## Causal Inference Using Tractable Circuits
## Adnan Darwiche
Computer Science Department University of California, Los Angeles darwiche@cs.ucla.edu
## Abstract
The aim of this paper is to discuss a recent result which shows that probabilistic inference in the presence of (unknown) causal mechanisms can be tractable for models that have traditionally been viewed as intractable. This result was reported recently in [15] to facilitate model-based supervised learning but it can be interpreted in a causality context as follows. One can compile a non-parametric causal graph into an arithmetic circuit that supports inference in time linear in the circuit size. The circuit is also non-parametric so it can be used to estimate parameters from data and to further reason (in linear time) about the causal graph parametrized by these estimates. Moreover, the circuit size can sometimes be bounded even when the treewidth of the causal graph is not, leading to tractable inference on models that have been deemed intractable previously. This has been enabled by a new technique that can exploit causal mechanisms computationally but without needing to know their identities (the classical setup in causal inference). Our goal is to provide a causality-oriented exposure to these new results and to speculate on how they may potentially contribute to more scalable and versatile causal inference.
## 1 Introduction
Tractable arithmetic circuits have been receiving an increased attention in AI and computer science more broadly; see [16] for a recent survey. These circuits represent real-valued functions and are called tractable because they allow one to answer some hard queries about these functions through linear-time, feed-forward passes on the circuit structure. These circuits were initially compiled from Bayesian networks as proposed in [13, 12] to facilitate probabilistic reasoning. They were later learned from data, starting with [23], and even handcrafted as initially proposed in [30]. Traditional methods for exact probabilistic inference have a complexity which is exponential in treewidth (a graph-theoretic parameter that measures the model's connectivity). With the introduction of compiled circuits, one could practically do inference on models whose treewidth can be in the hundreds; see, e.g., [5, 8, 7]. A recent comprehensive, empirical evaluation of at least a dozen probabilistic inference algorithms showed that methods based on circuits are at the forefront in terms of efficiency [1]; see also [18]. What stands behind the efficacy of circuit-based methods is their ability to aggresively exploit the parametric structure of models such as functional dependencies and context-specific independence [4]. This however has limited their utility in contexts where one does not know the model parameters, which is a common case in causal inference. A circuit compilation method was recently proposed in [15] which can exploit a more abstract form of parametric structure: functional dependencies (i.e., causal mechanisms) whose identities are unknown which is the classical setup in causal inference. This new method was able to compile circuits for models whose treewidth is very large without needing to know the model parameters, leading to a major advance on earlier techniques. Our aim in this paper is to provide an intuitive exposure to this new algorithm and its underlying techniques, while placing it in a causality context. Our belief is that a discussion of these new results could lead to a synthesis on how they can further advance causal inference in terms of scalability and versatility. We start first with a review of some core concepts in causality and circuit-based inference and then follow by a discussion of the new results and their significance.
## 2 Causal Models and Queries
Variables are discrete and denoted by uppercase letters (e.g., X ) and their values by lowercase letters (e.g., x ). Sets of variables are denoted by boldface, uppercase letters (e.g., X ) and their instantiations by boldface, lowercase letters (e.g., x ). We will write x ∈ x to mean that x is the value of variable X in instantiation x . For a binary variable X , we will use x and ¯ x to denote X =1 and X =0 , respectively.
We next define Structural Causal Models (SCMs) following the treatment in [21]; see also [26].
Definition 1. An SCM is a tuple M = ( U , V , F , { Pr ( U ) } U ∈ U ) where U and V are disjoint sets of variables called 'exogenous' and 'endogenous,' respectively; F contains exactly one function f V for each endogenous variable V ; and Pr ( U ) are distributions over exogenous variables U . A function f V is called a 'causal mechanism' and it determines the value of variable V based on two sets of inputs, U V ⊆ U and V V ⊆ V . That is, the mechanism f V is a mapping U V , V V ↦→ V .
The causal graph G of an SCM contains variables U ∪ V as its nodes. It also contains edges X → V for each endogenous variable V and each variable X that is an input to function f V . We will only deal with SCMs that produce acyclic causal graphs (the inputs of a function cannot depend on its output). Moreover, we will assume that exogenous variables are independent and that only endogenous variables can be observed. A key observation about SCMs is that once we fix the values of exogenous variables, the values of all endogenous variables are also fixed by the causal mechanisms.
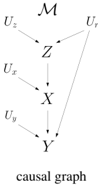
We will next consider an example SCM from [2] where all variables are binary. The endogenous variables are V = X,Y,Z , representing a treatment, the outcome and the presence of hypertension, respectively. The exogenous variables are U = U r , U x , U y , U z , representing natural resistance to disease ( U r ) and sources of variation affecting endogenous variables ( U x , U y , U z ). The distributions of exogenous variables are Pr ( u r ) = 0 . 25 , Pr ( u x ) = 0 . 9 , Pr ( u y ) = 0 . 7 and Pr ( u z ) = 0 . 95 . The three causal mechanisms are given next and they lead to the causal graph on the right:
<!-- formula-not-decoded -->
<details>
<summary>Image 1 Details</summary>
### Visual Description
\n
## Diagram: Causal Graph
### Overview
The image depicts a directed acyclic graph representing causal relationships between variables. The graph consists of nodes labeled with letters (M, Z, X, Y) and variables (U<sub>x</sub>, U<sub>y</sub>, U<sub>z</sub>, U). Arrows indicate the direction of causal influence. The diagram is labeled "causal graph" at the bottom.
### Components/Axes
The diagram consists of the following nodes:
* **M**: A top-level node.
* **Z**: A node influenced by M and U<sub>z</sub>.
* **X**: A node influenced by Z and U<sub>x</sub>.
* **Y**: A node influenced by X and U<sub>y</sub>.
* **U<sub>x</sub>**: A variable influencing X.
* **U<sub>y</sub>**: A variable influencing Y.
* **U<sub>z</sub>**: A variable influencing Z.
* **U**: A variable influencing M and Z.
The diagram uses directed edges (arrows) to represent causal relationships.
### Detailed Analysis or Content Details
The causal relationships are as follows:
* M influences Z.
* U influences M.
* U influences Z.
* U<sub>z</sub> influences Z.
* Z influences X.
* U<sub>x</sub> influences X.
* X influences Y.
* U<sub>y</sub> influences Y.
There are no numerical values or scales present in the diagram. It is a purely structural representation of causal relationships.
### Key Observations
The graph shows a hierarchical structure with M at the top and Y at the bottom. Each node (except M) is influenced by at least two other nodes or variables. The variables U<sub>x</sub>, U<sub>y</sub>, and U<sub>z</sub> represent unobserved confounders or exogenous variables influencing the respective nodes X, Y, and Z. The variable U influences both M and Z, suggesting a common cause.
### Interpretation
This causal graph illustrates a system where variables are interconnected through causal pathways. The presence of unobserved variables (U<sub>x</sub>, U<sub>y</sub>, U<sub>z</sub>, U) suggests that the observed relationships between M, Z, X, and Y may be subject to confounding. The graph can be used to reason about the effects of interventions on different variables. For example, if we were to intervene on M, we could predict the downstream effects on Z, X, and Y, taking into account the influence of the unobserved variables. The diagram is a simplified representation of a complex system, and the causal relationships may not be fully accurate or complete. It is a tool for modeling and understanding potential causal mechanisms.
</details>
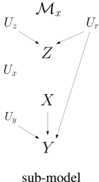
A key notion in causal inference is the sub-model M z of an SCM M , where z is an instantiation of some endogenous variables. This is another SCM obtained from M by replacing the function for each variable Z ∈ Z with the constant function f Z = z , where z ∈ z . Intuitively, the sub-model M z is used to reason about an intervention from outside the system modeled by M , which suppresses the causal mechanisms of variables Z and fixes the values of these variables to z . For example, if we intervene to administer a treatment ( x ) in the above SCM M , we get a sub-model M x in which the mechanism for variable X is replaced with the constant function f X = x . The causal graph of the resulting sub-model is shown on the right.
sub-model
<details>
<summary>Image 2 Details</summary>
### Visual Description
\n
## Diagram: Sub-Model Relationship
### Overview
The image presents a directed acyclic graph (DAG) representing relationships within a "sub-model". The diagram illustrates dependencies between variables labeled U<sub>x</sub>, U<sub>y</sub>, U<sub>z</sub>, X, Y, and Z, with M<sub>x</sub> appearing at the top. Arrows indicate the direction of influence or dependency.
### Components/Axes
The diagram consists of nodes (labeled with letters and subscripts) and directed edges (arrows). The nodes represent variables or components within the sub-model. The edges represent relationships between these components. The label "sub-model" is positioned at the bottom center of the diagram.
### Detailed Analysis or Content Details
The diagram shows the following relationships:
* **M<sub>x</sub>** is connected to **Z** and **U<sub>f</sub>** via directed arrows.
* **U<sub>z</sub>** is connected to **Z** via a directed arrow.
* **U<sub>x</sub>** is connected to **Z** via a directed arrow.
* **Z** is connected to **X** via a directed arrow.
* **U<sub>y</sub>** is connected to **X** via a directed arrow.
* **X** is connected to **Y** via a directed arrow.
* **U<sub>f</sub>** is connected to **Y** via a directed arrow.
There are no numerical values or scales present in the diagram. It is a purely structural representation of relationships.
### Key Observations
The diagram suggests a hierarchical structure. M<sub>x</sub> appears to be a root node influencing Z and U<sub>f</sub>. Z then influences X, and X influences Y. U<sub>y</sub> and U<sub>f</sub> also directly influence Y. The diagram does not indicate the nature of the relationships (e.g., causal, correlational).
### Interpretation
This diagram likely represents a simplified model of a system, where the variables represent different components or factors. The arrows indicate how changes in one variable might affect others. The structure suggests that M<sub>x</sub> and U<sub>z</sub>, U<sub>x</sub> are upstream influences on Z, which in turn influences X and ultimately Y. U<sub>y</sub> and U<sub>f</sub> provide additional direct influence on Y. The diagram is abstract and requires further context to understand the specific meaning of the variables and relationships. It could represent a Bayesian network, a causal diagram, or a similar type of model used in various fields like statistics, machine learning, or engineering. The absence of quantitative data suggests this is a conceptual model rather than a quantitative analysis.
</details>
Three types of queries are normally posed on SCMs, which are referred to as associational, interventional and counterfactual queries. They correspond to what is known as the causal hierarchy [26, 28, 27], where each class of queries belongs to a rung [28] or layer [2] in the hierarchy. We next define the syntax and semantics of these queries, using a slightly different notation than is customary-this will allow us to provide a more uniform and general treatment of these queries.
Definition 2. An 'observational event' has the form x where X is a set of endogenous variables. An 'interventional event' has the form y x where X and Y are sets of endogenous variables. A 'counterfactual event' has the form η 1 , . . . , η n where η i is an observational or interventional event.
An observational event x says that variables X took the value x . For example, a patient did not take the treatment and died ( ¯ x, ¯ y ). An interventional event y x says that variables Y took the value y after setting variables X to x by an intervention. For example, a patient survived after they were given the treatment ( y x ). A counterfactual event is a conjunction of events where each could be observational or interventional. For example, a patient who responds to treatment did not take it and died ( y x , ¯ x, ¯ y ).
Definition 3. A 'world' for SCM M is an instantiation of its exogenous variables. The worlds of an observational event W M ( x ) are those worlds that fix the values of variables X to x . The worlds of an interventional event W M ( y x ) are defined as W M x ( y ) . The worlds of a counterfactual event W M ( η 1 , . . . , η n ) are defined as W M ( η 1 ) ∩ . . . ∩ W M ( η n ) .
The following table, borrowed from [2], shows all sixteen worlds of the above SCM M , together with their probabilities and the unique states they entail for endogenous variables.
| | Ur | Uz | xn | Uy | Z | X | Y | P(u) | | Ur | ²n | xn | Uy | Z | X | Y | P(u) |
|----|------|------|------|------|-----|-----|-----|----------|----|------|------|------|------|-----|-----|-----|----------|
| 1 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0.001125 | 9 | 1 | 0 | 0 | 0 | 0 | 1 | 1 | 0.000375 |
| 2 | 0 | 0 | 0 | 1 | 0 | 1 | 0 | 0.002625 | 10 | 1 | 0 | 0 | 1 | 0 | 1 | 1 | 0.000875 |
| 3 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 0.010125 | 11 | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 0.003375 |
| 4 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0.023625 | 12 | 1 | 0 | 1 | 1 | 0 | 0 | 1 | 0.007875 |
| 5 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 0.021375 | 13 | 1 | 1 | 0 | 0 | 1 | 0 | 0 | 0.007125 |
| 6 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | 0.049875 | 14 | 1 | 1 | 0 | 1 | 1 | 0 | 1 | 0.016625 |
| 7 | 0 | 1 | 1 | 0 | 0 | 0 | 1 | 0.192375 | 15 | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 0.064125 |
| 8 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 0.448875 | 16 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0.149625 |
Using this table and a similar one for sub-model M x , we obtain W M (¯ x, ¯ y ) = { u 4 , u 8 , u 11 , u 13 } and W M ( y x ) = { u 9 , . . . , u 16 } which leads to W M ( y x , ¯ x, ¯ y ) = W M (¯ x, ¯ y ) ∩ W M ( y x ) = { u 11 , u 13 } .
We are now ready to define the probability of any SCM event.
Definition 4. Let M be an SCM with exogenous variables U and distributions Pr ( U ) for U ∈ U . The probability of event η with respect to SCM M is defined as:
<!-- formula-not-decoded -->
Hence, Pr (¯ x, ¯ y ) = Pr ( u 4 ) + Pr ( u 8 ) + Pr ( u 11 ) + Pr ( u 13 ) = 0 . 4830 and Pr ( y x , ¯ x, ¯ y ) = 0 . 0105 . We can now compute the probability that a patient who did not take the treatment and died would have been alive had they been given the treatment, Pr ( y x | ¯ x, ¯ y ) = Pr ( y x , ¯ x, ¯ y ) / Pr (¯ x, ¯ y ) = 0 . 0217 . We will focus next on associational and interventional queries, leaving counterfactuals to future work.
## 3 Causal Inference Using Circuits
As we just saw, one can answer sophisticated causal queries based on a fully specified SCM. However, such a model may not be available, particularly the identities of causal mechanisms and the distributions over exogenous variables. What is more common is to have the causal graph of an underlying SCM in addition to observational data about endogenous variables. A key task of causal inference is then to draw conclusions based on this limited input, particularly about interventional probabilities which is the task we shall focus on. We will next show how this cross-layer inference can be realized using feed-forward circuits that are compiled from the non-parametric causal graphs of SCMs. The discussion will reveal the significance of compiling the smallest possible circuit for a causal graph. It will also motivate the new compilation algorithm in [15] which is particularly relevant to the causal graphs of SCMs (in contrast to Bayesian networks). We shall discuss and study further this algorithm in Sections 4-6 where we will also contribute to understanding its complexity.
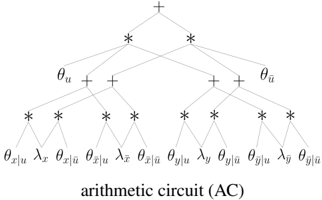
The Circuits of Causal Graphs Consider the causal graph X ← U → Y over binary variables and its compiled arithmetic circuit (AC) shown on the right. This circuit has two types of inputs: symbolic parameters θ and indicators λ . There are two parameters for exogenous variable U ( θ u and θ ¯ u ) which specify its distribution. There are four parameters for endogenous variable X ( θ x | u , θ ¯ x | u , θ x | ¯ u , θ ¯ x | ¯ u ) which specify its causal mechanism. The remaining four parameters specify the mechanism for endogenous variable Y . The indicators correspond to the
<details>
<summary>Image 3 Details</summary>
### Visual Description
\n
## Diagram: Arithmetic Circuit (AC)
### Overview
The image depicts a binary tree diagram representing an arithmetic circuit (AC). The circuit performs a series of multiplications and additions, culminating in a final addition at the root. The diagram is structured hierarchically, with leaf nodes representing variables and intermediate nodes representing operations.
### Components/Axes
The diagram consists of the following components:
* **Leaf Nodes:** Represented by variables: θ<sub>xu</sub>, λ<sub>x</sub>, x<sub>r|a</sub>, θ<sub>r|u</sub>, λ<sub>x</sub>, x<sub>r|a</sub>, θ<sub>yu</sub>, λ<sub>y</sub>, y<sub>r|a</sub>, θ<sub>g|u</sub>, λ<sub>g</sub>, g<sub>r|a</sub>.
* **Multiplication Nodes:** Represented by the symbol "*". These nodes take two inputs and perform multiplication.
* **Addition Nodes:** Represented by the symbol "+". These nodes take two inputs and perform addition.
* **Root Node:** The topmost node, also an addition node, representing the final result of the circuit.
* **Label:** "arithmetic circuit (AC)" located at the bottom center of the diagram.
### Detailed Analysis / Content Details
The diagram can be described as follows, starting from the bottom and moving upwards:
1. **Level 1 (Leaf Nodes):** The bottom row consists of 12 variables: θ<sub>xu</sub>, λ<sub>x</sub>, x<sub>r|a</sub>, θ<sub>r|u</sub>, λ<sub>x</sub>, x<sub>r|a</sub>, θ<sub>yu</sub>, λ<sub>y</sub>, y<sub>r|a</sub>, θ<sub>g|u</sub>, λ<sub>g</sub>, g<sub>r|a</sub>.
2. **Level 2:** Six multiplication nodes, each taking two leaf nodes as input.
* θ<sub>xu</sub> * λ<sub>x</sub>
* λ<sub>x</sub> * x<sub>r|a</sub>
* θ<sub>r|u</sub> * λ<sub>x</sub>
* λ<sub>x</sub> * x<sub>r|a</sub>
* θ<sub>yu</sub> * λ<sub>y</sub>
* λ<sub>y</sub> * y<sub>r|a</sub>
* θ<sub>g|u</sub> * λ<sub>g</sub>
* λ<sub>g</sub> * g<sub>r|a</sub>
3. **Level 3:** Two addition nodes, each taking two multiplication nodes as input.
* (θ<sub>xu</sub> * λ<sub>x</sub>) + (λ<sub>x</sub> * x<sub>r|a</sub>)
* (θ<sub>r|u</sub> * λ<sub>x</sub>) + (λ<sub>x</sub> * x<sub>r|a</sub>)
* (θ<sub>yu</sub> * λ<sub>y</sub>) + (λ<sub>y</sub> * y<sub>r|a</sub>)
* (θ<sub>g|u</sub> * λ<sub>g</sub>) + (λ<sub>g</sub> * g<sub>r|a</sub>)
4. **Level 4:** Two multiplication nodes, each taking two addition nodes as input.
* [(θ<sub>xu</sub> * λ<sub>x</sub>) + (λ<sub>x</sub> * x<sub>r|a</sub>)] * [(θ<sub>r|u</sub> * λ<sub>x</sub>) + (λ<sub>x</sub> * x<sub>r|a</sub>)]
* [(θ<sub>yu</sub> * λ<sub>y</sub>) + (λ<sub>y</sub> * y<sub>r|a</sub>)] * [(θ<sub>g|u</sub> * λ<sub>g</sub>) + (λ<sub>g</sub> * g<sub>r|a</sub>)]
5. **Level 5 (Root Node):** A final addition node taking the outputs of the two multiplication nodes as input.
* [(θ<sub>xu</sub> * λ<sub>x</sub>) + (λ<sub>x</sub> * x<sub>r|a</sub>)] * [(θ<sub>r|u</sub> * λ<sub>x</sub>) + (λ<sub>x</sub> * x<sub>r|a</sub>)] + [(θ<sub>yu</sub> * λ<sub>y</sub>) + (λ<sub>y</sub> * y<sub>r|a</sub>)] * [(θ<sub>g|u</sub> * λ<sub>g</sub>) + (λ<sub>g</sub> * g<sub>r|a</sub>)]
### Key Observations
The circuit is symmetrical in its structure, with two identical branches converging at the root. The use of multiplication and addition suggests that the circuit is designed to compute a product of sums or a similar arithmetic expression. The variables with subscripts "r|a" likely represent some form of conditional or masked values.
### Interpretation
This diagram represents a computational model for an arithmetic operation. The circuit's structure suggests it's performing a weighted sum of products. The variables θ, λ, x, y, and g likely represent parameters, weights, and input values, respectively. The "r|a" notation could indicate a conditional application of the input value, potentially representing a routing or activation mechanism. The overall purpose of the circuit is to compute a complex function based on these inputs and weights. The diagram is a visual representation of a mathematical expression, and understanding the meaning of the variables is crucial to interpreting the circuit's functionality. The circuit could be part of a larger system, such as a neural network or a signal processing algorithm.
</details>
values of endogenous variables. Variable X has indicators λ x and λ ¯ x and variable Y has indicators λ y and λ ¯ y . This circuit can compute the probability of any observational event η = x in time linear in the circuit size. We simply set each indicator to 1 if its subscript is compatible with the event η , otherwise we set it to 0 , and then evaluate the circuit [13]. For the event η = ¯ x , the indicators are set to λ x =0 , λ ¯ x =1 , λ y =1 , λ ¯ y =1 . If we (symbolically) evaluate the circuit under this indicator setting, we get θ u θ ¯ x | u ( θ y | u + θ ¯ y | u ) + θ ¯ u θ ¯ x | ¯ u ( θ y | ¯ u + θ ¯ y | ¯ u ) which is the expected result, Pr (¯ x ) . The circuit can also compute the probability of any interventional event η = y x in time linear in the circuit size. This is remarkably simple as well. To compute Pr ( y x ) , known as the causal effect, we first set the
parameters of all variables in X to 1 and then evaluate the circuit at instantiation x , y . 1 Suppose that x = ¯ x and y = ¯ y in our running example. To compute Pr (¯ y ¯ x ) , we evaluate the circuit while setting the parameters for variable X to θ x | u = θ ¯ x | u = θ x | ¯ u = θ ¯ x | ¯ u = 1 , and setting the indicators to λ x =0 , λ ¯ x =1 , λ y =0 , λ ¯ y =1 . If we (symbolically) evaluate the circuit under these settings, we get Pr (¯ y ¯ x ) = θ u θ ¯ y | u + θ ¯ u θ ¯ y | ¯ u = Pr (¯ y ) which is the expected result ( X has no causal effect on Y ).
Exploiting Unknown Mechanisms Consider the endogenous variable X in the causal graph we just discussed and its parameters θ x | u , θ ¯ x | u , θ x | ¯ u , θ ¯ x | ¯ u . Since these parameters specify a causal mechanism, they must all be in { 0 , 1 } subject to the constraints θ x | u + θ ¯ x | u = 1 and θ x | ¯ u + θ ¯ x | ¯ u = 1 . The main contribution of the new compilation algorithm in [15] is that it can exploit these constraintswithout needing to know the specific values of θ x | u , θ ¯ x | u , θ x | ¯ u , θ ¯ x | ¯ u -to produce circuits whose size can be exponentially smaller compared to methods developed during the last two decades; see, e.g., [13, 5, 6, 11, 32] and [14, Chapters 12,13]. These earlier methods can exploit functional dependencies computationally but only if they know the specific values of parameters. This does not help in a causality context (or a learning context more generally) where we do not know these values. The details of how the algorithm does this are discussed in Sections 4-6.
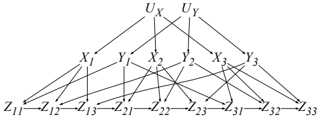
For now, consider the family of causal graphs on the right which generally has variables U X , U Y , X i , Y j , Z ij for i, j = 1 , . . . , n . These models have treewidth ≥ n +1 and hence are not accessible to non-parametric inference methods as they take time exponential in n . Moreover, the causal effect of X 1 on Z nn has the only back-door
<details>
<summary>Image 4 Details</summary>
### Visual Description
\n
## Diagram: Factor Graph
### Overview
The image depicts a factor graph, a bipartite graph used to represent the factorization of a function. The graph consists of variable nodes and factor nodes, connected by edges. This particular graph appears to represent a system with three variables (X and Y indexed by 1, 2, and 3) and several intermediate and output variables (Z indexed by two numbers).
### Components/Axes
The diagram consists of the following nodes:
* **Variable Nodes:** X<sub>1</sub>, Y<sub>1</sub>, X<sub>2</sub>, Y<sub>2</sub>, X<sub>3</sub>, Y<sub>3</sub>, Z<sub>11</sub>, Z<sub>12</sub>, Z<sub>13</sub>, Z<sub>21</sub>, Z<sub>22</sub>, Z<sub>23</sub>, Z<sub>31</sub>, Z<sub>32</sub>, Z<sub>33</sub>, U<sub>x</sub>, U<sub>y</sub>
* **Factor Nodes:** Implicitly represented by the connections between variable nodes.
The graph is structured in three layers:
1. Top Layer: U<sub>x</sub>, U<sub>y</sub>
2. Middle Layer: X<sub>1</sub>, Y<sub>1</sub>, X<sub>2</sub>, Y<sub>2</sub>, X<sub>3</sub>, Y<sub>3</sub>
3. Bottom Layer: Z<sub>11</sub>, Z<sub>12</sub>, Z<sub>13</sub>, Z<sub>21</sub>, Z<sub>22</sub>, Z<sub>23</sub>, Z<sub>31</sub>, Z<sub>32</sub>, Z<sub>33</sub>
Edges connect nodes between adjacent layers.
### Detailed Analysis / Content Details
The connections (factors) are as follows:
* U<sub>x</sub> is connected to X<sub>1</sub>, X<sub>2</sub>, X<sub>3</sub>.
* U<sub>y</sub> is connected to Y<sub>1</sub>, Y<sub>2</sub>, Y<sub>3</sub>.
* X<sub>1</sub> is connected to Z<sub>11</sub>, Z<sub>12</sub>, Z<sub>13</sub>.
* Y<sub>1</sub> is connected to Z<sub>11</sub>, Z<sub>21</sub>, Z<sub>31</sub>.
* X<sub>2</sub> is connected to Z<sub>12</sub>, Z<sub>22</sub>, Z<sub>32</sub>.
* Y<sub>2</sub> is connected to Z<sub>21</sub>, Z<sub>22</sub>, Z<sub>23</sub>.
* X<sub>3</sub> is connected to Z<sub>13</sub>, Z<sub>23</sub>, Z<sub>33</sub>.
* Y<sub>3</sub> is connected to Z<sub>31</sub>, Z<sub>32</sub>, Z<sub>33</sub>.
The graph is fully connected between layers, indicating that each factor depends on all variables in the adjacent layers.
### Key Observations
The structure suggests a system where U<sub>x</sub> and U<sub>y</sub> represent some input or prior information that influences the variables X and Y, respectively. The Z variables represent the combined effect of X and Y. The indexing of the Z variables (e.g., Z<sub>ij</sub>) suggests a relationship between the i-th X variable and the j-th Y variable.
### Interpretation
This factor graph likely represents a probabilistic model where the joint probability distribution can be factored into a product of functions (factors) involving pairs of variables. The U variables could represent prior distributions or constraints on the X and Y variables. The Z variables could represent observations or measurements that depend on the values of X and Y.
The graph's structure suggests that the variables X and Y are conditionally independent given the Z variables. The connections between U<sub>x</sub> and X<sub>i</sub>, and U<sub>y</sub> and Y<sub>i</sub>, indicate that the U variables influence the distributions of the X and Y variables.
The specific meaning of the variables and factors would depend on the context of the application. However, the factor graph provides a clear and concise representation of the relationships between the variables and factors in the model. This is a common representation in areas like Bayesian networks, Markov random fields, and graphical models.
</details>
X 2 , . . . , X n . By exploiting (unknown) causal mechanisms, we can now compile these causal graphs into circuits of size O ( n 2 ) (linear in the number of variables), allowing us to compute associational and interventional queries in O ( n 2 ) time. We will say more about this model and back-doors later.
Cross-Layer Inference We next show how to perform cross-layer causal inference using circuits. In a nutshell, we will use the circuit to estimate maximum-likelihood parameters for both exogenous and endogenous variables in the causal graph. We will then plug the estimates into the circuit and use it to compute associational and interventional probabilities in time linear in the circuit size as shown earlier. Suppose V is the set of observed endogenous variables. Since the causal graph has hidden variables, the maximum-likelihood parameters are not unique so they are not identifiable. However, the distribution Pr ( V ) is identifiable in this case. That is, even though we may have multiple maximum-likelihood estimates, they all lead to the same distribution Pr ( V ) . 2 This approach will need to be applied carefully though as its validity depends on (1) the specific query of interest, (2) the set of observed variables and (3) the causal graph structure. We will discuss this in detail after elaborating further on the estimation of maximum-likelihood parameters.
Estimation Since each example in a dataset corresponds to an observational event, one can use arithmetic circuits to compute the likelihood function by simply evaluating the circuit at each example (in linear time) and then multiplying the results. The algorithm in [15] facilitates this computation further as it compiles causal graphs into circuits in the form of tensor graphs. These are computation graphs in which nodes represent tensor operations instead of arithmetic operations, allowing one to evaluate and differentiate the circuit significantly more efficiently (think of a tensor operation as doing a bulk of arithmetic operations in parallel). Tensor graphs can lead to orders of magnitude speedups in estimation and inference time, especially that they allow batch (parallel) processing of examples-see [15, 9] which compiled circuits with tens of millions of nodes, leading to evaluation times in milliseconds. Backpropagation on arithmetic circuits takes time linear in the circuit size. Moreover, the partial derivatives with respect to circuit parameters correspond to marginals over families in the causal graph (nodes and their parents) [13], which is all that one needs to compute parameter updates for the EM algorithm; see, for example, [14, Eq. 17.7]. Hence, one can use methods such as gradient descent and EM to seek maximum-likelihood estimates, but the efficacy of these methods needs further investigation under the stated conditions including finite data.
1 This method follows directly from the mutilation semantics of interventions [26, Section 1.3.1] and the polynomial semantics of arithmetic circuits [13, 10]. It is a slight variation on [31] which also emulates interventions by adjusting model parameters; see also [37].
2 Ying Nian Wu provided the following argument for infinite data. Let Pr D ( V ) be the data distribution and Pr θ ( U , V ) be the model so Pr θ ( V ) = ∑ U Pr θ ( U , V ) . Maximum-likelihood estimation is equivalent to minimizing the KL divergence KL ( Pr D ( V ) | Pr θ ( V )) . If θ is not identifiable, then all solutions of θ belong to an equivalence class that minimizes the KL divergence and they all give the same marginal Pr θ ( V ) .
Identifiability A central question in causal inference is whether interventional probabilities can be identified based on a causal graph and (infinite) observational data on the endogenous variables V . Intuitively, identifiability means that interventional probabilities can be computed using any parameterization of the causal graph (or circuit) that yields the true marginal distribution Pr ( V ) ; see [26, Def 3.2.4] for a formal definition. A well behaved case arises when each exogenous variable feeds into at most one causal mechanism. These models are said to be Markovian and the case is termed no unobserved confounders. Interventional probabilities are always identifiable for Markovian models, so we can always compute the causal effect Pr ( y x ) for these models using circuits parameterized by maximum-likelihood parameters. 3
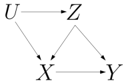
If an exogenous variable feeds into more than one causal mechanism, the model is said to be semi-Markovian . In this case, interventional probabilities are identifiable only when certain conditions are met. The causal graph on the right corresponds to a semi-Markovian model since U feeds into the causal mechanisms for both X and Z . The causal effect Pr ( y x ) is identifiable since Pr ( y x ) = ∑ z Pr ( y | x, z ) Pr ( z ) .
<details>
<summary>Image 5 Details</summary>
### Visual Description
\n
## Diagram: Directed Graph
### Overview
The image depicts a directed graph with six nodes labeled U, X, Y, Z, and two intermediate nodes. The graph illustrates relationships between these nodes through directed edges (arrows).
### Components/Axes
The diagram consists of nodes and directed edges. There are no explicit axes or scales. The nodes are labeled with single capital letters. The edges indicate a directional relationship from the source node to the destination node.
### Detailed Analysis or Content Details
The diagram shows the following relationships:
* U -> Z (U points to Z)
* U -> X (U points to X)
* Z -> Y (Z points to Y)
* X -> Y (X points to Y)
* X -> Z (X points to Z)
The graph forms a triangular structure with U at the top, and X and Z as intermediate nodes, both pointing to Y.
### Key Observations
The graph is acyclic, meaning there are no closed loops. Node Y has two incoming edges (from Z and X), making it a sink node. Node U has two outgoing edges (to Z and X), making it a source node. Node X has three outgoing edges (to Y, Z, and Z).
### Interpretation
This diagram likely represents a dependency graph, a workflow, or a causal relationship between the labeled entities. For example:
* U could represent an initial condition or input.
* X and Z could represent intermediate steps or processes.
* Y could represent a final outcome or output.
The multiple paths from U to Y (U -> Z -> Y and U -> X -> Y) suggest that there are multiple ways to achieve the outcome Y. The edge from X to Z indicates a feedback or iterative relationship between these two nodes. The diagram does not provide quantitative data, but rather a qualitative representation of relationships. It is a structural representation of connections, not a measurement of values.
</details>
However, the causal effect Pr ( x z ) is not identifiable as it cannot be uniquely determined based on the causal graph and the distribution Pr ( X,Y,Z ) . The do-calculus provides a complete and efficient characterization of identifiable, interventional probabilities based on observational data [25, 35, 34]; see also [20]. 4 It is based on a set of rules that can be used to transform an interventional probability into a formula that includes only associational probabilities (we will call this an identifiability formula ). If the rules fail to make such a derivation, then the interventional probability is not identifiable. Other methods such as back-door [24] and front-door [29] provide simpler but incomplete tests and lead to simple identifiability formulas. For example, the back-door and front-door formulas have the forms Pr ( y x ) = ∑ z Pr ( y | x , z ) Pr ( z ) and Pr ( y x ) = ∑ z Pr ( z | x ) ∑ x ′ Pr ( y | x ′ , z ) Pr ( x ′ ) , where Z is called a back-door or front-door, respectively. A more refined identifiability test that utilizes contextspecific independence relations was proposed recently [36], thus expanding the reach of cross-layer causal inference. These relations correspond to equating certain parameters and can be integrated into circuits [5] to reduce the number of estimands. In summary, we can compute causal effects on semi-Markovian models using circuits parameterized by maximum-likelihood estimates, but only for identifiable queries as licensed by the do-calculus or a more refined identifiability procedure.
Identifiability Formulas vs Circuits Most identifiability procedures yield formulas (also called the effect estimand ) which play three roles: they provide a proof of identifiability; they point to endogenous variables whose measurement guarantees identifiability; and they allow one to evaluate the causal effect by estimating quantities that populate such formulas. Back-door and front-door formulas have simple forms (albeit exponential sums) but the do-calculus and the procedure in [36] may generate identifiability formulas that are much more complex. Some procedures do not even aim to produce identifiability formulas; e.g., [19]. To evaluate causal effects using circuits, one only needs an identifiability test not a formula. That is, one estimates circuit parameters only once and then uses the parametrized circuit to answer any identifiable query in time linear in the circuit size-regardless of how complex the identifiability formula may be and without needing to have access to one. 5 Additional knowledge such as context-specific independence and known mechanisms can be directly integrated into the circuit [5], which can only improve the quality of estimates under finite data. At its core, this use of circuits amounts to computing causal effects based on the classical method of mutilating causal graphs, armed by an observation and an advance. The observation is that using maximum-likelihood parameters is sound if the causal effect is identifiable. The advance is that we can now perform this computation much more efficiently due to exploiting unknown mechanisms.
The Circuit Compilation Process Wewill next discuss the circuit compilation algorithm introduced recently in [15]. We will focus on the key insights behind the algorithm and slightly adjust it to suit our current objectives (the original algorithm targetted specific queries as is typically demanded in a supervised learning setting). In a nutshell, the algorithm is based on the classical algorithm of variable elimination (VE) with two exceptions. First, we will use VE symbolically by working with symbolic parameters instead of numeric ones. Second, we will empower VE by two new theorems based on unknown causal mechanisms which can reduce its complexity exponentially. We will review VE in Section 4 and then present the new theorems and compilation algorithm in Sections 5 and 6.
3 See [34, Corollary 4] for a weaker, necessary condition that guarantees identifiability of all causal effects.
4 See also [22] for a treatment of identifiability based on both observational and interventional data and [3] for a survey that considers other tasks such as the evaluation of soft interventions.
5 Using circuits in this manner requires fixing the cardinality of variables including exogenous ones.
## 4 The Variable Elimination Algorithm ( VE )
VE operates on causal graphs which are parameterized by factors. A factor over variables X is a function f ( X ) that maps each instantiation x into a number f ( x ) . For each node X and its parents P in the causal graph, we need a factor f X ( X, P ) where f X ( x, p ) = Pr ( x | p ) . For example, factor f ( U ) on the right specifies the distribution Pr ( U ) for exogenous variable U and factor g ( XY ) specifies the mechanism for endogenous variable Y : x 0 ↦→ y 1 , x 1 ↦→ y 0 . VE is based
| U u | X Y x 0 y 0 |
|-------|---------------|
two factor operations: multiplication and sum-out. The product of factors f ( X ) and g ( Y ) is another factor h ( Z ) , where Z = X ∪ Y and h ( z ) = f ( x ) g ( y ) for the unique instantiations x and y that are compatible with instantiation z . Summing-out variables Y ⊆ X from factor f ( X ) yields another factor g ( Z ) , where Z = X \ Y and g ( z ) = ∑ y f ( yz ) . We use ∑ Y f to denote the resulting factor g .
The joint distribution of a parametrized causal graph is simply the product of its factors. The causal graph in Figure 1(b) has factors f A ( A ) , f B ( AB ) , f C ( AC ) , f D ( BCD ) and f E ( CE ) . Its joint distribution is Pr ( ABCDE ) = f A f B f C f D f E . To record an observation X = x , we use an auxiliary evidence factor λ X ( X ) with λ X ( x ) = 1 and λ X ( x ′ ) = 0 for x ′ = x . A posterior distribution is obtained by normalizing the product of all factors in the causal graph including evidence factors. Suppose we have evidence e on variables A and E in Figure 1(b). The posterior Pr ( D | e ) is obtained by evaluating then normalizing the expression ∑ ABCE λ A λ E f A f B f C f D f E . VE tries to evaluate such expressions efficiently [38, 17] based on two theorems; see, e.g., [14, Chapter 6].
The first theorem allows us to sum out variables in any order. The second theorem allows us to pull out factors from sums, which can lead to exponential savings in time and space.
<!-- formula-not-decoded -->
Theorem 2. If variables X appear in factor f but not in factor g , then ∑ X f · g = g ∑ X f .
Consider the expression ∑ ABDE f ( ACE ) f ( BCD ) . A direct evaluation multiplies the two factors to yield f ( ABCDE ) then sums out variables ABDE . Using Theorem 1, we can arrange the expression into ∑ AE ∑ BD f ( ACE ) f ( BCD ) . Using Theorem 2, we can arrange it further into ∑ AE f ( ACE ) ∑ BD f ( BCD ) which is more efficient to evaluate. If we eliminate all variables using order π , and if the largest factor constructed in the process has w +1 variables, then w is called the width of order π . The smallest width attained by any elimination order corresponds to the treewidth of the causal graph. The best time complexity that can be attained by VE is O ( n exp( w )) , where n is the number of variables and w is the causal graph treewidth. This holds for any other non-parametric method known today (i.e., inference methods that do not exploit the graph parameters).
## 5 Variable Elimination with Causal Mechanisms
We next present two recent results that allow us to simplify expressions beyond what is permitted by Theorems 1 and 2, leading to a tighter complexity based on what we shall call the causal treewidth. We will use F , G , H to denote sets of factors, where each set is interpreted as a product of its factors.
Definition 5. A factor f ( X, P ) is said to be a 'mechanism' for variable X iff all numbers in the factor are in { 0 , 1 } and ∑ x f ( x, p ) = 1 for every instantiation p .
Theorem 3 ([15]) . Let f be a mechanism for variable X . If f ∈ G and f ∈ H , then G·H = G ∑ X H .
According to this result, if a mechanism for X appears in both parts of a product, then variable X can be summed out from one part without changing the value of the product. This has a key corollary.
Corollary 1. Let f be a mechanism for X . If f ∈ G and f ∈ H , then ∑ X G·H = ( ∑ X G ) ( ∑ X H ) .
That is, if a mechanism for X appears in both parts of a product, we can sum out variable X from the product by independently summing it out from each part. This is a remarkable addition to the algorithm of variable elimination which has been under study for a few decades now. Corollary 1 may appear unusable as it is predicated on multiple occurrences of a mechanism whereas the factors of a causal graph contain a single mechanism for each endogenous variable. This is where the second result comes in: replicating mechanisms in a product does not change the product value.
Theorem 4 ([15]) . For mechanism f , if f ∈ G , then f · G = G .
For an example that uses these theorems, consider the expression α = ∑ X f ( XY ) g ( XZ ) h ( XW ) . VE has to multiply all three factors before summing out variable X , leading to a factor over four variables XYZW . However, if factor f is a mechanism for variable X , then we can replicate it by Theorem 4: α = f ( XY ) g ( XZ ) f ( XY ) h ( XW ) . Corollary 1 then gives α = ∑ X f ( XY ) g ( XZ ) ∑ X f ( XY ) h ( XW ) . Hence, we can now evaluate expression α without having to construct any factor over more than three variables. This technique can more generally lead to exponential savings since the size of a factor is exponential in the number of its variables.
We will refer to the extension of VE with Theorems 3 and 4 as VEC ( V ariable E limination for C ausality). We emphasize that these new theorems do not require the values of parameters (i.e., specific mechanisms). They only need to know if a variables is functionally determined by its parents.
## 6 Compiling Causal Graphs Into Circuits
We next show how VE/VEC can be used symbolically to compile non-parametric causal graphs into arithmetic circuits with symbolic parameters. We will first show this concretely on a small example using VE, then discuss a general compilation algorithm based on VE and finally based on VEC.
Consider the causal graph U → V with binary variables. The parameters of this graph are given by the factors f ( U ) and g ( UV ) shown on the right. We also added an evidence factor for endogenous variable V as it will be measured. These factors have symbolic parameters instead of numeric ones.
| U | f ( U ) | U u | V g ( UV v θ v | u | V | h ( V ) |
|-----|-----------|-------|---------------------------|-----|-----------|
| u | θ u | u | ¯ v θ ¯ v | u | v | λ v |
| ¯ u | θ ¯ u | ¯ u | v θ v | ¯ u | ¯ v | λ ¯ v |
| | | ¯ u | ¯ v θ ¯ v | ¯ u | | |
We will further overload the + and ∗ operators so they now construct circuit nodes instead of performing numeric operations. That is, each entry in the above factors can be viewed as a leaf circuit node. When multiplying, say, node θ u with node θ v | u , we construct a circuit node with ∗ as its label and nodes θ u , θ v | u as its children. And similarly for addition. We can now get a circuit for the causal graph by multiplying all its factors, including evidence factors, and then summing out all variables, AC = ∑ UV f ( U ) g ( UV ) h ( V ) . The resulting factor AC will have a single entry which contains the root of compiled circuit λ v ∗ θ u ∗ θ v | u + λ ¯ v ∗ θ u ∗ θ ¯ v | u + λ v ∗ θ ¯ u ∗ θ v | ¯ u + λ ¯ v ∗ θ ¯ u ∗ θ ¯ v | ¯ u . The equivalent expression AC = ∑ V h ( V ) ∑ U f ( U ) g ( UV ) gives the circuit λ v ∗ ( θ u ∗ θ v | u + θ ¯ u ∗ θ v | ¯ u ) + λ ¯ v ∗ ( θ u ∗ θ ¯ v | u + θ ¯ u ∗ θ ¯ v | ¯ u ) . Hence, the size and shape of a compiled circuit depend on how we schedule factor operations (multiplication and sum-out). The symbolic use of VE to compile circuits was initially proposed in [6]. This method was recently refined in [15] by (1) scheduling factor operations based on a specific class of binary jointrees [33] and (2) allowing one to compile circuits using VEC by thinning the jointree. We will explain this advance after a brief review of (binary) jointrees.
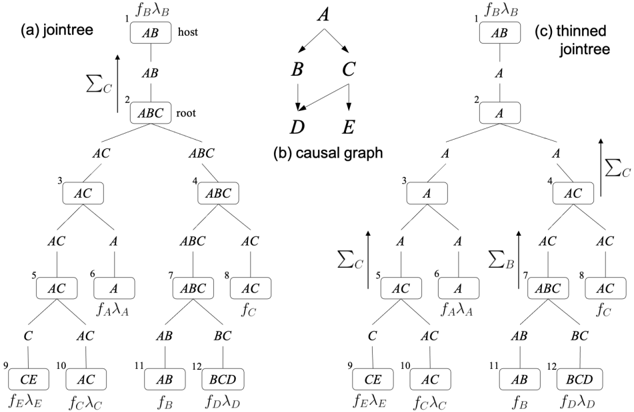
Jointrees A binary jointree is a tree in which each node is either a leaf (has a single neighbor) or internal (has three neighbors). We will require the leaf nodes to be in one-to-one correspondence with the factors of a causal graph, including replicated factors but excluding evidence factors. As we show next, the topology of a binary jointree determines all its properties, including the set of variables attached to each node, called a cluster, and the set of variables attached to each edge, called a separator. Figure 1(a) depicts a binary jointree for the causal graph in Figure 1(b). Following the convention in [15], this jointree is layed out so that each internal node has two neighbors below it (children) and the third neighbor above it (parent). The leaf nodes of this jointree are numbered 1 , 6 , 8 , 9 , 10 , 11 , 12 and correspond to the causal graph factors, f B , f A , f C , f E , f C , f B , f D (we replicated the factors for B and C ). The separator for edge ( i, j ) between node i and its parent j is denoted sep ( i ) and contains variables that are shared between factors on both sides of the edge ( i, j ) . For example, sep (3) = { A,C } as these are the variables shared between factors at leaves { 6 , 9 , 10 } and factors at leaves { 1 , 8 , 11 , 12 } . The cluster of node i is denoted cls ( i ) . The cluster of a leaf node is the variables of its associated factor. The cluster of an internal node is the union of separators connected to its children. For example, for leaf node 9 with factor f E , cls (9) = vars ( f E ) = { C, E } . Moreover, for internal node 7 with children 11 and 12 , cls (7) = sep (11) ∪ sep (12) = { A,B,C } .
Scheduling Given a binary jointree, VE (and later VEC) schedules its operations as follows. Visiting nodes bottom-up in the jointree, each node i computes a factor f ( i ) and sends it to its parent. A leaf node i computes f ( i ) by projecting its associated factor on sep ( i ) . For example, f (9) = ∑ E f E λ E . An internal node i computes f ( i ) by multiplying the factors it receives from its children and then projecting the product on sep ( i ) . For example, f (2) = ∑ C f (3) f (4) . This process terminates at the
Figure 1: A causal graph (middle) with a jointree (left) and a thinned jointree (right). The mechanisms for variables B and C , f B and f C , are replicated twice in the jointrees.
<details>
<summary>Image 6 Details</summary>
### Visual Description
\n
## Diagram: Jointree, Causal Graph, and Thinned Jointree
### Overview
The image presents three diagrams: a jointree (a), a causal graph (b), and a thinned jointree (c). All three diagrams represent relationships between variables A, B, C, D, and E. The diagrams use nodes (rectangles) to represent combinations of variables and arrows to indicate dependencies or conditional probabilities. Each node is labeled with a combination of variables and a number (1-12) indicating a specific step or state in the process. Each arrow is labeled with a function representing a conditional probability.
### Components/Axes
The diagrams share the following components:
* **Nodes:** Rectangular boxes representing combinations of variables (e.g., AB, AC, ABC).
* **Arrows:** Indicate dependencies or conditional probabilities between nodes.
* **Labels on Arrows:** Functions like f<sub>B∧B</sub>, f<sub>A</sub>, f<sub>C</sub>, f<sub>D</sub>, f<sub>E</sub>, etc., representing conditional probabilities.
* **Variables:** A, B, C, D, and E.
* **Σ:** Symbol indicating summation or marginalization.
* **Labels:** "host", "root".
### Detailed Analysis or Content Details
**Diagram (a): Jointree**
* **Node 1:** `AB` labeled "host", with an incoming arrow from `f<sub>B∧B</sub>`.
* **Node 2:** `ABC` labeled "root", with an incoming arrow from `Σ`.
* **Node 3:** `AC`, with an incoming arrow from `ABC`.
* **Node 4:** `ABC`, with an incoming arrow from `AC`.
* **Node 5:** `AC`, with an incoming arrow from `ABC`.
* **Node 6:** `A`, with an incoming arrow from `f<sub>A∧A</sub>`.
* **Node 7:** `ABC`, with an incoming arrow from `AC`.
* **Node 8:** `AC`, with an incoming arrow from `f<sub>C</sub>`.
* **Node 9:** `CE`, with an incoming arrow from `f<sub>E∧E</sub>`.
* **Node 10:** `AC`, with an incoming arrow from `f<sub>C∧C</sub>`.
* **Node 11:** `AB`, with an incoming arrow from `f<sub>B</sub>`.
* **Node 12:** `BCD`, with an incoming arrow from `f<sub>D∧D</sub>`.
* An arrow from `Σ` points to `ABC` (Node 2).
**Diagram (b): Causal Graph**
* **Node A:** At the top, with arrows pointing to B, C, D, and E.
* **Node B:** Receives an arrow from A.
* **Node C:** Receives an arrow from A.
* **Node D:** Receives an arrow from A.
* **Node E:** Receives an arrow from A.
* **Node 5:** `C`, with an incoming arrow from `Σ`.
* **Node 6:** `A`, with an incoming arrow from `f<sub>A∧A</sub>`.
* **Node 7:** `ABC`, with an incoming arrow from `AB`.
* **Node 8:** `AC`, with an incoming arrow from `f<sub>C</sub>`.
* **Node 9:** `CE`, with an incoming arrow from `f<sub>E∧E</sub>`.
* **Node 10:** `AC`, with an incoming arrow from `f<sub>C∧C</sub>`.
* **Node 11:** `AB`, with an incoming arrow from `f<sub>B</sub>`.
* **Node 12:** `BCD`, with an incoming arrow from `f<sub>D∧D</sub>`.
* An arrow from `Σ` points to `C` (Node 5).
**Diagram (c): Thinned Jointree**
* **Node 1:** `AB`, with an incoming arrow from `f<sub>B∧B</sub>`.
* **Node 2:** `A`, with an incoming arrow from `Σ`.
* **Node 3:** `A`, with an incoming arrow from `A`.
* **Node 4:** `AC`, with an incoming arrow from `AC`.
* **Node 5:** `Σ`, points to `A` (Node 2).
* **Node 6:** `A`, with an incoming arrow from `f<sub>A∧A</sub>`.
* **Node 7:** `ABC`, with an incoming arrow from `AC`.
* **Node 8:** `AC`, with an incoming arrow from `f<sub>C</sub>`.
* **Node 9:** `CE`, with an incoming arrow from `f<sub>E∧E</sub>`.
* **Node 10:** `AC`, with an incoming arrow from `f<sub>C∧C</sub>`.
* **Node 11:** `AB`, with an incoming arrow from `f<sub>B</sub>`.
* **Node 12:** `BCD`, with an incoming arrow from `f<sub>D∧D</sub>`.
* An arrow from `Σ` points to `A` (Node 2).
### Key Observations
* All three diagrams represent the same set of variables (A, B, C, D, E).
* The causal graph (b) shows a clear hierarchical structure with A as the root cause influencing B, C, D, and E.
* The jointree (a) and thinned jointree (c) represent the same relationships but with different levels of detail and complexity. The thinned jointree (c) appears to be a simplified version of the jointree (a).
* The functions labeled on the arrows (e.g., f<sub>B∧B</sub>, f<sub>A</sub>) likely represent conditional probabilities.
* The summation symbol (Σ) indicates marginalization or summing over possible values of other variables.
### Interpretation
These diagrams likely illustrate a process of probabilistic inference or belief propagation in a Bayesian network. The causal graph (b) represents the underlying causal relationships between the variables. The jointree (a) is a graphical representation of the factorization of the joint probability distribution over all variables. The thinned jointree (c) is a simplification of the jointree, potentially used for more efficient computation.
The functions on the arrows represent conditional probabilities, and the summation symbol indicates the process of marginalizing out variables to obtain the probability of a particular event. The diagrams demonstrate how to represent and manipulate probabilistic relationships between variables in a graphical model. The progression from causal graph to jointree to thinned jointree suggests a process of model construction and simplification for computational efficiency. The labels "host" and "root" likely refer to specific roles or states within the network. The numbers (1-12) likely represent a sequence of steps in an algorithm or process.
</details>
top leaf node r which multiplies the factor it receives from its single child c with its own factor and then projects the product on the empty set. In Figure 1(a), the final computation is ∑ AB f B λ B f (2) . Denoting the factor at leaf node i by F i , this process yields the factor AC = ∑ cls ( r ) F r f ( c ) , where
<!-- formula-not-decoded -->
The final factor AC has a single entry which contains the root of the compiled circuit as shown earlier. Moreover, the size of this circuit is determined by the jointree clusters and separators. Each cluster/separator contributes a number of multiplication/addition nodes that is exponential in the cluster/separator size. In a jointree, the largest cluster dominates the largest separator and the size of the largest cluster minus 1 is called the jointree width. Furthermore, the smallest width attained by any jointree corresponds to the treewidth of the causal graph; see, [14, Chapter 9]. Hence, the complexity of this compilation method is exponential in the treewidth of the causal graph.
Thinning This complexity was recently significantly improved by exploiting (unknown) causal mechanisms [15]. The basic idea is to thin the jointree by shrinking its separators (and hence clusters) while maintaining the correctness of compiled circuit. The thinning process is based on Theorems 3 and 4 and can lead to an exponential reduction in the circuit size. To see the key insight behind this thinning process, consider node 2 in the jointree of Figure 1(a). The separators sep (3) and sep (4) of its children both contain variable C . Hence, the factors f (3) and f (4) sent by these children to node 2 both contain variable C . Since C does not appear in sep (2) it gets summed out at node 2 so it does not appear in the factor f (2) that this node sends to its parent. However, since we have two replicas of the mechanism for variable C at leaf nodes 8 and 10 (as licensed by Theorem 4), we can sum out C earlier, at nodes 4 and 5 (as licensed by Theorem 3). This means that C can be removed from sep (4) , sep (5) and also sep (3) . We can similarly sum out variable B at node 7 , which removes it from sep (7) , sep (4) and sep (2) . The shrinking of separators causes clusters to shrink as well, leading to the thinned jointree in Figure 1(c) and a corresponding smaller circuit compilation.
Causal Treewidth The attained reduction in complexity depends on (1) the number of replicas for each mechanism; (2) the used binary jointree; and (3) how the jointree is thinned. Corresponding heuristics were proposed in [15] and the resulting algorithm was shown to yield exponential reductions in the size of compiled circuits on a number of benchmarks (elimination orders and jointrees are also constructed using heuristics since finding optimal ones is NP-hard). This motivates a new measure of complexity which we call the causal treewidth. We define this as the smallest width attained by any thinned jointree for a given causal graph. We next complement the empirical findings in [15] by showing that causal treewidth dominates treewidth and can be bounded when the treewidth is not.
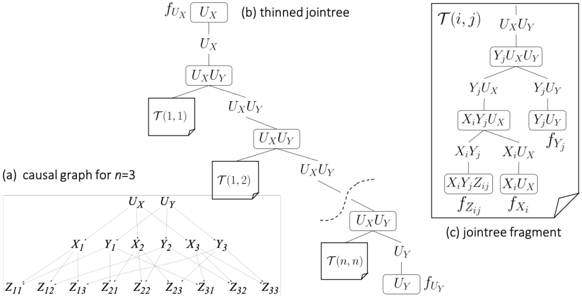
Figure 2: A causal graph and its thinned jointree. Mechanisms f X i and f Y j are replicated n times.
<details>
<summary>Image 7 Details</summary>
### Visual Description
\n
## Diagram: Causal Graph and Jointree Representations
### Overview
The image presents a diagram illustrating the relationship between a causal graph and its corresponding jointree representation. It shows a causal graph for n=3, a thinned jointree derived from it, and a fragment of the jointree. The diagram uses nodes and edges to represent variables and their dependencies.
### Components/Axes
The diagram is divided into three main sections:
**(a) causal graph for n=3:** This section displays a graph with nodes labeled U<sub>x</sub>, U<sub>y</sub>, X<sub>i</sub>, Y<sub>i</sub>, Z<sub>ij</sub>, and T(1,1), T(1,2), T(n,n). Edges connect these nodes, representing causal relationships.
**(b) thinned jointree:** This section shows a tree-like structure with nodes labeled f<sub>U<sub>x</sub></sub>, U<sub>x</sub>, U<sub>x</sub>U<sub>y</sub>, T(1,1), T(1,2), U<sub>x</sub>U<sub>y</sub>, U<sub>y</sub>, and f<sub>U<sub>y</sub></sub>. The tree structure represents the factorization of the joint distribution.
**(c) jointree fragment:** This section displays a portion of the jointree, with nodes labeled T(i,j), U<sub>x</sub>U<sub>y</sub>, Y<sub>j</sub>U<sub>x</sub>, Y<sub>j</sub>U<sub>y</sub>, X<sub>i</sub>Y<sub>j</sub>U<sub>x</sub>, Y<sub>j</sub>U<sub>y</sub>, X<sub>i</sub>Y<sub>j</sub>Z<sub>ij</sub>, X<sub>i</sub>U<sub>x</sub>, f<sub>Z<sub>ij</sub></sub>, f<sub>X<sub>i</sub></sub>, f<sub>Y<sub>j</sub></sub>.
### Detailed Analysis or Content Details
**(a) Causal Graph for n=3:**
* Nodes: U<sub>x</sub>, U<sub>y</sub>, X<sub>1</sub>, X<sub>2</sub>, X<sub>3</sub>, Y<sub>1</sub>, Y<sub>2</sub>, Y<sub>3</sub>, Z<sub>11</sub>, Z<sub>12</sub>, Z<sub>13</sub>, Z<sub>21</sub>, Z<sub>22</sub>, Z<sub>23</sub>, Z<sub>31</sub>, Z<sub>32</sub>, Z<sub>33</sub>, T(1,1), T(1,2), T(n,n).
* Edges: U<sub>x</sub> is connected to X<sub>1</sub>, X<sub>2</sub>, X<sub>3</sub>. U<sub>y</sub> is connected to Y<sub>1</sub>, Y<sub>2</sub>, Y<sub>3</sub>. X<sub>i</sub> is connected to Z<sub>ij</sub> and Y<sub>j</sub>. Y<sub>j</sub> is connected to Z<sub>ij</sub>. T(1,1) is connected to U<sub>x</sub>U<sub>y</sub>. T(1,2) is connected to U<sub>x</sub>U<sub>y</sub>. T(n,n) is connected to U<sub>y</sub>.
**(b) Thinned Jointree:**
* Root Node: f<sub>U<sub>x</sub></sub>
* Branches:
* f<sub>U<sub>x</sub></sub> -> U<sub>x</sub>
* U<sub>x</sub> -> U<sub>x</sub>U<sub>y</sub>
* U<sub>x</sub>U<sub>y</sub> -> T(1,1)
* U<sub>x</sub>U<sub>y</sub> -> T(1,2)
* U<sub>x</sub>U<sub>y</sub> -> U<sub>x</sub>U<sub>y</sub>
* U<sub>x</sub>U<sub>y</sub> -> U<sub>y</sub>
* U<sub>y</sub> -> f<sub>U<sub>y</sub></sub>
**(c) Jointree Fragment:**
* Root Node: T(i,j)
* Branches:
* T(i,j) -> U<sub>x</sub>U<sub>y</sub>
* U<sub>x</sub>U<sub>y</sub> -> Y<sub>j</sub>U<sub>x</sub>
* U<sub>x</sub>U<sub>y</sub> -> Y<sub>j</sub>U<sub>y</sub>
* Y<sub>j</sub>U<sub>x</sub> -> X<sub>i</sub>Y<sub>j</sub>U<sub>x</sub>
* Y<sub>j</sub>U<sub>y</sub> -> X<sub>i</sub>Y<sub>j</sub>Z<sub>ij</sub>
* X<sub>i</sub>Y<sub>j</sub>U<sub>x</sub> -> X<sub>i</sub>U<sub>x</sub>
* X<sub>i</sub>Y<sub>j</sub>Z<sub>ij</sub> -> f<sub>Z<sub>ij</sub></sub>
* X<sub>i</sub>U<sub>x</sub> -> f<sub>X<sub>i</sub></sub>
* Y<sub>j</sub>U<sub>y</sub> -> f<sub>Y<sub>j</sub></sub>
### Key Observations
* The causal graph represents the dependencies between variables in a probabilistic model.
* The jointree is a tree-like structure that represents the factorization of the joint probability distribution.
* The thinned jointree is a simplified version of the jointree, obtained by eliminating redundant nodes.
* The jointree fragment shows a portion of the complete jointree, illustrating the relationships between specific variables.
* Nodes labeled 'f' likely represent factors in the factorization.
### Interpretation
The diagram illustrates the process of converting a causal graph into a jointree representation, which is a key step in performing inference in probabilistic graphical models. The jointree provides a compact and efficient way to represent the joint probability distribution, allowing for efficient computation of marginal and conditional probabilities. The thinned jointree further simplifies the representation by eliminating redundant nodes, reducing the computational cost of inference. The fragment shows how the jointree can be broken down into smaller components, making it easier to understand and analyze. The diagram demonstrates a fundamental concept in Bayesian networks and graphical models, showing how complex dependencies can be represented and manipulated using tree structures. The use of 'f' nodes suggests a factorization approach, where the joint distribution is expressed as a product of factors associated with the cliques in the jointree.
</details>
Theorem 5. The causal treewidth is no greater than treewidth. Moreover, there is a family of causal graphs with n 2 +2 n +1 variables, treewidth n +1 and causal treewidth 2 where n is an integer ≥ 1 .
Proof Sketch. Consider a causal graph G with treewidth w . Without replicating mechanisms, we can always get a (thinned) jointree with width w . Hence, the causal treewidth of G is ≤ w . To show the second part of the theorem, consider the family of causal graphs G n with exogenous variables U X , U Y and endogenous variables X i , Y j , Z ij for i, j = 1 , . . . , n ( 2 + 2 n + n 2 variables), and edges U X → X i , U Y → Y j , X i → Z ij , Y j → Z ij . Figure 2(a) depicts G 3 . We next show that G n has treewidth n +1 based on standard techniques for treewidth; see, e.g., [14, Chapter 9]. The moral graph of G n is obtained by dropping edge directions and connecting every pair of nodes X i and Y j by an undirected edge. Each Z ij has only two (connected) neighbors in the moral graph ( X i and Y j ) so it is a simplicial node. Hence, there must exist an optimal elimination order that starts with nodes Z ij [14, Section 9.3.2]. After eliminating all Z ij , nodes U X and U Y will each have n neighbors, and nodes X i and Y j will each have n +1 neighbors. A simple argument shows that eliminating these variables in any order from the moral graph will create a clique over n +1 variables so the treewidth is ≥ n +1 . One can easily verify that the elimination order Z 11 , . . . , Z nn , U X , Y 1 , . . . , Y n , U Y , X 1 , . . . , X n has width n +1 so the treewidth of G n is n +1 . Figure 2(b) depicts a thinned jointree for G n with width 2 , which results from cascading n 2 instances of the jointree fragment T ( i, j ) in Figure 2(c). This fragment contains the mechanism for Z ij and replicas of the mechanisms for X i and Y j . This thinned jointree is optimal since the mechanism for Z ij contains 3 variables so any thinned jointree must have a cluster of size ≥ 3 . Hence, the causal treewidth of G n is 2 .
Theorem 5 effectively says that circuits compiled by VEC are no larger than those compiled by VE and can be exponentially smaller. We finally note that a variation G ′ n on G n was shown in Section 3 with additional edges between variables Z ij . The treewidth of G ′ n must be ≥ n +1 yet has a thinned jointree of width 4 (constructed by the algorithm in [15]) so its causal treewidth is ≤ 4 . Recall that for this family of causal graphs G ′ n , the causal effect of X 1 on Z nn has the only back-door X 2 , . . . , X n so it has a back-door formula with a sum that is exponential in n and a circuit of size O ( n 2 ) .
## 7 Conclusion
We discussed recent techniques that can exploit causal mechanisms computationally without having to know their identities, which is the classical setup in causal inference. We also showed how one can use these techniques to compile non-parametric causal graphs into circuits that can be used to estimate parameters from data and to perform cross-layer causal inference in time linear in the circuit size. Our aim was to provide an intuitive exposure to these techniques to a causality audience who may not be as familiar with them, with the hope that this may lead to a synthesis on how tractable arithmetic circuits can aid causal inference in reaching higher levels of scalability and versatility.
Acknowledgements I wish to thank Elias Bareinboim, Yizuo Chen, Scott Mueller, Judea Pearl and Jin Tian for useful discussions and feedback. This work has been partially supported by NSF grant #ISS-1910317 and ONR grant #N00014-18-1-2561.
## References
- [1] Durgesh Agrawal, Yash Pote, and Kuldeep S. Meel. Partition function estimation: A quantitative study. In IJCAI , pages 4276-4285. ijcai.org, 2021.
- [2] E. Bareinboim, Juan David Correa, D. Ibeling, and Thomas F. Icard. On Pearl's hierarchy and the foundations of causal inference. 2021. Technical Report, R-60, Colombia University.
- [3] Elias Bareinboim and Judea Pearl. Causal inference and the data-fusion problem. Proc. Natl. Acad. Sci. USA , 113(27):7345-7352, 2016.
- [4] Craig Boutilier, Nir Friedman, Moisés Goldszmidt, and Daphne Koller. Context-specific independence in Bayesian networks. CoRR , abs/1302.3562, 2013.
- [5] Mark Chavira and Adnan Darwiche. Compiling Bayesian networks with local structure. In Leslie Pack Kaelbling and Alessandro Saffiotti, editors, IJCAI-05, Proceedings of the Nineteenth International Joint Conference on Artificial Intelligence, Edinburgh, Scotland, UK, July 30 August 5, 2005 , pages 1306-1312. Professional Book Center, 2005.
- [6] Mark Chavira and Adnan Darwiche. Compiling Bayesian networks using variable elimination. In Proceedings of the 20th International Joint Conference on Artificial Intelligence (IJCAI) , pages 2443-2449, 2007.
- [7] Mark Chavira and Adnan Darwiche. On probabilistic inference by weighted model counting. Artif. Intell. , 172(6-7):772-799, 2008.
- [8] Mark Chavira, Adnan Darwiche, and Manfred Jaeger. Compiling relational Bayesian networks for exact inference. Int. J. Approx. Reason. , 42(1-2):4-20, 2006.
- [9] Yizuo Chen, Arthur Choi, and Adnan Darwiche. Supervised learning with background knowledge. In PGM , 2020.
- [10] Arthur Choi and Adnan Darwiche. On relaxing determinism in arithmetic circuits. In Proceedings of the Thirty-Fourth International Conference on Machine Learning (ICML) , pages 825-833, 2017.
- [11] Arthur Choi, Doga Kisa, and Adnan Darwiche. Compiling probabilistic graphical models using sentential decision diagrams. In ECSQARU , volume 7958 of Lecture Notes in Computer Science , pages 121-132. Springer, 2013.
- [12] Adnan Darwiche. A logical approach to factoring belief networks. In Dieter Fensel, Fausto Giunchiglia, Deborah L. McGuinness, and Mary-Anne Williams, editors, Proceedings of the Eights International Conference on Principles and Knowledge Representation and Reasoning (KR-02), Toulouse, France, April 22-25, 2002 , pages 409-420. Morgan Kaufmann, 2002.
- [13] Adnan Darwiche. A differential approach to inference in Bayesian networks. J. ACM , 50(3):280305, 2003.
- [14] Adnan Darwiche. Modeling and Reasoning with Bayesian Networks . Cambridge University Press, 2009.
- [15] Adnan Darwiche. An advance on variable elimination with applications to tensor-based computation. In ECAI , volume 325 of Frontiers in Artificial Intelligence and Applications , pages 2559-2568. IOS Press, 2020.
- [16] Adnan Darwiche. Tractable Boolean and arithmetic circuits. In Pascal Hitzler and Md Kamruzzaman Sarker, editors, Neuro-symbolic Artificial Intelligence: The State of the Art . Frontiers in Artificial Intelligence and Applications. IOS Press, 2022. In print.
- [17] Rina Dechter. Bucket elimination: A unifying framework for probabilistic inference. In Proceedings of the Twelfth Annual Conference on Uncertainty in Artificial Intelligence (UAI) , pages 211-219, 1996.
- [18] Paulius Dilkas and Vaishak Belle. Weighted model counting with conditional weights for Bayesian networks. In UAI , 2021.
- [19] Joseph Y. Halpern. Axiomatizing causal reasoning. J. Artif. Intell. Res. , 12:317-337, 2000.
- [20] Yimin Huang and Marco Valtorta. Identifiability in causal bayesian networks: A sound and complete algorithm. In AAAI , pages 1149-1154. AAAI Press, 2006.
- [21] Madelyn Glymour Judea Pearl and Nicholas P. Jewell. Causal Inference in Statistics: A Primer . Wiley, 2016.
- [22] Sanghack Lee, Juan D. Correa, and Elias Bareinboim. General identifiability with arbitrary surrogate experiments. In UAI , volume 115 of Proceedings of Machine Learning Research , pages 389-398. AUAI Press, 2019.
- [23] Daniel Lowd and Pedro M. Domingos. Learning arithmetic circuits. In Proceedings of the 24th Conference in Uncertainty in Artificial Intelligence (UAI) , pages 383-392, 2008.
- [24] Judea Pearl. [bayesian analysis in expert systems]: Comment: Graphical models, causality and intervention. Statistical Science , 8(3):266-269, 1993.
- [25] Judea Pearl. Causal diagrams for empirical research. Biometrika , 82(4):669-688, 1995.
- [26] Judea Pearl. Causality . Cambridge University Press, 2000.
- [27] Judea Pearl. The seven tools of causal inference, with reflections on machine learning. Commun. ACM , 62(3):54-60, 2019.
- [28] Judea Pearl and Dana Mackenzie. The Book of Why: The New Science of Cause and Effect . Basic Books, 2018.
- [29] Judea Pearl and James M. Robins. Probabilistic evaluation of sequential plans from causal models with hidden variables. In Philippe Besnard and Steve Hanks, editors, UAI '95: Proceedings of the Eleventh Annual Conference on Uncertainty in Artificial Intelligence, Montreal, Quebec, Canada, August 18-20, 1995 , pages 444-453. Morgan Kaufmann, 1995.
- [30] Hoifung Poon and Pedro M. Domingos. Sum-product networks: A new deep architecture. In UAI , pages 337-346. AUAI Press, 2011.
- [31] Biao Qin. Differential semantics of intervention in Bayesian networks. In IJCAI , pages 710-716. AAAI Press, 2015.
- [32] Yujia Shen, Arthur Choi, and Adnan Darwiche. Tractable operations for arithmetic circuits of probabilistic models. In NIPS , pages 3936-3944, 2016.
- [33] Prakash P. Shenoy. Binary join trees. In UAI , pages 492-499. Morgan Kaufmann, 1996.
- [34] Ilya Shpitser and Judea Pearl. Identification of joint interventional distributions in recursive semi-markovian causal models. In AAAI , pages 1219-1226. AAAI Press, 2006.
- [35] Jin Tian and Judea Pearl. A general identification condition for causal effects. In AAAI/IAAI , pages 567-573. AAAI Press / The MIT Press, 2002.
- [36] Santtu Tikka, Antti Hyttinen, and Juha Karvanen. Identifying causal effects via context-specific independence relations. In NeurIPS , pages 2800-2810, 2019.
- [37] Benjie Wang, Clare Lyle, and Marta Kwiatkowska. Provable guarantees on the robustness of decision rules to causal interventions. In IJCAI , pages 4258-4265. ijcai.org, 2021.
- [38] Nevin Lianwen Zhang and David Poole. Exploiting causal independence in bayesian network inference. Journal of Artificial Intelligence Research , 5:301-328, 1996.