artificial Life II , p. 371, Addison-Wesley.
- [30] Standish, R. K. (2000) The role of innovation within economics. Barnett, W., Chiarella, C., Keen, S., Marks, R., and Schnabl, H. (eds.), Commerce, Complexity and Evolution , vol. 11 of International Symposia in Economic Theory and Econometrics , pp. 61-79, Cambridge UP.
- [31] Standish, R. K. (1994) Population models with random embryologies as a paradigm for evolution. Complexity International , 2 .
- [32] Standish, R. K. (2001) On complexity and emergence. Complexity International , 9 , arXiv:nlin.AO/0101006.
- [33] Standish, R. K. (2003) Open-ended artificial evolution. International Journal of Computational Intelligence and Applications , 3 , 167, arXiv:nlin.AO/0210027.
- [34] Standish, R. K. (2004) Ecolab, Webworld and self-organisation. Pollack et al. (eds.), Artificial Life IX , Cambridge, MA, p. 358, MIT Press.
- [35] Standish, R. K. (2004) The influence of parsimony and randomness on complexity growth in Tierra. Bedau et al. (eds.), ALife IX Workshop and Tutorial Proceedings , pp. 51-55, arXiv:nlin.AO/0604026.
- [36] Standish, R. K. (2005) Complexity of networks. Abbass et al. (eds.), Recent Advances in Artificial Life , Singapore, vol. 3 of Advances in Natural Computation , pp. 253-263, World Scientific, arXiv:cs.IT/0508075.
- [37] Standish, R. K. (2010) Network complexity of foodwebs. Fellerman et al. [15], pp. 337-343.
- [38] Standish, R. K. (2010) SuperNOVA: a novel algorithm for graph automorphism calculations. Journal of Algorithms - Algorithms in Cognition, Informatics and Logic , submitted, arXix: 0905.3927.
- [39] Ulanowicz, R., Bondavalli, C., and Egnotovich, M. (1998) Network analysis of trophic dynamics in South Florida ecosystem, FY 97: The Florida Bay ecosystem. Tech. rep., Chesapeake Biological Laboratory, Solomons, MD 20688-0038 USA, [UMCES]CBL 98-123.
- [40] Ulanowicz, R., Heymans, J., and Egnotovich., M. (2000) Network analysis of trophic dynamics in South Florida ecosystems, FY 99: The graminoid ecosystem. Tech. rep., Chesapeake Biological Laboratory, Solomons, MD 20688-0038 USA, [UMCES] CBL 00-0176.
- [41] Watts, D. J. and Strogatz, S. H. (1998) Collective dynamics of 'small-world' networks. Nature , 393 , 409-10.
- [42] Wilhelm, T. and Hollunder, J. (2007) Information theoretic description of networks. Physica A , 385 , 385-396.
Table 1: Complexity values of several freely available network datasets. celegansneural, lesmis and adjnoun are available from Mark Newman's website, representing the neural network of the C. elegans nematode[41], the coappearance of characters in the novel Les Mis´ erables by Victor Hugo[20] and the adjacency network of common adjectives and nouns in the novel David Copperfield by Charles Dickens[26]. The metabolic data of C. elegans [13] and protein interaction network in yeast[18] are available from Duncan Watt's website. PA1 and PA3 are networks generated via preferential attachment with in degree of one or three respectively, and uniformly distributed link weights. The other datasets are food webs available from the Pajek website [7, 39, 40, 17, 2, 1, 3]. For each network, the number of nodes and links are given, along with the computed complexity C . In the fourth column, the original network is shuffled 1000 times, and the logarithm of the complexity is averaged ( 〈 ln C ER 〉 ). The fifth column gives the difference between these two values, which represents the information content of the specific arrangement of links. The final column gives a measure of the significance of this difference in terms of the number of standard deviations ('sigmas') of the distribution of shuffled networks. In two examples, the distributions of shuffled networks had zero standard deviation, so ∞ appears in this column.
| Dataset | nodes | links | C | e 〈 ln C ER 〉 | C - e 〈 ln C ER 〉 | | ln C-〈 ln C ER 〉| σ ER |
|-------------------|---------|---------|---------|-----------------|---------------------|--------------------------------------|
| celegansneural | 297 | 2345 | 442.7 | 251.6 | 191.1 | 29 |
| lesmis | 77 | 508 | 199.7 | 114.2 | 85.4 | 24 |
| adjnoun | 112 | 850 | 3891 | 3890 | 0.98 | ∞ |
| yeast | 2112 | 4406 | 33500.6 | 30218.2 | 3282.4 | 113.0 |
| celegansmetabolic | 453 | 4050 | 25421.8 | 25387.2 | 34.6 | ∞ |
| baydry | 128 | 2138 | 126.6 | 54.2 | 72.3 | 22 |
| baywet | 128 | 2107 | 128.3 | 51 | 77.3 | 20 |
| cypdry | 71 | 641 | 85.7 | 44.1 | 41.5 | 13 |
| cypwet | 71 | 632 | 87.4 | 42.3 | 45 | 14 |
| gramdry | 69 | 911 | 47.4 | 31.6 | 15.8 | 10 |
| gramwet | 69 | 912 | 54.5 | 32.7 | 21.8 | 12 |
| Chesapeake | 39 | 177 | 66.8 | 45.7 | 21.1 | 10.4 |
| ChesLower | 37 | 178 | 82.1 | 62.5 | 19.6 | 10.6 |
| ChesMiddle | 37 | 208 | 65.2 | 48 | 17.3 | 9.3 |
| ChesUpper | 37 | 215 | 81.8 | 60.7 | 21.1 | 10.2 |
| CrystalC | 24 | 126 | 31.1 | 24.2 | 6.9 | 6.4 |
| CrystalD | 24 | 100 | 31.3 | 24.2 | 7 | 6.2 |
| Everglades | 69 | 912 | 54.5 | 32.7 | 21.8 | 11.8 |
| Florida | 128 | 2107 | 128.4 | 51 | 77.3 | 20.1 |
| Maspalomas | 24 | 83 | 70.3 | 61.7 | 8.6 | 5.3 |
| Michigan | 39 | 219 | 47.6 | 33.7 | 14 | 9.5 |
| Mondego | 46 | 393 | 45.2 | 32.2 | 13 | 10.0 |
| Narragan | 35 | 219 | 58.2 | 39.6 | 18.6 | 11.0 |
| Rhode | 19 | 54 | 36.3 | 30.3 | 6 | 5.3 |
| StMarks | 54 | 354 | 110.8 | 73.6 | 37.2 | 16.0 |
| PA1 | 100 | 99 | 98.9 | 85.4 | 13.5 | 2.5 |
| PA3 | 100 | 177 | 225.9 | 207.3 | 18.6 | 3.0 |