# DiReCT: Diagnostic Reasoning for Clinical Notes via Large Language Models
**Authors**: {, , }, , {, , }
## Abstract
Large language models (LLMs) have recently showcased remarkable capabilities, spanning a wide range of tasks and applications, including those in the medical domain. Models like GPT-4 excel in medical question answering but may face challenges in the lack of interpretability when handling complex tasks in real clinical settings. We thus introduce the diagnostic reasoning dataset for clinical notes (DiReCT), aiming at evaluating the reasoning ability and interpretability of LLMs compared to human doctors. It contains 511 clinical notes, each meticulously annotated by physicians, detailing the diagnostic reasoning process from observations in a clinical note to the final diagnosis. Additionally, a diagnostic knowledge graph is provided to offer essential knowledge for reasoning, which may not be covered in the training data of existing LLMs. Evaluations of leading LLMs on DiReCT bring out a significant gap between their reasoning ability and that of human doctors, highlighting the critical need for models that can reason effectively in real-world clinical scenarios Code are available https://github.com/wbw520/DiReCT. Data will be released through PhysioNet..
footnotetext: Corresponding author.
## 1 Introduction
Recent advancements of large language models (LLMs) [Zhao et al., 2023] have ushered in new possibilities and challenges for a wide range of natural language processing (NLP) tasks [Min et al., 2023]. In the medical domain, these models have demonstrated remarkable prowess [Anil et al., 2023, Han et al., 2023], particularly in medical question answering (QA) [Jin et al., 2021]. Leading-edge models, such as GPT-4 [OpenAI, 2023a], exhibit profound proficiency in understanding and generating text [Bubeck et al., 2023], even achieved high scores on the United States Medical Licensing Examination (USMLE) questions [Nori et al., 2023].
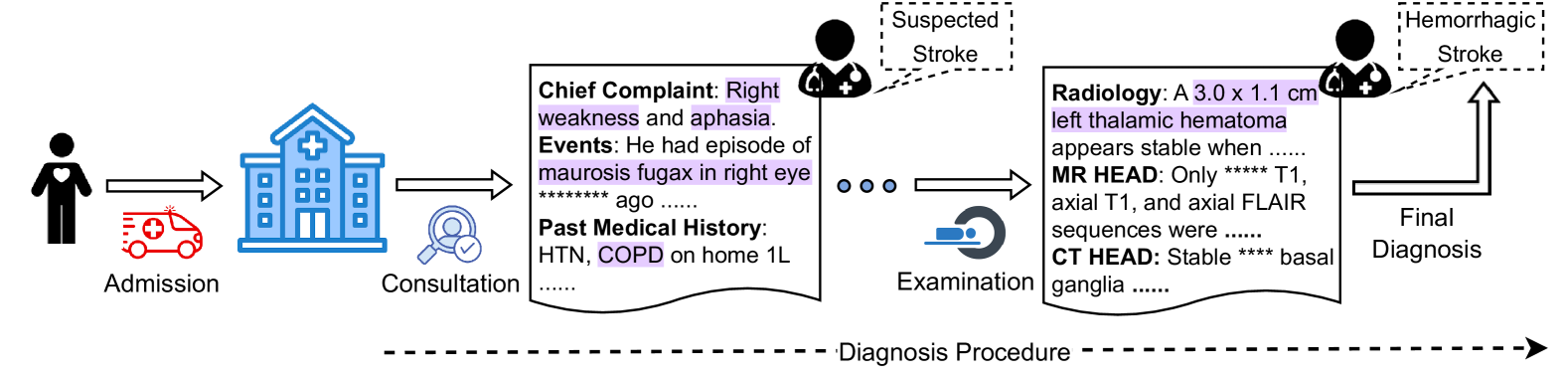
Despite the advancements, interpretability is critical, particularly in medical NLP tasks [Liévin et al., 2024]. Some studies assess this capability over medical QA [Pal et al., 2022, Li et al., 2023, Chen et al., 2024] or natural language inference (NLI) [Jullien et al., 2023]. Putting more attention on interpretability, they use relatively simple tasks as testbeds, taking short text as input. However, tasks in real clinical settings can be more complex [Gao et al., 2023a]. As shown in Figure 1, a typical diagnosis requires comprehending and combining various information, such as health records, physical examinations, and laboratory tests, for further reasoning of possible diseases in a step-by-step manner following the established guidelines. This observation suggests that both perception, or reading, (e.g., finding necessary information in medical record) and reasoning (determining the disease based on the observations) should be counted when evaluating interpretability in LLM-based medical NLP tasks.
For a more comprehensive evaluation of LLMs for supporting diagnosis in a more realistic setting, we propose a Di agnostic Re asoning dataset for C linical no T es (DiReCT). The task basically is predicting the diagnosis from a clinical note of a patient, which is a collection of various medical records, written in natural language. Our dataset contains 511 clinical notes spanning 25 disease categories, sampled from a publicly available database, MIMIC-IV [Johnson et al., 2023]. Each clinical note undergoes fine-grained annotation by professional physicians. The annotators (i.e., the physicians) are responsible for identifying the text, or the observation, in the note that leads to a certain diagnosis, as well as the explanation. The dataset also provides a diagnostic knowledge graph based on existing diagnostic guidelines to facilitate more consistent annotations and to supply a model with essential knowledge for reasoning that might not be encompassed in its training data.
To underscore the challenge offered by our dataset, we evaluate a simple AI-agent based baseline [Xi et al., 2023, Tang et al., 2023] that utilizes the knowledge graph to decompose the diagnosis into a sequence of diagnoses from a smaller number of observations. Our experimental findings indicate that current state-of-the-art LLMs still fall short of aligning well with human doctors.
Contribution. DiReCT offers a new challenge in diagnosis from a complex clinical note with explicit knowledge of established guidelines. This challenge aligns with a realistic medical scenario that doctors are experiencing. In the application aspect, the dataset facilitates the development of a model to support doctors in diagnosis, which is error-prone [Middleton et al., 2013, Liu et al., 2022]. From the technical aspect, the dataset can benchmark models’ ability to read long text and find necessary observations for multi-evidence entailment tree reasoning. As shown in Figure 3, this is not trivial because of the variations in writing; superficial matching does not help, and medical knowledge is vital. Meanwhile, reasoning itself is facilitated by the knowledge graph. The model does not necessarily have the knowledge of diagnostic guidelines. With this choice, the knowledge graph explains the reasoning process, which is also beneficial when deploying such a diagnosis assistant system in practical uses.
<details>
<summary>x1.png Details</summary>
### Visual Description
## Medical Diagnosis Procedure Diagram: Hemorrhagic Stroke Case
### Overview
This image is a horizontal flowchart diagram illustrating the clinical pathway for diagnosing a hemorrhagic stroke in a patient. It depicts a sequential process from patient admission to final diagnosis, using icons, text boxes, and directional arrows to show the flow of information and decision-making. The diagram is designed to show how clinical information, patient history, and diagnostic imaging converge to reach a specific diagnosis.
### Components/Axes
The diagram is structured as a linear process flow from left to right, with a dashed arrow at the bottom labeled **"Diagnosis Procedure"** indicating the overall direction.
**Key Components (from left to right):**
1. **Admission Stage:**
* Icon: A person silhouette with a heart symbol.
* Label: **"Admission"**.
* Connected via an arrow to an ambulance icon, which then points to a hospital building icon.
2. **Consultation Stage:**
* Icon: A magnifying glass over a document.
* Label: **"Consultation"**.
* Leads to the first major information box.
3. **Initial Clinical Information Box:**
* A rectangular text box containing patient history.
* **Text Content:**
* **Chief Complaint:** Right weakness and aphasia.
* **Events:** He had episode of maurosis fugax in right eye ******** ago ......
* **Past Medical History:** HTN, COPD on home 1L ......
* *Note: "maurosis fugax" is likely a misspelling of "amaurosis fugax." Asterisks (********) and ellipses (......) indicate omitted or generalized text.*
4. **Suspected Diagnosis Point:**
* Icon: A doctor silhouette with a stethoscope.
* Speech Bubble: **"Suspected Stroke"**.
* This icon is positioned above and connected to the flow after the initial information box.
5. **Examination Stage:**
* Icon: A person lying down with a scanner (representing imaging).
* Label: **"Examination"**.
* Connected via an arrow to the second major information box.
6. **Radiology Findings Box:**
* A rectangular text box containing diagnostic imaging results.
* **Text Content:**
* **Radiology:** A 3.0 x 1.1 cm left thalamic hematoma appears stable when ......
* **MR HEAD:** Only ***** T1, axial T1, and axial FLAIR sequences were ......
* **CT HEAD:** Stable **** basal ganglia ......
* *Note: Asterisks (*****) and ellipses (......) indicate omitted or generalized text.*
7. **Final Diagnosis Point:**
* Icon: A doctor silhouette with a stethoscope (identical to the first).
* Speech Bubble: **"Hemorrhagic Stroke"**.
* An arrow points from this icon to the label **"Final Diagnosis"**.
**Highlighted Text (in purple):**
* "Right weakness and aphasia"
* "maurosis fugax"
* "HTN, COPD"
* "3.0 x 1.1 cm left thalamic hematoma"
* "basal ganglia"
### Detailed Analysis
The diagram maps a specific clinical case:
1. **Patient Presentation:** The patient is admitted with right-sided weakness and aphasia (inability to speak). A past event of amaurosis fugax (temporary vision loss) in the right eye is noted, along with a history of hypertension (HTN) and chronic obstructive pulmonary disease (COPD) requiring home oxygen.
2. **Initial Clinical Assessment:** Based on the presentation, a stroke is suspected.
3. **Diagnostic Workup:** The patient undergoes examination, specifically neuroimaging (MRI and CT of the head).
4. **Key Radiological Finding:** The imaging reveals a **3.0 x 1.1 cm hematoma (bleed) in the left thalamus**. The report notes it is "stable." Additional findings mention stable changes in the basal ganglia.
5. **Final Diagnosis:** The presence of a thalamic hematoma confirms the diagnosis of a **Hemorrhagic Stroke** (a stroke caused by bleeding in the brain), as opposed to an ischemic stroke (caused by a clot).
### Key Observations
* **Linear, Unidirectional Flow:** The process is shown as a straightforward sequence with no feedback loops or decision branches, simplifying the complex diagnostic process.
* **Information Synthesis:** The final diagnosis is not based on a single data point but on the synthesis of clinical symptoms (weakness, aphasia), patient history (HTN, amaurosis fugax), and definitive imaging evidence (thalamic hematoma).
* **Use of Placeholders:** The asterisks and ellipses indicate this is a template or generalized example, not a complete, specific patient record. The purple highlights draw attention to the critical data points that drive the diagnosis.
* **Spatial Layout:** The two doctor icons are placed at the top of the flow, visually representing the clinician's judgment points ("Suspected" and "Final" diagnosis) that bookend the objective data-gathering stages (Consultation and Examination).
### Interpretation
This diagram serves as an educational or procedural model for the **diagnostic pathway of a hemorrhagic stroke**. It demonstrates the **Peircean investigative process**:
* **Abduction (Inference to the Best Explanation):** The initial symptoms (right weakness, aphasia) and risk factors (HTN) lead to the abductive hypothesis: "Suspected Stroke."
* **Deduction:** If the hypothesis is true (stroke), then specific diagnostic tests (brain imaging) should reveal a corresponding pathology.
* **Induction:** The imaging results (finding a left thalamic hematoma) confirm the hypothesis, leading to the specific inductive conclusion: "Hemorrhagic Stroke."
The diagram emphasizes that the **definitive diagnosis is radiologically confirmed**. The clinical suspicion is necessary to initiate the correct diagnostic pathway, but the objective finding of a brain bleed is what solidifies the final classification. The highlighted terms represent the **critical data chain**: from symptom (right-sided deficit) to risk factor (HTN) to pathological finding (thalamic bleed), all of which are logically connected in the context of stroke diagnosis. The "stable" notations on the imaging reports are important clinical observations, suggesting the bleed is not actively expanding at the time of the scan.
</details>
Figure 1: When a patient is admitted, an initial consultation takes place to collect subjective information. Subsequent observations may then require further examination to confirm the diagnosis.
## 2 Related Works
Natural language explanation. Recent advancements in NLP have led to significant achievements [Min et al., 2023]. However, existing models often lack explainability, posing potential risks [Danilevsky et al., 2020, Gurrapu et al., 2023]. Numerous efforts have been made to address this challenge. One effective approach is to provide a human-understandable plain text explanation alongside the model’s output [Camburu et al., 2018, Rajani et al., 2019]. Another strategy involves identifying evidence within the input that serves as a rationale for the model’s decisions, aligning with human reasoning [DeYoung et al., 2020]. Expanding on this concept, [Jhamtani and Clark, 2020] introduces chain-structured explanations, given that a diagnosis can demand multi-hop reasoning. This idea is further refined by ProofWriter [Tafjord et al., 2021] through a proof stage for explanations, and by [Zhao et al., 2021] through retrieval from a corpus. [Dalvi et al., 2021] proposes the entailment tree, offering more detailed explanations and facilitating inspection of the model’s reasoning. More recently, [Zhang et al., 2024] employed cumulative reasoning to tap into the potential of LLMs to provide explanation via a directed acyclic graph. Although substantial progress has been made, interpreting NLP tasks in medical domains remains an ongoing challenge [Liévin et al., 2024].
Benchmarks of interpretability in the medical domain Several datasets are designed to assess a model’s reasoning together with its interpretability in medical NLP (Table 1). MedMCQA [Pal et al., 2022] and other medical QA datasets [Li et al., 2023, Chen et al., 2024] provide plain text as explanations for QA tasks. NLI4CT [Jullien et al., 2023] uses clinical trial reports, focusing on NLI supported by multi-hop reasoning. N2N2 [Gao et al., 2022] proposes a summarization (Sum) task for a diagnosis based on multiple pieces of evidence in the input clinical note. NEJM CPC [Zack et al., 2023] interprets clinicians’ diagnostic reasoning as plain text for reasoning clinical diagnosis (CD). DR.BENCH [Gao et al., 2023b] aggregates publicly available datasets to assess the diagnostic reasoning of LLMs. Utilizing an multi-evidence entailment tree explanation, DiReCT introduces a more rigorous task to assess whether LLMs can align with doctors’ reasoning in real clinical settings.
Table 1: Comparison of existing datasets for medical reasoning tasks and ours. “t” and “w” mean tokens and words for the length of input, respectively.
| Dataset | Task | Data Source | Length | Explanation | # Cases |
| --- | --- | --- | --- | --- | --- |
| MedMCQA [Pal et al., 2022] | QA | Examination | 9.93 t | Plain Text | 194,000 |
| ExplainCPE [Li et al., 2023] | QA | Examination | 37.79 w | Plain Text | 7,000 |
| JAMA Challenge [Chen et al., 2024] | QA | Clinical Cases | 371 w | Plain Text | 1,524 |
| Medbullets [Chen et al., 2024] | QA | Online Questions | 163 w | Plain Text | 308 |
| N2N2 [Gao et al., 2022] | Sum | Clinical Notes | 785.46 t | Evidences | 768 |
| NLI4CT [Jullien et al., 2023] | NLI | Clinical Trail Reports | 10-35 t | Multi-hop | 2,400 |
| NEJM CPC [Zack et al., 2023] | CD | Clinical Cases | - | Plain Text | 2,525 |
| DiReCT (Ours) | CD | Clinical Notes | 1074.6 t | Entailment Tree | 511 |
## 3 A benchmark for Clinical Notes Diagnosis
This section first detail clinical notes (Section 3.1). We also describes the knowledge graph that encodes existing guidelines (Section 3.2). Our task definition, which tasks a clinical note and the knowledge graph as input is given in Section 3.4. We then present our annotation process for clinical notes (Section 3.3) and the evaluation metrics (Section 3.5).
### 3.1 Clinical Notes
Clinical notes used in DiReCT are stored in the SOAP format [Weed, 1970]. A clinical note comprises four components: In the subjective section, the physician records the patient’s chief complaint, the history of present illness, and other subjective experiences reported by the patient. The objective section contains structural data obtained through examinations (inspection, auscultation, etc.) and other measurable means. The assessment section involves the physician’s analysis and evaluation of the patient’s condition. This may include a summary of current status, etc. Finally, the plan section outlines the physician’s proposed treatment and management plan. This may include prescribed medications, recommended therapies, and further investigations. A clinical note also includes a primary discharge diagnosis (PDD) in the assessment section.
DiReCT’s clinical notes are sourced from the MIMIC-IV dataset [Johnson et al., 2023] (PhysioNet Credentialed Health Data License 1.5.0), which encompasses over 40,000 patients admitted to the intensive care units. Each note contains clinical data for a patient. To construct DiReCT, we curated a subset of 511 notes whose PDDs fell within one of 25 disease categories $i$ in 5 medical domains.
In our task, a note $R=\{r\}$ is an excerpt of 6 clinical data in the subjective and objective sections (i.e., $|R|=6$ ): chief complaint, history of present illness, past medical history, family history, physical exam, and pertinent results. We excluded data, such as review system and social history, because they are often missing in the original clinical notes and are less relevant to the diagnosis. We also identified the PDD $d^⋆$ associated with $R$ . All clinical notes in DiReCT are related to only one PDD, and there is no secondary discharge diagnosis. The set of $d^⋆$ ’s for all $R$ ’s collectively forms $D^⋆$ . We manually removed any descriptions that disclose the PDD in $R$ .
### 3.2 Diagnostic Knowledge Graph
Existing knowledge graphs for the medical domain, e.g., UMLS KG [Bodenreider, 2004], lack the ability to provide specific clinical decision support (e.g., diagnostic threshold, context-specific data, dosage information, etc.), which are critical for accurate diagnosis.
Our knowledge graphs $K=\{K_i\}_i$ is a collection of graph $K_i$ for disease category $i$ . $K_i$ is based on the diagnosis criteria in existing guidelines (refer to supplementary material for details). $K_i$ ’s nodes are either premise $p∈P_i$ (medical statement, e.g., Headache is a symptom of) and diagnoses $d∈D_i$ (e.g., Suspected Stroke). $K_i$ consists of two different types of edges. One is premise-to-diagnosis edges $S_i=\{(p,d)\}$ , where $p∈P_i$ and $d∈D_i$ ; an edge is from $p$ to $d$ . This edge represents the necessary premise $p$ to make a diagnosis $d$ . We refer to them as supporting edges. The other is diagnosis-to-diagnosis edges $F_i=\{(d,d^\prime)\}$ , where $d,d^\prime∈D_i$ and the edge is from $d$ to $d^\prime$ , which represents the diagnostic flow. These edges are referred to as procedural edges.
A disease category is defined according to an existing guideline, which starts from a certain diagnosis; therefore, a procedural graph $G_i=(D_i,F_i)$ has only one root node and arbitrarily branches toward multiple leaf nodes that represent PDDs (i.e., the clinical notes in DiReCT are chosen to cover all leaf nodes of $G_i$ ). Thus, $G_i$ is a tree. We denote the set of the leaf nodes (or PDDs) as $D^⋆_i⊂D_i$ . The knowledge graph is denoted by $K_i=(D_i,P_i,S_i,F_ i)$ .
<details>
<summary>x2.png Details</summary>
### Visual Description
## Clinical Decision Pathway: Acute Coronary Syndrome (ACS) Diagnosis
### Overview
The image is a clinical flowchart or decision pathway diagram illustrating the diagnostic process for Acute Coronary Syndrome (ACS). It maps the progression from initial symptoms and clinical findings to specific ACS diagnoses, using a combination of blue boxes (representing symptoms, signs, or criteria) and gray boxes (representing diagnostic states or categories). The flow moves generally from left to right, with arrows indicating the logical progression and branching based on specific criteria.
### Components/Axes
The diagram is structured into three main conceptual regions:
1. **Left Region (Initial Presentation):** Contains blue boxes listing symptoms and signs that lead to a suspicion of ACS.
2. **Central Region (Diagnostic Progression):** Contains gray boxes representing escalating levels of diagnostic certainty ("Suspected ACS" -> "Strongly Suspected ACS") and the primary branching point for ACS types.
3. **Right Region (Final Classifications):** Contains gray boxes for the final diagnostic categories (STEMI-ACS, NSTE-ACS, NSTEMI-ACS, UA) and blue boxes listing the specific criteria that lead to each.
**Legend/Color Coding:**
* **Blue Boxes:** Clinical symptoms, signs, or diagnostic criteria (e.g., "Breathlessness is a symptom...", "ST Elevation is criteria...").
* **Gray Boxes:** Diagnostic states or categories (e.g., "Suspected ACS", "STEMI-ACS").
* **Arrows:** Indicate the flow of clinical reasoning. Black arrows show contributing factors or criteria leading to a state. Red arrows show the primary diagnostic pathway progression.
### Detailed Analysis
**Textual Content & Flow:**
**1. Initial Symptoms & Signs (Left Region - Blue Boxes):**
* "Breathlessness is a symptom ..." (Top-left, points to "Suspected ACS")
* "Arrhythmias is ..." (Far-left, points to "Suspected ACS")
* "Third Heart Sound ..." (Bottom-left, points to "Suspected ACS")
* "Any Severe Presentations ..." (Bottom-center, points to "Strongly Suspected ACS")
**2. Diagnostic States (Gray Boxes - Central Flow):**
* **Suspected ACS:** The initial diagnostic state, reached from the symptoms listed above.
* **Strongly Suspected ACS:** The next state, reached from "Suspected ACS" (red arrow) and informed by "Any Severe Presentations..." (black arrow).
**3. Branching Criteria & Final Diagnoses (Right Region):**
From "Strongly Suspected ACS," the pathway branches via red arrows:
* **Branch 1 (Top Path):**
* **Criteria (Blue Box):** "ST Elevation is criteria ..." (Points to "STEMI-ACS")
* **Diagnosis (Gray Box):** **STEMI-ACS** (ST-Elevation Myocardial Infarction - Acute Coronary Syndrome)
* **Branch 2 (Bottom Path):**
* **Criteria (Blue Box):** "non-ST Elevation ..." (Points to "NSTE-ACS")
* **Diagnosis (Gray Box):** **NSTE-ACS** (Non-ST-Elevation Acute Coronary Syndrome)
From **NSTE-ACS**, the pathway further differentiates:
* **Sub-Branch 2a (Top Path from NSTE-ACS):**
* **Criteria (Blue Boxes):**
* "hs-cTn Exceeded ..." (High-sensitivity cardiac Troponin)
* "Cardiac Troponin ↑" (Increased)
* **Diagnosis (Gray Box):** **NSTEMI-ACS** (Non-ST-Elevation Myocardial Infarction - ACS)
* **Sub-Branch 2b (Bottom Path from NSTE-ACS):**
* **Criteria (Blue Box):** "No Obvious ECG ..." (Electrocardiogram)
* **Diagnosis (Gray Box):** **UA** (Unstable Angina)
### Key Observations
1. **Hierarchical Diagnosis:** The flowchart presents a clear hierarchy: from general suspicion ("Suspected ACS") to strong suspicion, and then to specific, mutually exclusive final diagnoses (STEMI, NSTEMI, UA).
2. **Critical Decision Points:** The two major branching points are:
* The presence or absence of **ST Elevation** on ECG, which separates STEMI from NSTE-ACS.
* Within NSTE-ACS, the status of **cardiac troponin** levels, which separates NSTEMI (elevated troponin) from Unstable Angina (no obvious ECG changes, implying troponin is not elevated or the presentation is primarily ischemic without biomarker rise).
3. **Symptom vs. Criteria:** The initial blue boxes list non-specific symptoms (breathlessness, arrhythmias, third heart sound) that raise suspicion. The later blue boxes list more specific diagnostic criteria (ST elevation, troponin levels) that confirm a specific diagnosis.
4. **Incomplete Text:** Several blue boxes contain ellipses ("..."), indicating that the text is truncated. The full criteria or descriptions are not visible in this diagram.
### Interpretation
This flowchart represents a standardized clinical algorithm for triaging and diagnosing patients with suspected Acute Coronary Syndrome. It visually encodes the **Peircean investigative logic** of medical diagnosis:
* **Abduction:** Initial symptoms (breathlessness, arrhythmias) lead to the abductive inference of a possible ACS ("Suspected ACS").
* **Deduction:** Based on the established diagnostic rules (e.g., "If ST elevation is present, then it is STEMI"), the clinician deduces the specific type of ACS from the "Strongly Suspected" state.
* **Induction:** The final diagnosis (e.g., NSTEMI) is confirmed by specific, observable facts (elevated hs-cTn), which inductively support the general category.
The diagram emphasizes that ACS is not a single entity but a spectrum. The critical differentiation between **STEMI** (a full-thickness heart attack requiring immediate reperfusion) and **NSTE-ACS** (which includes NSTEMI and UA) is driven primarily by the ECG finding of ST elevation. The subsequent split between **NSTEMI** and **UA** hinges on myocardial necrosis, as evidenced by elevated cardiac troponins. The pathway underscores the importance of integrating clinical presentation, ECG findings, and biomarker results to arrive at an accurate diagnosis, which is essential for determining the correct and urgent treatment strategy. The truncated text suggests this is a simplified overview, and the full criteria would be detailed in accompanying medical guidelines.
</details>
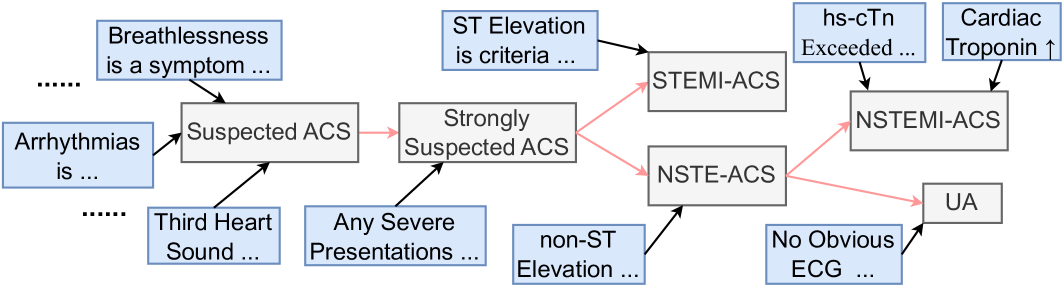
Figure 2: A part of $K_i$ for $i$ being Acute Coronary Syndromes.
Figure 2 shows a part of $K_i$ , where $i$ is Acute Coronary Syndromes (ACS). Premises in $P_i$ and diagnoses in $D_i$ are given in the blue and gray boxes, while PDDs in $D^⋆_i$ are ones without outgoing edges (i.e., STEMI-ACS and NSTEMI-ACS, and UA). The black and red arrows are edges in $S$ and $F$ , respectively, where the black arrows indicate the supporting edges.
$K$ serves two essential functions: (1) They serve as the gold standard for annotation, guiding doctors in the precise and uniform interpretation of clinical notes. (2) Our task also allows a model to use them to ensure the output from an LLM can be closely aligned with the reasoning processes of medical professionals.
### 3.3 Data Annotation
Let $d^⋆∈D^⋆_i$ denote the PDD of disease category $i$ associated with $R$ . We can find a subgraph $K_i(d^⋆)$ of $K_i$ that contains all ancestors of $d^⋆$ , including premises in $P_i$ . We also denote the set of supporting edges in $K_i(d^⋆)$ as $S_i(d^⋆)$ . Our annotation process is, for each supporting edge $(p,d)∈S_i(d^⋆)$ , to extract observation $o∈O$ in $R$ (highlighted text in the clinical note in Figure 3) and provide rationalization $z$ of this deduction why $o$ is a support for $d$ or corresponds to $p$ . All annotations strictly follow the procedural flow in $K_i$ , and each observation is only related to one diagnostic node. If $R$ does not provide sufficient observations for the PDD (which may happen when a certain test is omitted), the annotators were asked to add plausible observations to $R$ . This choice compromises the fidelity of our dataset to the original clinical notes, but we chose it for the completeness of the dataset. They form the explanation $E=\{(o,z,d)\}$ for $(R,d^⋆)$ . This annotation process was carried out by 9 clinical physicians and subsequently verified for accuracy and completeness by three senior medical experts.
Table 2: Statistics of DiReCT.
| Medical domain | # cat. | # samples | $|D_i|$ | $|D^⋆_i|$ | $|O|$ | Length |
| --- | --- | --- | --- | --- | --- | --- |
| Cardiology | 7 | 184 | 27 | 16 | 8.7 | 1156.6 t |
| Gastroenterology | 4 | 103 | 11 | 7 | 4.3 | 1026.0 t |
| Neurology | 5 | 77 | 17 | 11 | 11.9 | 1186.3 t |
| Pulmonology | 5 | 92 | 26 | 17 | 10.7 | 940.7 t |
| Endocrinology | 4 | 55 | 20 | 14 | 6.9 | 1063.5 t |
| Overall | 25 | 511 | 101 | 65 | 8.5 | 1074.6 t |
Table 2 summarizes statistics of our dataset. The second and third columns (“# cats.” and “# samples”) show the numbers of disease categories and samples in the respective medical domains. $|D_i|$ and $|D_i^⋆|$ are the total numbers of diagnoses (diseases) and PDDs, summed over all diagnostic categories in the medical domain, respectively. $|O|$ is the average number of annotated observations. “Length” is the average number of tokens in $R$ .
<details>
<summary>x3.png Details</summary>
### Visual Description
## Clinical Document: Heart Failure Diagnostic Pathway
### Overview
The image is a composite technical diagram illustrating the diagnostic reasoning process for a patient presenting with symptoms suggestive of heart failure. It integrates a clinical note (left), a rationale section linking findings to diagnostic criteria (center), and a diagnostic flowchart (right). The document demonstrates how specific patient data points map to established medical criteria to arrive at a diagnosis of Heart Failure with mildly reduced Ejection Fraction (HFmrEF).
### Components/Axes
The image is segmented into three primary vertical regions:
1. **Left Region: Clinical Note**
* **Structure:** A structured medical note with labeled sections.
* **Sections & Content:**
* **Chief Complaint:** "Scrotal and leg swelling ..."
* **History of Present Illness:** Describes 3 days of worsening swelling, similar to a prior admission for acute CHF. Notes consistent EKG, mildly enlarged left ventricle, and good response to IV diuretics (I/V diuretics).
* **Past Medical History:** Lists Diabetes, Hypertension, CKD (stage 3), GERD, history of [REDACTED] embolization, Pneumonia, Osteoarthritis, Asthma.
* **Family History:** "There is no family history of ... artery ..."
* **Physical Exam:**
* LUNG: "bibasilar rales that do not clear with deep inspiration ..."
* ABDOMEN: "soft, nontender, ... all quadrants."
* EXTREMITIES: "bilateral pitting edema to the sacrum extending to the low abdomen. Warm. Well perfused."
* CV: "RRR, no EOMI, PERRLA."
* **Recent Results:**
* "1:50PM BLOOD WBC-8.0 *RBC-3.26* Hgb-9.3* Hct-30.9* MCHC-29.9* 11:30AM BLOOD proBNP-3843 ..."
* "Overall left ventricular systolic function is mildly depressed (LVEF= 45-50 %) without regional wall motion abnormalities. ... imaging suggests an increased ... filling pressure (PCWP=... mmHg)."
2. **Center Region: Rationale**
* **Structure:** A series of text boxes with arrows pointing from specific findings in the Clinical Note to explanatory diagnostic statements.
* **Boxes & Connections (from top to bottom):**
* Arrow from "Scrotal and leg swelling" and "bilateral pitting edema..." → Box: "Peripheral edema is a sign of heart failure."
* Arrow from "Hypertension" in Past Medical History → Box: "Hypertension is the risk factor of heart failure."
* Arrow from "proBNP-3843" in Recent Results → Box: "NT-proBNP 3843≥125pg/ml is a diagnostic criteria of strong HF."
* Arrow from "mildly enlarged" left ventricle and "increased ... filling pressure" → Box: "Cardiac structure abnormalities are diagnostic criteria of heart failure."
* Arrow from "LVEF= 45-50 %" → Box: "Cardiac systolic dysfunction ~49% can lead to the diagnosis of HFmrEF."
3. **Right Region: Diagnosis Flowchart**
* **Structure:** A vertical flowchart with rectangular boxes connected by downward-pointing arrows.
* **Flow:** Suspected HF → Strongly Suspected HF → HF → HFmrEF.
### Detailed Analysis
**Clinical Data Extraction:**
* **Key Lab Value:** NT-proBNP = 3843 pg/ml. The rationale explicitly states this meets the diagnostic criterion of ≥125 pg/ml for "strong HF."
* **Echocardiogram Finding:** Left Ventricular Ejection Fraction (LVEF) is 45-50%. The rationale interprets this as "~49%" systolic dysfunction, leading to the specific diagnosis of HFmrEF (typically defined as LVEF 41-49%).
* **Supporting Signs:** Peripheral edema (scrotal, leg, sacral pitting edema), bibasilar rales, history of hypertension, and evidence of increased cardiac filling pressure (PCWP value redacted).
* **Redacted Information:** Several terms are obscured with asterisks (`...`), including specific medications, a prior embolization site, and the exact PCWP value.
**Diagnostic Logic Flow:**
The rationale section creates a direct, evidence-based link between patient findings and diagnostic rules:
1. **Symptom (Edema)** → Sign of HF.
2. **Risk Factor (Hypertension)** → Predisposes to HF.
3. **Biomarker (NT-proBNP 3843)** → Meets quantitative criterion for strong HF.
4. **Structural Abnormality (Enlarged LV, increased filling pressure)** → Meets structural criterion for HF.
5. **Functional Abnormality (LVEF 45-50%)** → Specifies the subtype as HFmrEF.
This cumulative evidence progresses the diagnosis from "Suspected" to "Strongly Suspected," confirms "HF," and finally specifies "HFmrEF."
### Key Observations
1. **Quantitative Thresholds:** The document explicitly references diagnostic thresholds (NT-proBNP ≥125 pg/ml, LVEF ~49% for HFmrEF), showing a criteria-based diagnostic approach.
2. **Multi-Factorial Diagnosis:** The diagnosis is not based on a single finding but on the convergence of symptoms (edema), signs (rales), risk factors (HTN), biomarkers (BNP), and imaging (echo).
3. **Specificity in Classification:** The pathway doesn't stop at a general "Heart Failure" diagnosis but uses the LVEF value to specify the subtype (HFmrEF), which has different therapeutic implications.
4. **Data Gaps:** The redacted information (PCWP value, specific medications, embolization history) represents missing data that would be relevant for a complete clinical picture but is not essential for the demonstrated diagnostic logic.
### Interpretation
This diagram serves as a pedagogical or clinical decision-support tool that visualizes the **Peircean abductive reasoning** common in medicine: starting from an observation (symptoms/signs), applying known medical rules (diagnostic criteria), and inferring the most plausible explanation (HFmrEF diagnosis).
* **What the Data Suggests:** The patient's clinical profile is a classic presentation of heart failure, specifically the mildly reduced ejection fraction subtype. The elevated NT-proBNP is a strong objective indicator of cardiac stress, corroborating the subjective symptoms and physical exam findings.
* **How Elements Relate:** The three panels form a logical chain: **Evidence (Note)** → **Reasoning (Rationale)** → **Conclusion (Diagnosis Flowchart)**. The arrows in the Rationale section are critical, as they explicitly map raw data to medical knowledge, transforming information into actionable diagnosis.
* **Notable Patterns/Anomalies:** The pattern is one of **convergent validity**—multiple independent lines of evidence point to the same conclusion. There are no apparent anomalies or contradictory findings in the presented data. The LVEF of 45-50% is the key data point that refines the diagnosis from generic HF to the specific HFmrEF category, highlighting the importance of precise measurement in guiding classification and subsequent treatment.
</details>
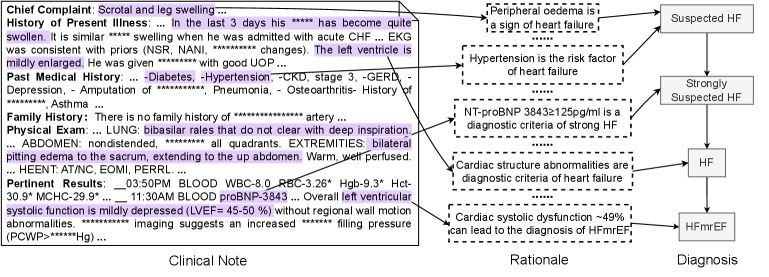
Figure 3: An annotation sample of Heart Failure (HF). The left part is the clinical note alongside extracted observations by a doctor. The middle part outlines the steps of the rationale for the premise corresponding to each diagnostic node shown in the right part.
### 3.4 Task Definition
We propose two tasks with different levels of supplied external knowledge. The first task is, given $R$ and $G$ , to predict the associated PDD $d^⋆$ and generate an explanation $E$ that explains the model’s diagnostic procedure from $R$ to $d^⋆$ , i.e., letting $M$ denote a model:
$$
\displaystyle\hat{d}^⋆,\hat{E}=M(R,G), \tag{1}
$$
where $\hat{d}^⋆∈∪_iD^⋆_i$ and $\hat{E}$ are predictions for the PDD and explanation, respectively. With this task, the knowledge of specific diagnostic procedures in existing guidelines can be used for prediction, facilitating interpretability. The second task takes $K$ as input instead of $G$ , i.e.,:
$$
\displaystyle\hat{d}^⋆,\hat{E}=M(R,K). \tag{2}
$$
This task allows for the use of broader knowledge of premises for prediction. One may also try a task without any external knowledge.
### 3.5 Evaluation Metrics
We designed three metrics to quantify the predictive performance over our benchmark.
(1) Accuracy of diagnosis $\textit{Acc}^diag$ evaluates if a model can correctly identify the diagnosis. $\textit{Acc}^diag=1$ if $d^⋆=\hat{d}$ , and $\textit{Acc}^diag=0$ otherwise. The average is reported.
(2) Completeness of observations $\textit{Obs}^comp$ evaluates whether a model extracts all and only necessary observations for the prediction. Let $O$ and $\hat{O}$ denote the sets of observations in $E$ and $\hat{E}$ , respectively. The metric is defined as $\textit{Obs}^comp=|O∩\hat{O}|/|O∪ \hat{O}|$ , where the numerator is the number of observations that are common in both $O$ and $\hat{O}$ . We find the common observations with an LLM (refer to the supplementary material for more detail). This metric simultaneously evaluates the correctness of each observation and the coverage. To supplement it, we also report the precision $\textit{Obs}^pre$ and recall $\textit{Obs}^rec$ , given by $\textit{Obs}^pre=|O∩\hat{O}|/|\hat{O}|$ and $\textit{Obs}^rec=|O∩\hat{O}|/|O|$ .
(3) Faithfulness of explanations Faith evaluates if the diagnostic flow toward the PDD is fully supported by observations with faithful rationalizations. This involves establishing a one-to-one correspondence between deductions in the prediction and the ground truth. We use the correspondences established for computing $\textit{Obs}^comp$ . Let $o∈O$ and $\hat{o}∈\hat{O}$ denote corresponding observations. This correspondence is considered successful if $z$ and $\hat{z}$ as well as $d$ and $\hat{d}$ associated with $o$ and $\hat{o}$ matches. Let $m(E,\hat{E})$ denote the number of successful matches. We use the ratio of $m(E,\hat{E})$ to $|O∩\hat{O}|$ and $|O∪\hat{O}|$ as evaluation metrics $\textit{Exp}^com$ and $\textit{Exp}^all$ , respectively, to see failures come from observations or explanations and diagnosis.
## 4 Baseline
<details>
<summary>x4.png Details</summary>
### Visual Description
## [System Architecture Diagram]: Clinical Diagnostic Reasoning Workflow
### Overview
This image is a technical flowchart illustrating a multi-stage system for processing clinical notes to arrive at a medical diagnosis. The diagram depicts a workflow that transforms unstructured clinical text into structured observations, integrates them with a diagnostic knowledge graph (KG), and uses iterative reasoning to generate diagnostic hypotheses. The process is cyclical and incorporates feedback loops.
### Components/Axes
The diagram is organized into several interconnected functional blocks, flowing generally from left to right.
**1. Input Source (Leftmost):**
* **Component:** `Clinical Note` (represented by a clipboard icon with a medical cross and a waveform).
* **Output:** Feeds into two parallel processing paths.
**2. Initial Processing Blocks:**
* **`Narrowing-down`** (Green box, top-left): Receives input from the Clinical Note.
* **`Perception`** (Purple box, center-left): Receives input from both the Clinical Note and the `Narrowing-down` block. An arrow labeled `i` points from `Narrowing-down` to `Perception`.
**3. Structured Data Extraction (Center):**
* **`Observations`** (Large white box, center): This is a key output of the `Perception` block. It contains a numbered list of extracted clinical findings.
* **`Diagnostic KG`** (Teal-bordered box, bottom-center): Represents a Diagnostic Knowledge Graph. It contains a network of nodes (blue circles) and directed edges (black and red arrows) connecting labeled nodes (`a1`, `a2`, `a3`, `a4`, `a5`).
**4. Reasoning Engine (Right Side):**
* **`Reasoning`** (Two grey boxes with gear icons, stacked vertically on the right): These blocks represent the core reasoning process. They receive inputs from both the `Observations` list and the `Diagnostic KG`.
* **Iterative Process:** A circular arrow (`↻`) between the two `Reasoning` blocks indicates an iterative or cyclical reasoning process.
* **Output Diagrams:** Each `Reasoning` block outputs a small diagram showing a subset of the observations (numbered circles ①-⑤) connected to diagnostic nodes (`a1`, `a2`, `a4`), illustrating the formation of diagnostic hypotheses.
**5. Connecting Elements:**
* **Arrows:** Solid black arrows indicate the primary data flow. Teal-colored arrows specifically show data flowing from the `Diagnostic KG` into the `Reasoning` blocks.
* **`Rationale`** (Label on an arrow): An arrow labeled "Rationale" connects the `Observations` box to the first output diagram of the reasoning process.
### Detailed Analysis
**Textual Content Transcription:**
* **Observations List:**
1. Elevated blood pressures
2. CXR showed mild pulmonary edema
3. CHF/Cardiomyopathy
4. Severe LV diastolic dysfunction
5. BPs: 148/98, 156/93
* `......` (Ellipsis indicates the list is not exhaustive).
* **Clinical Note Components (in the rounded rectangle below the note icon):**
* `r1: Chief Complaint`
* `r2: History of Present Illness`
* `r3: Past Medical History`
* `r4: Family History`
* `r5: Physical Examination`
* `r6: Pertinent Results`
* **Diagnostic KG Nodes:** The visible labeled nodes are `a1`, `a2`, `a3`, `a4`, and `a5`. The graph shows directed relationships, with some edges highlighted in red (e.g., `a1` -> `a2`, `a2` -> `a3`, `a2` -> `a4`).
* **Reasoning Output Diagrams:** These show the mapping of observations to diagnostic nodes.
* **Top Diagram:** Observations ①, ②, ③, ④, ⑤ all point to node `a1`.
* **Middle Diagram:** Observations ①, ②, ③, ④, ⑤ point to `a1`; a red arrow connects `a1` to `a2`.
* **Bottom Diagram:** Observations ①, ②, ③, ④, ⑤ point to `a1`; red arrows connect `a1` to `a2` and `a2` to `a4`.
### Key Observations
1. **Structured Extraction:** The system explicitly extracts discrete, numbered observations from free-text clinical notes.
2. **Knowledge Integration:** The reasoning process is not based solely on the extracted observations but is actively informed by a pre-existing Diagnostic Knowledge Graph (`Diagnostic KG`).
3. **Iterative Hypothesis Refinement:** The presence of two `Reasoning` blocks connected by a cycle symbol suggests the system refines its diagnostic hypotheses over multiple iterations.
4. **Evidence Linking:** The output diagrams visually demonstrate how specific clinical observations (evidence) are linked to form a chain of diagnostic reasoning (e.g., evidence leads to `a1`, which then implies `a2`, which then implies `a4`).
5. **Comprehensive Input:** The system considers a wide range of clinical note sections (`r1` through `r6`), indicating a holistic approach to data ingestion.
### Interpretation
This diagram represents a **hybrid AI system for medical diagnosis** that combines natural language perception with symbolic reasoning.
* **What it demonstrates:** The workflow shows a pipeline for converting unstructured medical text into actionable, structured knowledge. It emphasizes that diagnosis is not a single-step classification but a **reasoning process** that connects evidence (Observations) to a network of medical concepts (Diagnostic KG) through iterative logic.
* **Relationships between elements:** The `Perception` module acts as a bridge between raw text and structured data. The `Diagnostic KG` provides the medical ontology and causal/associative relationships necessary for reasoning. The `Reasoning` engine is the core that performs abductive and deductive inference, using the KG to explain the observations and generate a differential diagnosis.
* **Notable patterns:** The flow from a single set of observations to progressively more complex diagnostic chains (`a1` -> `a1→a2` -> `a1→a2→a4`) illustrates how the system builds a coherent explanatory model for the patient's condition. The "Rationale" arrow underscores that the system's output is meant to be interpretable, linking conclusions back to the original evidence.
* **Underlying principle:** The architecture embodies a **Peircean investigative approach**—moving from the surprising fact (the clinical note) to the observation of signs (extracted findings), and then to the formulation of an explanatory hypothesis (the diagnostic chain) that best accounts for those signs, using a structured knowledge base to guide and validate the reasoning.
</details>
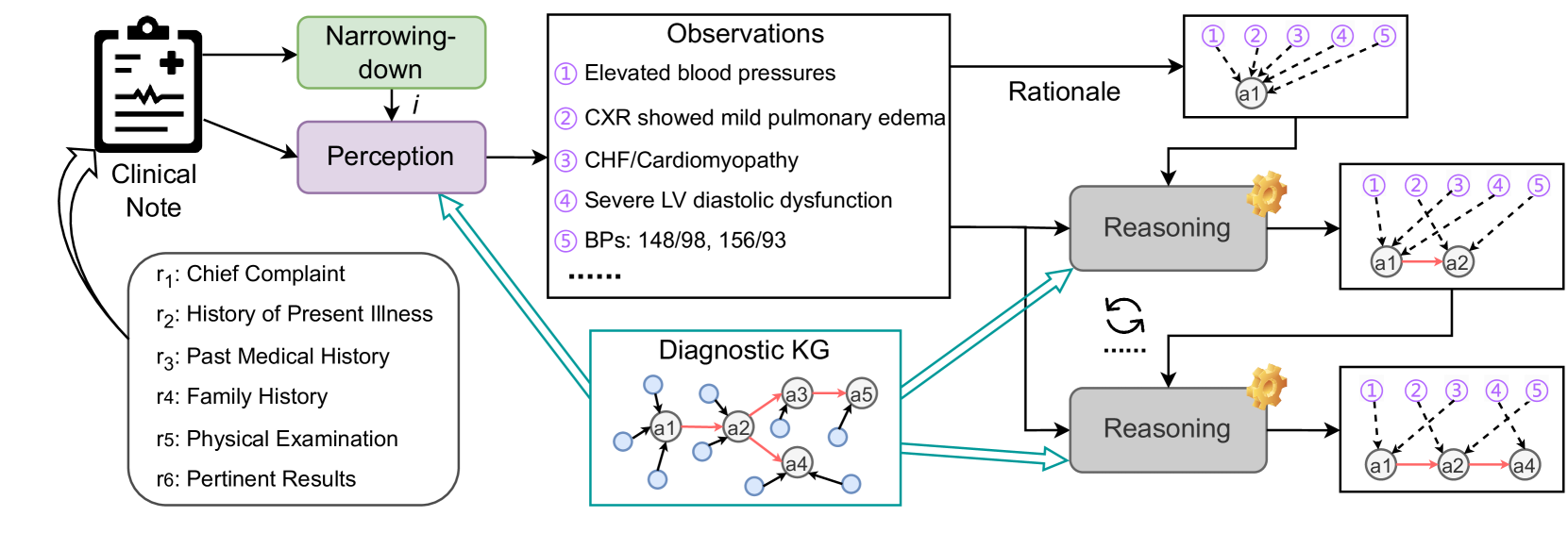
Figure 4: Pipeline of our baseline. The dotted line in the right-most boxes means deductions from an observation to a diagnosis.
Figure 4 shows an overview of our baseline with three LLM-based modules narrowing-down, perception, and reasoning (refer to the supplementary material for more details). The narrowing-down module $U$ takes $R$ as input to make a prediction $\hat{i}$ of the disease category, i.e., $\hat{i}=U(R)$ .
Let $d_t∈D_\hat{i}$ be the diagnosis that has been reached with $t$ iterations over $K_\hat{i}$ , where $t$ corresponds to the depth of node $d_t$ and so is less than or equal to the depth of $K_i$ . $d_0$ is the root node of $K_\hat{i}$ . For $d_0$ , we apply the perception module to extract all observations in $R$ and explanation $E_0$ to support $d_0$ as
$$
\displaystyle\hat{O},\hat{E_0}=W(d_0,K_\hat
{i}). \tag{3}
$$
$K_\hat{i}$ is supplied to facilitate the model to extract all observations for the following reasoning process. We used only pairs of an observation and a premise. We abuse $K$ to mean this for notation simplicity.
Diagnosis $d_t$ identifies the set $\{d_n\}_n$ of its children and so the set $P_\hat{i}(\{d_n\}_n)=\{p∈P_i|(p,d_n)∈ S_i,d_n∈\{d_n\}\}$ of premises that support $d_n$ . Therefore, our reasoning module $V$ iteratively and greedily identifies the next step’s diagnosis (i.e., $d_t+1$ ) from $\{d_n\}_n$ , making a rationalization for each deduction. That is, $V$ verifies whether there exist $o$ ’s in $\hat{O}$ that supports one $d_n$ . If $d_n$ is fully supported, $d_n$ is identified as $d_t+1$ for the $(t+1)$ -th iteration, i.e.,
$$
\displaystyle d_t+1,\hat{E}_t+1=V(\hat{O},\{d_n\},
P_\hat{i}(\{d_n\}_n)). \tag{4}
$$
Otherwise, the reasoning module fails. $V$ is repeated until $d_t^\prime$ in $D^⋆_\hat{i}$ is found or it fails. In our annotation, each observation contributes to deducing only one $d_t$ . Therefore, if an observation in $\hat{E}_t+1$ is included in the preceding sets of explanations $\hat{E}_0$ to $\hat{E}_t$ , the corresponding explanation in the preceding sets is removed.
## 5 Experiments
### 5.1 Experimental Setup
We assess the reasoning capabilities of 7 recent LLMs from diverse families and model sizes, including 5 instruction-tuned models that are openly accessible: LLama3 8B and 70B [AI@Meta, 2024], Zephyr 7B [Tunstall et al., 2023], Mistral 7B [Jiang et al., 2023], and Mixtral 8 $×$ 7B [Jiang et al., 2023]. We have also obtained access to private versions of the GPT-3.5 turbo [OpenAI, 2023b] and GPT-4 turbo [OpenAI, 2023a] These two models are housed on a HIPPA-compliant instance within Microsoft Azure AI Studio. No data is transferred to either Microsoft or OpenAI. This secure environment enables us to safely conduct experiments with the MIMIC-IV dataset, in compliance with the Data Use Agreement., which are high-performance closed-source models. Each LLM is utilized to implement our baseline’s narrowing-down, perception, and reasoning modules. The temperature is set to 0. For computing evaluation metrics, we use LLama3 8B with few-shot prompts to make correspondences between $O$ and $\hat{O}$ as well as to verify a match between predicted and ground-truth explanations (refer to the supplementary material for more details).
### 5.2 Results
Comparison among LLMs. Table 3 shows the performance of our baseline built on top of various LLMs. We first evaluate a variant of our task that takes graph $G=\{G_i\}$ consisting of only procedural flow as external knowledge instead of $K$ . Comparison between $G$ and $K$ demonstrates the importance of supplying premises with the model and LLMs’ capability to make use of extensive external knowledge that may be superficially different from statements in $R$ . Subsequently, some models are evaluated with our task using $K$ . In addition to the metrics in Section 3.5, we also adopt the accuracy of disease category $\textit{Acc}^cat$ , which gives 1 when $\hat{i}=i$ , as our baseline’s performance depends on it.
Table 3: Diagnostic reasoning ability of different LLMs under the proposed baseline method.
| | | Diagnosis | Observation | Explanation | | | | |
| --- | --- | --- | --- | --- | --- | --- | --- | --- |
| Task | Models | Acc ${}^cat$ | Acc ${}^diag$ | $\textit{Obs}^pre$ | $\textit{Obs}^rec$ | $\textit{Obs}^comp$ | $\textit{Exp}^com$ | $\textit{Exp}^all$ |
| With $G$ | Zephyr 7B | 0.274 | 0.151 | 0.123 ${}_±0.200$ | 0.115 ${}_±0.166$ | 0.092 ${}_±0.108$ | 0.071 ${}_±0.139$ | 0.014 ${}_±0.037$ |
| Mistral 7B | 0.507 | 0.306 | 0.211 ${}_±0.190$ | 0.317 ${}_±0.253$ | 0.173 ${}_±0.157$ | 0.230 ${}_±0.312$ | 0.062 ${}_±0.088$ | |
| Mixtral 8 $×$ 7B | 0.413 | 0.237 | 0.147 ${}_±0.165$ | 0.266 ${}_±0.261$ | 0.124 ${}_±0.138$ | 0.144 ${}_±0.268$ | 0.029 ${}_±0.056$ | |
| LLama3 8B | 0.576 | 0.321 | 0.253 ${}_±0.156$ | 0.437 ${}_±0.207$ | 0.219 ${}_±0.137$ | 0.232 ${}_±0.316$ | 0.071 ${}_±0.093$ | |
| LLama3 70B | 0.752 | 0.540 | 0.277 ${}_±0.146$ | 0.537 ${}_±0.192$ | 0.256 ${}_±0.142$ | 0.395 ${}_±0.320$ | 0.112 ${}_±0.110$ | |
| GPT-3.5 turbo | 0.679 | 0.455 | 0.389 ${}_±0.212$ | 0.351 ${}_±0.192$ | 0.275 ${}_±0.167$ | 0.331 ${}_±0.366$ | 0.103 ${}_±0.127$ | |
| GPT-4 turbo | 0.772 | 0.572 | 0.446 ${}_±0.207$ | 0.491 ${}_±0.180$ | 0.371 ${}_±0.186$ | 0.475 ${}_±0.363$ | 0.199 ${}_±0.181$ | |
| With $K$ | LLama3 8B | 0.576 | 0.344 | 0.235 ${}_±0.162$ | 0.394 ${}_±0.227$ | 0.199 ${}_±0.142$ | 0.327 ${}_±0.375$ | 0.087 ${}_±0.114$ |
| LLama3 70B | 0.735 | 0.581 | 0.262 ${}_±0.146$ | 0.501 ${}_±0.208$ | 0.236 ${}_±0.131$ | 0.463 ${}_±0.374$ | 0.125 ${}_±0.117$ | |
| GPT-3.5 turbo | 0.652 | 0.413 | 0.347 ${}_±0.241$ | 0.279 ${}_±0.203$ | 0.232 ${}_±0.184$ | 0.374 ${}_±0.408$ | 0.121 ${}_±0.152$ | |
| GPT-4 turbo | 0.781 | 0.614 | 0.431 ${}_±0.207$ | 0.458 ${}_±0.187$ | 0.353 ${}_±0.170$ | 0.633 ${}_±0.338$ | 0.247 ${}_±0.201$ | |
Table 4: Evaluation of diagnostic reasoning ability of LLMs when no external knowledge is provided.
| | | | Observation | Explanation | | | |
| --- | --- | --- | --- | --- | --- | --- | --- |
| Task | Models | Acc ${}^diag$ | $\textit{Obs}^pre$ | $\textit{Obs}^rec$ | $\textit{Obs}^comp$ | $\textit{Exp}^com$ | $\textit{Exp}^all$ |
| With $D^⋆$ | LLama3 8B | 0.070 | 0.154 ${}_±0.139$ | 0.330 ${}_±0.244$ | 0.135 ${}_±0.122$ | 0.020 ${}_±0.100$ | 0.004 ${}_±0.016$ |
| LLama3 70B | 0.502 | 0.257 ${}_±0.150$ | 0.509 ${}_±0.213$ | 0.237 ${}_±0.145$ | 0.138 ${}_±0.209$ | 0.034 ${}_±0.054$ | |
| GPT-3.5 turbo | 0.223 | 0.164 ${}_±0.242$ | 0.149 ${}_±0.212$ | 0.116 ${}_±0.174$ | 0.091 ${}_±0.231$ | 0.025 ${}_±0.065$ | |
| GPT-4 turbo | 0.636 | 0.461 ${}_±0.206$ | 0.482 ${}_±0.160$ | 0.378 ${}_±0.174$ | 0.186 ${}_±0.221$ | 0.074 ${}_±0.090$ | |
| No Knowledge | LLama3 8B | 0.023 | 0.137 ${}_±0.159$ | 0.258 ${}_±0.274$ | 0.119 ${}_±0.141$ | 0.018 ${}_±0.083$ | 0.006 ${}_±0.026$ |
| LLama3 70B | 0.037 | 0.246 ${}_±0.148$ | 0.504 ${}_±0.222$ | 0.227 ${}_±0.148$ | 0.022 ${}_±0.093$ | 0.007 ${}_±0.030$ | |
| GPT-3.5 turbo | 0.059 | 0.161 ${}_±0.238$ | 0.148 ${}_±0.215$ | 0.113 ${}_±0.171$ | 0.036 ${}_±0.131$ | 0.011 ${}_±0.039$ | |
| GPT-4 turbo | 0.074 | 0.410 ${}_±0.208$ | 0.443 ${}_±0.191$ | 0.324 ${}_±0.182$ | 0.047 ${}_±0.143$ | 0.019 ${}_±0.058$ | |
With $G$ , we can see that GPT-4 achieves the best performance in most metrics, especially related to observations and explanations, surpassing LLama3 70B by a large margin. In terms of accuracy (in both category and diagnosis levels), LLama3 70B is comparable to GPT-4. LLama3 70B also has a higher $\textit{Obs}^rec$ but low $\textit{Obs}^pre$ and $\textit{Obs}^comp$ , which means that this model tends to extract many observations. Models with high diagnostic accuracy are not necessarily excel in finding essential information in long text (i.e., observations) and generating reasons (i.e., explanations).
When $K$ is given, all models show better diagnostic accuracy (except GPT-3.5) and explanations, while observations are slightly degraded. GPT-4 with $K$ enhances Acc ${}^diag$ , $\textit{Exp}^com$ , and $\textit{Exp}^all$ scores. This suggests that premises and supporting edges are beneficial for diagnosis and explanation. Lower observational performance may indicate that the models lack the ability to associate premises and text in $R$ , which are often superficially different though semantically consistent.
LLMs may undergo inherent challenges for evaluation when no external knowledge is supplied. They may have the knowledge to diagnose but cannot make consistent observations and explanations that our task expects through $K$ . To explore this, we evaluate two settings: (1) giving $D^⋆$ and (2) no knowledge is supplied to a model (shown in Table 4). The prompts used for this setup are detailed in the supplementary material. We do not evaluate the accuracy of disease category prediction as it is basically the same as Table 3. We can clearly see that with $D^⋆$ , GPT-4’s diagnostic and observational scores are comparable to those of the task with $K$ , though explanatory performance is much worse. Without any external knowledge, the diagnostic accuracy is also inferior. We understand this comparison is unfair, as the prompts differ. We intend to give a rough idea about the challenge without external knowledge. The deteriorated performance can be attributed to inconsistent wording of diagnosis names, which makes evaluation tough. High observational scores imply that observations in $R$ can be identified without relying on external knowledge. There can be some cues to spot them.
<details>
<summary>x5.png Details</summary>
### Visual Description
## Grouped Bar Chart: Model Performance Across Medical Specialties
### Overview
The image displays a series of five grouped bar charts arranged horizontally. Each chart compares the performance of three large language models (LLMs) across three evaluation metrics within a specific medical specialty. The overall purpose is to benchmark model capabilities in specialized medical domains.
### Components/Axes
* **Chart Type:** Grouped Bar Chart (5 panels).
* **Legend:** Located at the top center of the entire figure.
* **Green Bar:** "Acc" (Likely abbreviation for Accuracy).
* **Beige/Light Orange Bar:** "Comp" (Likely abbreviation for Comprehensiveness or Completeness).
* **Teal/Light Blue Bar:** "Faith" (Likely abbreviation for Faithfulness).
* **Y-Axis:** Common to all five panels. Labeled with numerical values from `0.0` to `1.0` in increments of `0.2`. This represents a normalized score or probability.
* **X-Axis (Per Panel):** Lists three models: `LLama3`, `GPT-3.5`, `GPT-4`.
* **Panel Titles (Bottom Labels):** Each panel is labeled with a medical specialty:
1. Cardiology
2. Gastroenterology
3. Neurology
4. Pulmonology
5. Endocrinology
### Detailed Analysis
**Panel 1: Cardiology**
* **LLama3:** Acc ≈ 0.42, Comp ≈ 0.26, Faith ≈ 0.12.
* **GPT-3.5:** Acc ≈ 0.44, Comp ≈ 0.28, Faith ≈ 0.11.
* **GPT-4:** Acc ≈ 0.45, Comp ≈ 0.36, Faith ≈ 0.18.
* **Trend:** Acc scores are similar and moderate. Comp and Faith scores are lower, with GPT-4 showing a notable increase in Comp.
**Panel 2: Gastroenterology**
* **LLama3:** Acc ≈ 0.63, Comp ≈ 0.17, Faith ≈ 0.07.
* **GPT-3.5:** Acc ≈ 0.42, Comp ≈ 0.25, Faith ≈ 0.06.
* **GPT-4:** Acc ≈ 0.58, Comp ≈ 0.29, Faith ≈ 0.15.
* **Trend:** LLama3 has the highest Acc but the lowest Comp and Faith. GPT-4 shows a balanced profile with the highest Faith in this panel.
**Panel 3: Neurology**
* **LLama3:** Acc ≈ 0.79, Comp ≈ 0.33, Faith ≈ 0.20.
* **GPT-3.5:** Acc ≈ 0.71, Comp ≈ 0.30, Faith ≈ 0.19.
* **GPT-4:** Acc ≈ 0.80, Comp ≈ 0.45, Faith ≈ 0.34.
* **Trend:** This specialty shows the highest overall Acc scores. GPT-4 leads in all three metrics, with a particularly strong Comp score.
**Panel 4: Pulmonology**
* **LLama3:** Acc ≈ 0.61, Comp ≈ 0.33, Faith ≈ 0.10.
* **GPT-3.5:** Acc ≈ 0.35, Comp ≈ 0.29, Faith ≈ 0.10.
* **GPT-4:** Acc ≈ 0.70, Comp ≈ 0.44, Faith ≈ 0.19.
* **Trend:** GPT-4 significantly outperforms the other models in Acc and Comp. GPT-3.5 shows a notable dip in Acc compared to other specialties.
**Panel 5: Endocrinology**
* **LLama3:** Acc ≈ 0.44, Comp ≈ 0.26, Faith ≈ 0.11.
* **GPT-3.5:** Acc ≈ 0.38, Comp ≈ 0.26, Faith ≈ 0.12.
* **GPT-4:** Acc ≈ 0.47, Comp ≈ 0.38, Faith ≈ 0.21.
* **Trend:** Performance is generally lower and more uniform across models compared to other specialties. GPT-4 maintains a slight lead.
### Key Observations
1. **Metric Hierarchy:** Across all models and specialties, the `Acc` (green) score is consistently the highest, followed by `Comp` (beige), with `Faith` (teal) being the lowest. This suggests a potential trade-off or difficulty in achieving high faithfulness.
2. **Model Performance:** `GPT-4` consistently achieves the highest or near-highest scores in all three metrics across every specialty. `LLama3` often shows strong `Acc` but weaker `Comp` and `Faith`. `GPT-3.5` performance is more variable.
3. **Specialty Variance:** `Neurology` appears to be the specialty where models achieve the highest overall scores, particularly in `Acc`. `Cardiology` and `Endocrinology` show lower, more clustered performance.
4. **Notable Outlier:** In `Pulmonology`, `GPT-3.5`'s `Acc` score (~0.35) is significantly lower than its performance in other specialties and lower than both `LLama3` and `GPT-4` in the same panel.
### Interpretation
The data suggests a clear performance gradient among the evaluated LLMs in specialized medical question-answering or reasoning tasks, with `GPT-4` demonstrating superior capability. The consistent pattern of `Acc > Comp > Faith` indicates that while models can often arrive at correct answers (`Acc`), providing comprehensive (`Comp`) and, especially, faithful (`Faith`) justifications or information is more challenging. This has significant implications for clinical applications where explainability and reliability of the reasoning process are critical.
The variation across specialties implies that model knowledge or reasoning ability is not uniform across medicine. The high scores in Neurology might reflect a larger or more structured training corpus for that domain, while lower scores in Endocrinology could indicate a more complex or less represented knowledge base. The dip for `GPT-3.5` in Pulmonology warrants further investigation into potential dataset biases or model limitations for that specific domain. Overall, the chart provides a multi-faceted benchmark showing that model selection for medical AI should consider both the target specialty and the required balance between accuracy, comprehensiveness, and faithfulness.
</details>
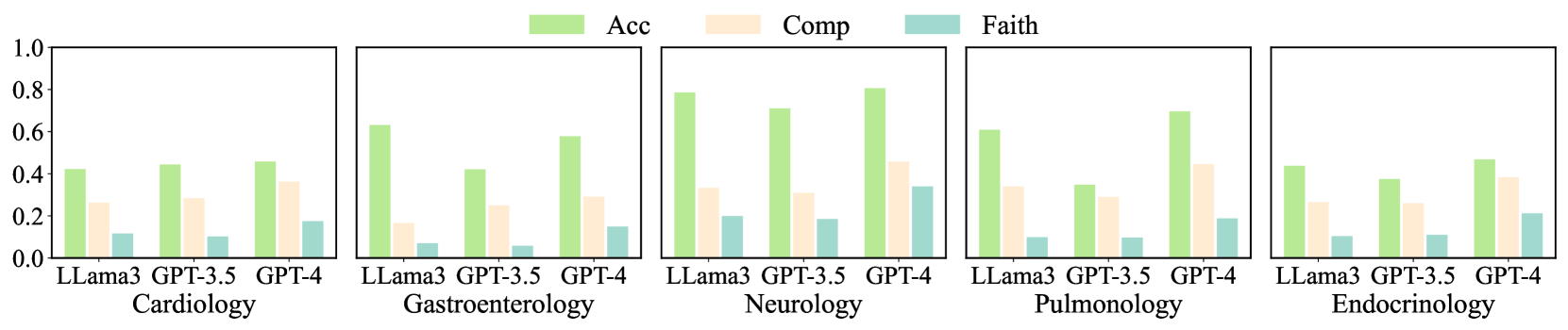
Figure 5: Performance of LLama3 70B, GPT-3.5, and GPT-4 under different medical domains. We use the task with $G$ .
Performance in individual domains. Figure 5 summarizes the performance of LLama3 70B, GPT-3.5, and GPT-4 across different medical domains, evaluated using Acc ${}^cat$ , $\textit{Obs}^comp$ , and $\textit{Exp}^all$ . Neurology gives the best diagnostic accuracy, where GPT-4 achieved an accuracy of 0.806. LLama3 also performed well (0.786). In terms of $\textit{Obs}^comp$ and $\textit{Exp}^all$ , GPT-4’s results were 0.458 and 0.340, respectively, with the smallest difference between the two scores among all domains. This smaller gap indicates that in Neurology, the common observations in prediction and ground truth lead to the correct diagnoses with faithful rationalizations. However, GPT-4 yields a higher diagnostic accuracy score while a lower explanatory score, suggesting that the observations captured by the model or their rationalizations differ from human doctors.
For Cardiology and Endocrinology, the diagnostic accuracy of the models is relatively low (GPT-4 achieved 0.458 and 0.468, respectively). Nevertheless, $\textit{Obs}^comp$ and $\textit{Exp}^all$ are relatively high. Endocrinology results in lower diagnostic accuracy and higher explanatory performance. A smaller gap may imply that in these two domains, successful predictions are associated with observations similar to those of human doctors, and the reasoning process may be analogous. Conversely, in Gastroenterology, higher Acc ${}^cat$ ) is accompanied by lower $\textit{Obs}^comp$ and $\textit{Exp}^all$ (especially for LLama3), potentially indicating a significant divergence in the reasoning process from human doctors. Overall, DiReCT demonstrates that the degree of alignment between the model’s diagnostic reasoning ability and that of human doctors varies across different medical domains.
Table 5: Consistency of automated evaluation metrics with human judgments.
| Model | Observation | Rationalization |
| --- | --- | --- |
| LLama3 8B | 0.887 | 0.801 |
| GPT-4 turbo | 0.902 | 0.836 |
Reliability of automatic evaluation. We randomly pick out 100 samples from DiReCT and their prediction by GPT-4 over the task with $G$ to assess the consistency of our automated metrics to evaluate the observational and explanatory performance in Section 3.3 to human judgments. Three physicians joined this experiment. For each prediction $\hat{o}∈\hat{O}$ , they are asked to find a similar observation in ground truth $O$ . For explanatory metrics, they verify if each prediction $\hat{z}∈\hat{E}$ for $\hat{o}∈\hat{O}$ align with ground-truth $z∈E$ corresponding to $o$ . A prediction and a ground truth are deemed aligned for both assessments if at least two specialists agree. We compare LLama3’s and GPT-4’s judgments to explore if there is a gap between these LLMs. As summarized in Table 5, GPT-4 achieves the best results, with LLama3 8B also displaying a similar performance. From these results, we argue that our automated evaluation metrics are consistent with human judgments, and LLama3 is sufficient for this evaluation, allowing the cost-efficient option.
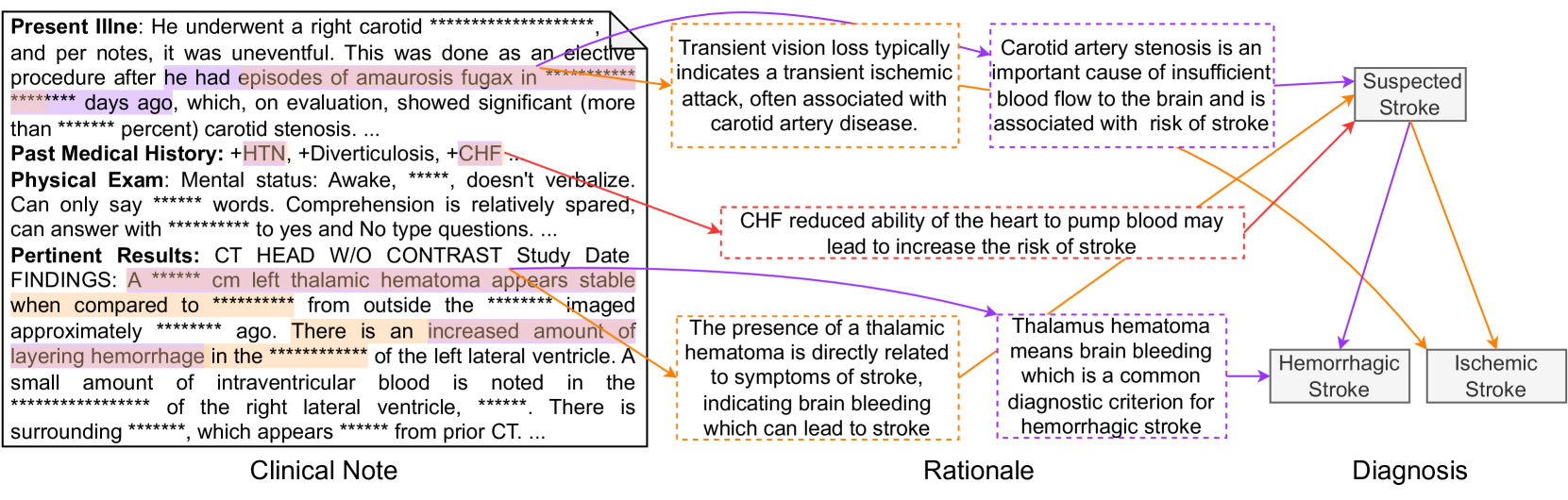
A prediction example. Figure 6 shows a sample generated by GPT-4. The ground-truth PDD of the input clinical note is Hemorrhagic Stroke. In this figure, purple, orange, and red indicate explanations only in the ground truth, only in prediction, and common in both, respectively; therefore, red is a successful prediction of an explanation, while purple and orange are a false negative and false positive. GPT-4 treats the observation of aurosis fugax as the criteria for diagnosing Ischemic Stroke. However, this observation only supports Suspected Stroke. Conversely, observation thalamic hematoma, which is the key indicator of Hemorrhagic Stroke, is regarded as a less important clue. Such observation-diagnosis correspondence errors lead to the model’s misdiagnosis. More samples are available in the supplementary material.
<details>
<summary>x6.png Details</summary>
### Visual Description
## Medical Case Analysis Diagram: Clinical Reasoning for Stroke Diagnosis
### Overview
The image is a structured medical reasoning diagram that traces a patient's clinical presentation from a clinical note through a rationale phase to a final diagnostic conclusion. It visually maps how specific findings in a patient's record lead to the suspicion and classification of a stroke. The diagram is divided into three vertical sections: **Clinical Note** (left), **Rationale** (center), and **Diagnosis** (right). Colored arrows connect specific text excerpts to explanatory rationales and then to diagnostic outcomes.
### Components/Axes
The diagram has three primary components arranged left to right:
1. **Clinical Note (Left Panel):** A text box containing excerpts from a patient's medical record. Sections include:
* **Present Illness:** Describes a right carotid procedure and past episodes of *amaurosis fugax* (transient vision loss) due to significant carotid stenosis.
* **Past Medical History:** Lists comorbidities: +HTN (Hypertension), +Diverticulosis, +CHF (Congestive Heart Failure).
* **Physical Exam:** Notes mental status (awake, non-verbal, limited comprehension).
* **Pertinent Results:** Details from a CT HEAD W/O CONTRAST scan, noting a stable left thalamic hematoma and increased layering hemorrhage in the left lateral ventricle.
* *Note: Several specific values (percentages, measurements, dates) are redacted with asterisks (`*`).*
2. **Rationale (Center Panel):** A series of text boxes with dashed borders, each providing a medical explanation linking a clinical finding to a stroke mechanism. These are connected by colored arrows to the source text and to the diagnoses.
* **Orange Dashed Box (Top):** "Transient vision loss typically indicates a transient ischemic attack, often associated with carotid artery disease."
* **Purple Dashed Box (Top):** "Carotid artery stenosis is an important cause of insufficient blood flow to the brain and is associated with risk of stroke."
* **Red Dashed Box (Middle):** "CHF reduced ability of the heart to pump blood may lead to increase the risk of stroke."
* **Orange Dashed Box (Bottom):** "The presence of a thalamic hematoma is directly related to symptoms of stroke, indicating brain bleeding which can lead to stroke."
* **Purple Dashed Box (Bottom):** "Thalamus hematoma means brain bleeding which is a common diagnostic criterion for hemorrhagic stroke."
3. **Diagnosis (Right Panel):** Three solid-bordered boxes representing diagnostic conclusions.
* **Suspected Stroke** (Top)
* **Hemorrhagic Stroke** (Bottom Left)
* **Ischemic Stroke** (Bottom Right)
### Detailed Analysis
**Textual Extraction from Clinical Note:**
* **Present Illness:** "He underwent a right carotid ***************** and per notes, it was uneventful. This was done as an elective procedure after he had episodes of amaurosis fugax in ******** days ago, which, on evaluation, showed significant (more than ****** percent) carotid stenosis. ..."
* **Past Medical History:** "+HTN, +Diverticulosis, +CHF."
* **Physical Exam:** "Mental status: Awake, ****, doesn't verbalize. Can only say ****** words. Comprehension is relatively spared, can answer with ********** to yes and No type questions. ..."
* **Pertinent Results:** "CT HEAD W/O CONTRAST Study Date FINDINGS: A ****** cm left thalamic hematoma appears stable when compared to ********** from outside the ****** imaged approximately ****** ago. There is an increased amount of layering hemorrhage in the *********** of the left lateral ventricle. A small amount of intraventricular blood is noted in the *************** of the right lateral ventricle, ******. There is surrounding *******, which appears ****** from prior CT. ..."
**Arrow Mapping & Logic Flow:**
* **Orange Arrows:** Trace from "amaurosis fugax" and "thalamic hematoma" text to the corresponding orange rationale boxes. These rationales then point to **Ischemic Stroke** (from carotid disease) and **Hemorrhagic Stroke** (from brain bleeding).
* **Purple Arrows:** Trace from "carotid stenosis" and "thalamic hematoma" text to the corresponding purple rationale boxes. These point to **Suspected Stroke** (general risk) and **Hemorrhagic Stroke** (specific criterion).
* **Red Arrow:** Traces from "+CHF" to the red rationale box, which points directly to **Suspected Stroke**.
### Key Observations
1. **Dual Pathways to Diagnosis:** The diagram shows two distinct etiological pathways leading to stroke diagnosis:
* An **ischemic pathway** (orange) originating from carotid artery disease (amaurosis fugax, stenosis).
* A **hemorrhagic pathway** (purple/orange) originating from direct evidence of brain bleeding (thalamic hematoma, intraventricular hemorrhage).
2. **CHF as a General Risk Factor:** Congestive Heart Failure (CHF) is presented not as a direct cause of a specific stroke type, but as a general condition that increases overall stroke risk, leading to the "Suspected Stroke" conclusion.
3. **Anatomical Specificity:** The clinical note specifies a **left thalamic hematoma** and hemorrhage in the **left lateral ventricle**, providing precise anatomical localization for the hemorrhagic event.
4. **Temporal Information:** The note indicates the hematoma is "stable" compared to a prior scan, suggesting an ongoing but not acutely expanding condition at the time of the note.
### Interpretation
This diagram is a visual representation of **clinical diagnostic reasoning**. It demonstrates how a clinician synthesizes disparate pieces of information from a patient's record to arrive at a diagnosis.
* **What the data suggests:** The patient presents with a complex picture involving both risk factors for ischemic stroke (carotid disease, CHF) and definitive evidence of hemorrhagic stroke (thalamic hematoma). The diagram logically concludes that a **stroke is highly suspected**, and further classifies it as having a **hemorrhagic component** based on imaging, while acknowledging an **ischemic risk profile**.
* **How elements relate:** The "Rationale" section acts as the critical bridge, translating raw clinical data (symptoms, history, imaging) into pathophysiological mechanisms. The color-coded arrows create a traceable audit trail from evidence to conclusion.
* **Notable patterns:** The most significant pattern is the **co-occurrence of ischemic risk factors and hemorrhagic evidence**. This is not uncommon in clinical practice, where patients may have multiple comorbidities. The diagram effectively shows that a single diagnosis ("Stroke") can have multiple contributing factors and subtypes. The redaction of specific values (stenosis percentage, hematoma size) highlights that the diagnostic logic often relies on the *presence* and *qualitative description* of findings rather than precise numbers alone.
</details>
Figure 6: An example prediction for a clinical note with PDD of Hemorrhagic Stroke by GPT-4.
## 6 Conclusion and Limitations
We proposed DiReCT as the first benchmark for evaluating the diagnostic reasoning ability of LLMs with interpretability by supplying external knowledge as a graph. Our evaluations reveal a notable disparity between current leading-edge LLMs and human experts, underscoring the urgent need for AI models that can perform reliable and interpretable reasoning in clinical environments. DiReCT can be easily extended to more challenging settings by removing the knowledge graph from the input, facilitating evaluations of future LLMs.
Limitations. DiReCT encompasses only a subset of disease categories and considers only one PDD, omitting the inter-diagnostic relationships due to their complexity—a significant challenge even for human doctors. Additionally, our baseline may not use optimal prompts, chain-of-thought reasoning, or address issues related to hallucinations in task responses. Our dataset is solely intended for model evaluation but not for use in clinical environments. The use of the diagnostic knowledge graph is also limited to serving merely as a part of input. Future work will focus on constructing a more comprehensive disease dataset and developing an extensive diagnostic knowledge graph.
## Acknowledgments and Disclosure of Funding
This work was supported by World Premier International Research Center Initiative (WPI), MEXT, Japan. This work is also supported by JSPS KAKENHI 24K20795 and Dalian Haichuang Project for Advanced Talents.
## References
- Zhao et al. [2023] Wayne Xin Zhao, Kun Zhou, Junyi Li, Tianyi Tang, Xiaolei Wang, Yupeng Hou, Yingqian Min, Beichen Zhang, Junjie Zhang, Zican Dong, Yifan Du, Chen Yang, Yushuo Chen, Zhipeng Chen, Jinhao Jiang, Ruiyang Ren, Yifan Li, Xinyu Tang, Zikang Liu, Peiyu Liu, Jian-Yun Nie, and Ji-Rong Wen. A survey of large language models. arXiv preprint arXiv:2303.18223, 2023.
- Min et al. [2023] Bonan Min, Hayley Ross, Elior Sulem, Amir Pouran Ben Veyseh, Thien Huu Nguyen, Oscar Sainz, Eneko Agirre, Ilana Heintz, and Dan Roth. Recent advances in natural language processing via large pre-trained language models: A survey. ACM Computing Surveys, 56(2):1–40, 2023.
- Anil et al. [2023] Rohan Anil, Andrew M Dai, Orhan Firat, Melvin Johnson, Dmitry Lepikhin, Alexandre Passos, Siamak Shakeri, Emanuel Taropa, Paige Bailey, Zhifeng Chen, et al. Palm 2 technical report. arXiv preprint arXiv:2305.10403, 2023.
- Han et al. [2023] Tianyu Han, Lisa C Adams, Jens-Michalis Papaioannou, Paul Grundmann, Tom Oberhauser, Alexander Löser, Daniel Truhn, and Keno K Bressem. Medalpaca–an open-source collection of medical conversational ai models and training data. arXiv preprint arXiv:2304.08247, 2023.
- Jin et al. [2021] Di Jin, Eileen Pan, Nassim Oufattole, Wei-Hung Weng, Hanyi Fang, and Peter Szolovits. What disease does this patient have? a large-scale open domain question answering dataset from medical exams. Applied Sciences, 11(14):6421, 2021.
- OpenAI [2023a] OpenAI. GPT-4 Technical Report. CoRR, abs/2303.08774, 2023a. doi: 10.48550/arXiv.2303.08774. URL https://doi.org/10.48550/arXiv.2303.08774.
- Bubeck et al. [2023] Sébastien Bubeck, Varun Chandrasekaran, Ronen Eldan, Johannes Gehrke, Eric Horvitz, Ece Kamar, Peter Lee, Yin Tat Lee, Yuanzhi Li, Scott Lundberg, et al. Sparks of artificial general intelligence: Early experiments with gpt-4. arXiv preprint arXiv:2303.12712, 2023.
- Nori et al. [2023] Harsha Nori, Nicholas King, Scott Mayer McKinney, Dean Carignan, and Eric Horvitz. Capabilities of gpt-4 on medical challenge problems. arXiv preprint arXiv:2303.13375, 2023.
- Liévin et al. [2024] Valentin Liévin, Christoffer Egeberg Hother, Andreas Geert Motzfeldt, and Ole Winther. Can large language models reason about medical questions? Patterns, 5(3), 2024.
- Pal et al. [2022] Ankit Pal, Logesh Kumar Umapathi, and Malaikannan Sankarasubbu. MedMCQA: A large-scale multi-subject multi-choice dataset for medical domain question answering. In Conference on health, inference, and learning, pages 248–260. PMLR, 2022.
- Li et al. [2023] Dongfang Li, Jindi Yu, Baotian Hu, Zhenran Xu, and Min Zhang. ExplainCPE: A free-text explanation benchmark of chinese pharmacist examination. arXiv preprint arXiv:2305.12945, 2023.
- Chen et al. [2024] Hanjie Chen, Zhouxiang Fang, Yash Singla, and Mark Dredze. Benchmarking large language models on answering and explaining challenging medical questions. arXiv preprint arXiv:2402.18060, 2024.
- Jullien et al. [2023] Mael Jullien, Marco Valentino, Hannah Frost, Paul O’Regan, Donal Landers, and André Freitas. Semeval-2023 task 7: Multi-evidence natural language inference for clinical trial data. arXiv preprint arXiv:2305.02993, 2023.
- Gao et al. [2023a] Yanjun Gao, Ruizhe Li, John Caskey, Dmitriy Dligach, Timothy Miller, Matthew M Churpek, and Majid Afshar. Leveraging a medical knowledge graph into large language models for diagnosis prediction. arXiv preprint arXiv:2308.14321, 2023a.
- Johnson et al. [2023] Alistair E. W. Johnson, Lucas Bulgarelli, Lu Shen, Alvin Gayles, Ayad Shammout, Steven Horng, Tom J. Pollard, Sicheng Hao, Benjamin Moody, Brian Gow, Li-wei H. Lehman, Leo A. Celi, and Roger G. Mark. MIMIC-IV, a freely accessible electronic health record dataset. Scientific data, 10(1):1, 2023.
- Xi et al. [2023] Zhiheng Xi, Wenxiang Chen, Xin Guo, Wei He, Yiwen Ding, Boyang Hong, Ming Zhang, Junzhe Wang, Senjie Jin, Enyu Zhou, Rui Zheng, Xiaoran Fan, Xiao Wang, Limao Xiong, Yuhao Zhou, Weiran Wang, Changhao Jiang, Yicheng Zou, Xiangyang Liu, Zhangyue Yin, Shihan Dou, Rongxiang Weng, Wensen Cheng, Qi Zhang, Wenjuan Qin, Yongyan Zheng, Xipeng Qiu, Xuanjing Huang, and Tao Gui. The rise and potential of large language model based agents: A survey, 2023.
- Tang et al. [2023] Xiangru Tang, Anni Zou, Zhuosheng Zhang, Yilun Zhao, Xingyao Zhang, Arman Cohan, and Mark Gerstein. Medagents: Large language models as collaborators for zero-shot medical reasoning. arXiv preprint arXiv:2311.10537, 2023.
- Middleton et al. [2013] Blackford Middleton, Meryl Bloomrosen, Mark A Dente, Bill Hashmat, Ross Koppel, J Marc Overhage, Thomas H Payne, S Trent Rosenbloom, Charlotte Weaver, and Jiajie Zhang. Enhancing patient safety and quality of care by improving the usability of electronic health record systems: recommendations from amia. Journal of the American Medical Informatics Association, 20(e1):e2–e8, 2013.
- Liu et al. [2022] Jinghui Liu, Daniel Capurro, Anthony Nguyen, and Karin Verspoor. “note bloat” impacts deep learning-based nlp models for clinical prediction tasks. Journal of biomedical informatics, 133:104149, 2022.
- Danilevsky et al. [2020] Marina Danilevsky, Kun Qian, Ranit Aharonov, Yannis Katsis, Ban Kawas, and Prithviraj Sen. A survey of the state of explainable AI for natural language processing. In Proceedings of the 1st Conference of the Asia-Pacific Chapter of the Association for Computational Linguistics and the 10th International Joint Conference on Natural Language Processing, pages 447–459, 2020.
- Gurrapu et al. [2023] Sai Gurrapu, Ajay Kulkarni, Lifu Huang, Ismini Lourentzou, and Feras A Batarseh. Rationalization for explainable nlp: A survey. Frontiers in Artificial Intelligence, 6, 2023.
- Camburu et al. [2018] Oana-Maria Camburu, Tim Rocktäschel, Thomas Lukasiewicz, and Phil Blunsom. e-snli: Natural language inference with natural language explanations. Advances in Neural Information Processing Systems, 31, 2018.
- Rajani et al. [2019] Nazneen Fatema Rajani, Bryan McCann, Caiming Xiong, and Richard Socher. Explain yourself! leveraging language models for commonsense reasoning. In Proceedings of the 57th Annual Meeting of the Association for Computational Linguistics, pages 4932–4942, Florence, Italy, 2019.
- DeYoung et al. [2020] Jay DeYoung, Sarthak Jain, Nazneen Fatema Rajani, Eric Lehman, Caiming Xiong, Richard Socher, and Byron C. Wallace. ERASER: A benchmark to evaluate rationalized NLP models. In Proceedings of the 58th Annual Meeting of the Association for Computational Linguistics, pages 4443–4458, 2020.
- Jhamtani and Clark [2020] Harsh Jhamtani and Peter Clark. Learning to explain: Datasets and models for identifying valid reasoning chains in multihop question-answering. In Proceedings of the 2020 Conference on Empirical Methods in Natural Language Processing, page 137–150, 2020.
- Tafjord et al. [2021] Oyvind Tafjord, Bhavana Dalvi Mishra, and Peter Clark. Proofwriter: Generating implications, proofs, and abductive statements over natural language. In Findings of the Association for Computational Linguistics: ACL-IJCNLP, page 3621–3634, 2021.
- Zhao et al. [2021] Chen Zhao, Chenyan Xiong, Jordan Boyd-Graber, and Hal Daumé III. Multi-step reasoning over unstructured text with beam dense retrieval. In Proceedings of the 2021 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, pages 4635–4641, 2021.
- Dalvi et al. [2021] Bhavana Dalvi, Peter Jansen, Oyvind Tafjord, Zhengnan Xie, Hannah Smith, Leighanna Pipatanangkura, and Peter Clark. Explaining answers with entailment trees. In Proceedings of the 2021 Conference on Empirical Methods in Natural Language Processing, pages 7358–7370, 2021.
- Zhang et al. [2024] Yifan Zhang, Jingqin Yang, Yang Yuan, and Andrew Chi-Chih Yao. Cumulative reasoning with large language models. In ICLR 2024 Workshop on Bridging the Gap Between Practice and Theory in Deep Learning, 2024. URL https://openreview.net/forum?id=XAAYyRxTlQ.
- Gao et al. [2022] Yanjun Gao, Dmitriy Dligach, Timothy Miller, Samuel Tesch, Ryan Laffin, Matthew M. Churpek, and Majid Afshar. Hierarchical annotation for building a suite of clinical natural language processing tasks: Progress note understanding. In Proceedings of the Thirteenth Language Resources and Evaluation Conference, pages 5484–5493, Marseille, France, 2022. European Language Resources Association.
- Zack et al. [2023] Travis Zack, Gurpreet Dhaliwal, Rabih Geha, Mary Margaretten, Sara Murray, and Julian C Hong. A clinical reasoning-encoded case library developed through natural language processing. Journal of General Internal Medicine, 38(1):5–11, 2023.
- Gao et al. [2023b] Yanjun Gao, Dmitriy Dligach, Timothy Miller, John Caskey, Brihat Sharma, Matthew M Churpek, and Majid Afshar. Dr. bench: Diagnostic reasoning benchmark for clinical natural language processing. Journal of Biomedical Informatics, 138:104286, 2023b.
- Weed [1970] L.L. Weed. Medical Records, Medical Education, and Patient Care: The Problem-oriented Record as a Basic Tool. Press of Case Western Reserve University, 1970. ISBN 9780815191889.
- Bodenreider [2004] Olivier Bodenreider. The unified medical language system (umls): integrating biomedical terminology. Nucleic acids research, 32(suppl_1):D267–D270, 2004.
- AI@Meta [2024] AI@Meta. Llama 3 model card. 2024. URL https://github.com/meta-llama/llama3/blob/main/MODEL_CARD.md.
- Tunstall et al. [2023] Lewis Tunstall, Edward Beeching, Nathan Lambert, Nazneen Rajani, Kashif Rasul, Younes Belkada, Shengyi Huang, Leandro von Werra, Clémentine Fourrier, Nathan Habib, et al. Zephyr: Direct distillation of lm alignment. arXiv preprint arXiv:2310.16944, 2023.
- Jiang et al. [2023] Albert Q Jiang, Alexandre Sablayrolles, Arthur Mensch, Chris Bamford, Devendra Singh Chaplot, Diego de las Casas, Florian Bressand, Gianna Lengyel, Guillaume Lample, Lucile Saulnier, et al. Mistral 7b. arXiv preprint arXiv:2310.06825, 2023.
- OpenAI [2023b] OpenAI. Introducing ChatGPT and Whisper APIs. 2023b. URL https://openai.com/blog/introducing-chatgpt-and-whisper-apis.
- Byrne et al. [2024] Robert A Byrne, Xavier Rossello, JJ Coughlan, Emanuele Barbato, Colin Berry, Alaide Chieffo, Marc J Claeys, Gheorghe-Andrei Dan, Marc R Dweck, Mary Galbraith, et al. 2023 esc guidelines for the management of acute coronary syndromes: developed by the task force on the management of acute coronary syndromes of the european society of cardiology (esc). European Heart Journal: Acute Cardiovascular Care, 13(1):55–161, 2024.
- Members et al. [2022] Writing Committee Members, Eric M Isselbacher, Ourania Preventza, James Hamilton Black III, John G Augoustides, Adam W Beck, Michael A Bolen, Alan C Braverman, Bruce E Bray, Maya M Brown-Zimmerman, et al. 2022 acc/aha guideline for the diagnosis and management of aortic disease: a report of the american heart association/american college of cardiology joint committee on clinical practice guidelines. Journal of the American College of Cardiology, 80(24):e223–e393, 2022.
- Joglar et al. [2024] José A Joglar, Mina K Chung, Anastasia L Armbruster, Emelia J Benjamin, Janice Y Chyou, Edmond M Cronin, Anita Deswal, Lee L Eckhardt, Zachary D Goldberger, Rakesh Gopinathannair, et al. 2023 acc/aha/accp/hrs guideline for the diagnosis and management of atrial fibrillation: a report of the american college of cardiology/american heart association joint committee on clinical practice guidelines. Circulation, 149(1):e1–e156, 2024.
- Ommen et al. [2020] Steve R Ommen, Seema Mital, Michael A Burke, Sharlene M Day, Anita Deswal, Perry Elliott, Lauren L Evanovich, Judy Hung, José A Joglar, Paul Kantor, et al. 2020 aha/acc guideline for the diagnosis and treatment of patients with hypertrophic cardiomyopathy: executive summary: a report of the american college of cardiology/american heart association joint committee on clinical practice guidelines. Journal of the American College of Cardiology, 76(25):3022–3055, 2020.
- Heidenreich et al. [2022] Paul A Heidenreich, Biykem Bozkurt, David Aguilar, Larry A Allen, Joni J Byun, Monica M Colvin, Anita Deswal, Mark H Drazner, Shannon M Dunlay, Linda R Evers, et al. 2022 aha/acc/hfsa guideline for the management of heart failure: a report of the american college of cardiology/american heart association joint committee on clinical practice guidelines. Journal of the American College of Cardiology, 79(17):e263–e421, 2022.
- Su et al. [2021] Lilly Su, Rea Mittal, Devyani Ramgobin, Rahul Jain, and Rohit Jain. Current management guidelines on hyperlipidemia: the silent killer. Journal of lipids, 2021(1):9883352, 2021.
- Unger et al. [2020] Thomas Unger, Claudio Borghi, Fadi Charchar, Nadia A Khan, Neil R Poulter, Dorairaj Prabhakaran, Agustin Ramirez, Markus Schlaich, George S Stergiou, Maciej Tomaszewski, et al. 2020 international society of hypertension global hypertension practice guidelines. Hypertension, 75(6):1334–1357, 2020.
- Shah et al. [2021] Shailja C Shah, M Blanca Piazuelo, Ernst J Kuipers, and Dan Li. Aga clinical practice update on the diagnosis and management of atrophic gastritis: expert review. Gastroenterology, 161(4):1325–1332, 2021.
- Gyawali et al. [2024] C Prakash Gyawali, Rena Yadlapati, Ronnie Fass, David Katzka, John Pandolfino, Edoardo Savarino, Daniel Sifrim, Stuart Spechler, Frank Zerbib, Mark R Fox, et al. Updates to the modern diagnosis of gerd: Lyon consensus 2.0. Gut, 73(2):361–371, 2024.
- Kavitt et al. [2019] Robert T Kavitt, Anna M Lipowska, Adjoa Anyane-Yeboa, and Ian M Gralnek. Diagnosis and treatment of peptic ulcer disease. The American journal of medicine, 132(4):447–456, 2019.
- Barkun et al. [2019] Alan N Barkun, Majid Almadi, Ernst J Kuipers, Loren Laine, Joseph Sung, Frances Tse, Grigorios I Leontiadis, Neena S Abraham, Xavier Calvet, Francis KL Chan, et al. Management of nonvariceal upper gastrointestinal bleeding: guideline recommendations from the international consensus group. Annals of internal medicine, 171(11):805–822, 2019.
- McKhann et al. [1984] Guy McKhann, David Drachman, Marshall Folstein, Robert Katzman, Donald Price, and Emanuel M Stadlan. Clinical diagnosis of alzheimer’s disease: Report of the nincds-adrda work group* under the auspices of department of health and human services task force on alzheimer’s disease. Neurology, 34(7):939–939, 1984.
- Igaku-Shoin-Ltd. [2018] Igaku-Shoin-Ltd. Clinical practice guidelines for epilepsy 2018. 2018.
- Lipton et al. [2001] Richard B Lipton, Seymour Diamond, Michael Reed, Merle L Diamond, and Walter F Stewart. Migraine diagnosis and treatment: results from the american migraine study ii. Headache: The Journal of Head and Face Pain, 41(7):638–645, 2001.
- Lublin [2005] Fred D Lublin. Clinical features and diagnosis of multiple sclerosis. Neurologic clinics, 23(1):1–15, 2005.
- Kleindorfer et al. [2021] Dawn O Kleindorfer, Amytis Towfighi, Seemant Chaturvedi, Kevin M Cockroft, Jose Gutierrez, Debbie Lombardi-Hill, Hooman Kamel, Walter N Kernan, Steven J Kittner, Enrique C Leira, et al. 2021 guideline for the prevention of stroke in patients with stroke and transient ischemic attack: a guideline from the american heart association/american stroke association. Stroke, 52(7):e364–e467, 2021.
- Qaseem et al. [2011] Amir Qaseem, Timothy J Wilt, Steven E Weinberger, Nicola A Hanania, Gerard Criner, Thys van der Molen, Darcy D Marciniuk, Tom Denberg, Holger Schünemann, Wisia Wedzicha, et al. Diagnosis and management of stable chronic obstructive pulmonary disease: a clinical practice guideline update from the american college of physicians, american college of chest physicians, american thoracic society, and european respiratory society. Annals of internal medicine, 155(3):179–191, 2011.
- Gupta et al. [2013] Dheeraj Gupta, Ritesh Agarwal, Ashutosh Nath Aggarwal, VN Maturu, Sahajal Dhooria, KT Prasad, Inderpaul S Sehgal, Lakshmikant B Yenge, Aditya Jindal, Navneet Singh, et al. Guidelines for diagnosis and management of chronic obstructive pulmonary disease: Joint ics/nccp (i) recommendations. Lung India, 30(3):228–267, 2013.
- Olson and Davis [2020] Gregory Olson and Andrew M Davis. Diagnosis and treatment of adults with community-acquired pneumonia. Jama, 323(9):885–886, 2020.
- Konstantinides et al. [2020] Stavros V Konstantinides, Guy Meyer, Cecilia Becattini, Héctor Bueno, Geert-Jan Geersing, Veli-Pekka Harjola, Menno V Huisman, Marc Humbert, Catriona Sian Jennings, David Jiménez, et al. 2019 esc guidelines for the diagnosis and management of acute pulmonary embolism developed in collaboration with the european respiratory society (ers) the task force for the diagnosis and management of acute pulmonary embolism of the european society of cardiology (esc). European heart journal, 41(4):543–603, 2020.
- Lewinsohn et al. [2017] David M Lewinsohn, Michael K Leonard, Philip A LoBue, David L Cohn, Charles L Daley, Ed Desmond, Joseph Keane, Deborah A Lewinsohn, Ann M Loeffler, Gerald H Mazurek, et al. Official american thoracic society/infectious diseases society of america/centers for disease control and prevention clinical practice guidelines: diagnosis of tuberculosis in adults and children. Clinical Infectious Diseases, 64(2):e1–e33, 2017.
- Charmandari et al. [2014] Evangelia Charmandari, Nicolas C Nicolaides, and George P Chrousos. Adrenal insufficiency. The Lancet, 383(9935):2152–2167, 2014.
- ElSayed et al. [2023] Nuha A ElSayed, Grazia Aleppo, Vanita R Aroda, Raveendhara R Bannuru, Florence M Brown, Dennis Bruemmer, Billy S Collins, Kenneth Cusi, Marisa E Hilliard, Diana Isaacs, et al. 4. comprehensive medical evaluation and assessment of comorbidities: Standards of care in diabetes—2023. Diabetes Care, 46(Suppl 1):s49, 2023.
- Tritos and Miller [2023] Nicholas A Tritos and Karen K Miller. Diagnosis and management of pituitary adenomas: a review. Jama, 329(16):1386–1398, 2023.
- AlexanderErik et al. [2017] K AlexanderErik, N PearceElizabeth, A BrentGregory, S BrownRosalind, A GrobmanWilliam, H LazarusJohn, J MandelSusan, P PeetersRobin, et al. 2017 guidelines of the american thyroid association for the diagnosis and management of thyroid disease during pregnancy and the postpartum. Thyroid, 2017.
## Appendix A Details of DiReCT
### A.1 Data Statistics
Table 6: Disease statistics of DiReCT.
| Domains | Categories | # samples | $|D_i|$ | $|D^⋆_i|$ | References |
| --- | --- | --- | --- | --- | --- |
| Cardiology | Acute Coronary Syndromes | 65 | 6 | 3 | [Byrne et al., 2024] |
| Aortic Dissection | 14 | 3 | 2 | [Members et al., 2022] | |
| Atrial Fibrillation | 10 | 3 | 2 | [Joglar et al., 2024] | |
| Cardiomyopathy | 9 | 5 | 4 | [Ommen et al., 2020] | |
| Heart Failure | 52 | 6 | 3 | [Heidenreich et al., 2022] | |
| Hyperlipidemia | 2 | 2 | 1 | [Su et al., 2021] | |
| Hypertension | 32 | 2 | 1 | [Unger et al., 2020] | |
| Gastroenterology | Gastritis | 27 | 5 | 3 | [Shah et al., 2021] |
| Gastroesophageal Reflux Disease | 41 | 2 | 1 | [Gyawali et al., 2024] | |
| Peptic Ulcer Disease | 28 | 3 | 2 | [Kavitt et al., 2019] | |
| Upper Gastrointestinal Bleeding | 7 | 2 | 1 | [Barkun et al., 2019] | |
| Neurology | Alzheimer | 10 | 2 | 1 | [McKhann et al., 1984] |
| Epilepsy | 8 | 3 | 2 | [Igaku-Shoin-Ltd., 2018] | |
| Migraine | 4 | 3 | 2 | [Lipton et al., 2001] | |
| Multiple Sclerosis | 27 | 6 | 4 | [Lublin, 2005] | |
| Stroke | 28 | 3 | 2 | [Kleindorfer et al., 2021] | |
| Pulmonology | Asthma | 13 | 7 | 5 | [Qaseem et al., 2011] |
| COPD | 19 | 6 | 4 | [Gupta et al., 2013] | |
| Pneumonia | 20 | 4 | 2 | [Olson and Davis, 2020] | |
| Pulmonary Embolism | 35 | 5 | 3 | [Konstantinides et al., 2020] | |
| Tuberculosis | 5 | 3 | 2 | [Lewinsohn et al., 2017] | |
| Endocrinology | Adrenal Insufficiency | 20 | 4 | 3 | [Charmandari et al., 2014] |
| Diabetes | 13 | 4 | 2 | [ElSayed et al., 2023] | |
| Pituitary | 12 | 4 | 3 | [Tritos and Miller, 2023] | |
| Thyroid Disease | 10 | 6 | 4 | [AlexanderErik et al., 2017] | |
Table 6 provides a detailed breakdown of the disease categories included in DiReCT. The column labeled # samples indicates the number of data points. The symbols $|D_i|$ and $|D^⋆_i|$ denote the total number of diagnoses (diseases) and PDDs, respectively. Existing guidelines for diagnosing diseases were used as References, forming the foundation for constructing the diagnostic knowledge graphs. As some premise may not included in the referred guidelines. During annotation, physicians will incorporate their own knowledge to complete the knowledge graph.
### A.2 Structure of Knowledge Graph
The entire knowledge graph, denoted as $K$ , is stored in separate JSON files, each corresponding to a specific disease category $i$ as $K_i$ . Each $K_i$ comprises a procedural graph $G_i$ and the corresponding premise $p$ for each disease. As illustrated in Figure 7, the procedural graph $G_i$ is stored under the key "Diagnostic" in a dictionary structure. A key with an empty list as its value indicates a leaf diagnostic node as $d^⋆$ . The premise for each disease is saved under the key of "Knowledge" with the corresponding disease name as an index. For all the root nodes (e.g., Suspected Heart Failure), we further divide the premise into "Risk Factors", "Symptoms", and "Signs". Note that each premise is separated by ";".
<details>
<summary>x7.png Details</summary>

### Visual Description
## Diagnostic Framework: Heart Failure Classification and Clinical Criteria
### Overview
The image displays a structured, text-based diagnostic framework for heart failure, presented in a nested JSON-like format. It is divided into two primary sections: **"Diagnostic"** and **"Knowledge"**. The "Diagnostic" section outlines a hierarchical classification system for heart failure, while the "Knowledge" section provides the clinical criteria (risk factors, symptoms, signs, and diagnostic thresholds) used to populate that classification.
### Components/Axes
The content is organized as a hierarchical data structure with the following top-level keys:
1. **"Diagnostic"**: Contains a nested tree for classifying heart failure cases.
2. **"Knowledge"**: Contains the clinical reference data used for diagnosis.
**Spatial Layout:** The text is left-aligned and indented to show hierarchy. The entire block is enclosed within a dashed border.
### Detailed Analysis / Content Details
#### 1. Diagnostic Section
This section defines a patient classification pathway:
* **Root:** `"Diagnostic"`
* **Level 1:** `"Suspected Heart Failure"`
* **Level 2:** `"Strongly Suspected Heart Failure"`
* **Level 3:** `"Heart Failure"`
* **Level 4 (Subtypes):**
* `"HFrEF": []` (Heart Failure with Reduced Ejection Fraction)
* `"HFmrEF": []` (Heart Failure with Mildly Reduced Ejection Fraction)
* `"HFpEF": []` (Heart Failure with Preserved Ejection Fraction)
*Note: The empty arrays `[]` at the leaf nodes suggest this is a template or schema where specific patient data or evidence would be inserted.*
#### 2. Knowledge Section
This section lists the clinical evidence required for each diagnostic level.
* **For "Suspected Heart Failure":**
* **"Risk Factors":** A comprehensive list including: CAD; Hypertension; Valve disease; Arrhythmias; CMPs; Congenital heart disease; Infective; Drug-induced; Infiltrative, Storage disorders, Endomyocardial disease, Pericardial disease, Metabolic, Neuromuscular disease.
* **"Symptoms":** A detailed list including: Breathlessness; Orthopnoea; Paroxysmal nocturnal dyspnoea; Reduced exercise tolerance; Fatigue; tiredness; increased time to recover after exercise; Ankle swelling; Nocturnal cough; Wheezing; Bloated feeling; Loss of appetite; Confusion (especially in the elderly); Depression; Palpitation; Dizziness; Syncope.
* **"Signs":** A detailed list including: Elevated jugular venous pressure; Hepatojugular reflux; Third heart sound (gallop rhythm); Laterally displaced apical impulse; Weight gain (>2 kg/week); Weight loss (in advanced HF); Tissue wasting (cachexia); Cardiac murmur; Peripheral edema (ankle, sacral, scrotal); Pulmonary crepitations; Pleural effusion; Tachycardia; Irregular pulse; Tachypnoea; Cheyne-Stokes respiration; Hepatomegaly; Ascites; Cold extremities; Oliguria; Narrow pulse pressure.
* **For "Strongly Suspected Heart Failure":**
* **Criterion:** `"NT-proBNP > 125 pg/mL; BNP > 35 pg/mL"`
* **For "Heart Failure" (Confirmation):**
* **Criterion:** `"Abnormal findings from echocardiography: LV mass index>95 g/m2 (Female), > 115 g/m2 (Male); Relative wall thickness >0.42; LA volume index>34 mL/m2; E/e ratio at rest >9; PA systolic pressure >35 mmHg; TR velocity at rest >2.8 m/s"`
* **For the Heart Failure Subtypes (Ejection Fraction Criteria):**
* **"HFrEF":** `"LVEF<40%"`
* **"HFmrEF":** `"LVEF41-49%"`
* **"HFpEF":** `"LVEF>50%"`
### Key Observations
1. **Hierarchical Logic:** The framework follows a clear diagnostic cascade: from initial suspicion based on risk factors and clinical presentation, to stronger suspicion based on biomarkers (BNP/NT-proBNP), to definitive diagnosis via imaging (echocardiography), and finally to sub-classification based on Left Ventricular Ejection Fraction (LVEF).
2. **Comprehensive Clinical Data:** The "Knowledge" section is exhaustive, listing a wide array of potential risk factors, symptoms, and physical signs, indicating a thorough clinical reference.
3. **Quantitative Thresholds:** Specific, numerical diagnostic thresholds are provided for biomarkers, echocardiographic parameters, and LVEF, making the framework actionable for clinical decision-making.
4. **Template Structure:** The empty arrays `[]` in the "Diagnostic" section strongly suggest this is a data schema or template designed to be populated with individual patient findings.
### Interpretation
This image represents a **clinical decision support algorithm or data model** for heart failure. It translates complex clinical guidelines into a structured, machine-readable (or at least highly organized) format.
* **What it demonstrates:** It maps the journey from a patient presenting with vague symptoms to a precise, sub-typed diagnosis. The "Knowledge" section acts as the lookup table or rule set, while the "Diagnostic" section is the output classification tree.
* **Relationships:** The "Knowledge" criteria are the inputs that determine a patient's position within the "Diagnostic" hierarchy. For example, a patient with risk factors and symptoms is "Suspected." If their BNP is elevated, they become "Strongly Suspected." If an echocardiogram shows specific abnormalities, they are confirmed to have "Heart Failure," and their LVEF value then places them in the HFrEF, HFmrEF, or HFpEF category.
* **Notable Anomalies/Outliers:** The framework is purely clinical and does not account for patient history, comorbidities, or treatment response, which are crucial in real-world diagnosis. The strict numerical cutoffs (e.g., LVEF 40% vs. 41%) are necessary for classification but represent clinical gray zones in practice.
* **Underlying Purpose:** This structure is likely intended for use in electronic health records (EHR), clinical research databases, or diagnostic software to standardize the diagnosis and classification of heart failure, ensuring consistency and facilitating data analysis.
</details>
Figure 7: A sample of knowledge graph for Heart Failure. Each premise under the key of "Knowledge" is separated with ";".
### A.3 Annotation and Tools
We have developed proprietary software for annotation purposes. As depicted in Figure 8, annotators are presented with the original text as observations $o$ and are required to provide rationales ( $z$ ) to explain why a particular observation $o$ supports a disease $d$ . The left section of the figure, labeled Input1 to Input6, corresponds to different parts of the clinical note, specifically the chief complaint, history of present illness, past medical history, family history, physical exam, and pertinent results, respectively. Annotators will add the raw text into the first layer by left-clicking and dragging to select the original text, then right-clicking to add it. After each observation, a white box will be used to record the rationales. Finally, a connection will be made from each rationale to a disease, represented in a grey box. The annotation process strictly follow the knowledge graph. Both the final annotation and the raw clinical note will be saved in a JSON file. We provide the code to compile these annotations and detailed instructions for using our tool on GitHub.
<details>
<summary>extracted/5772958/Images_spp/annotation.png Details</summary>
### Visual Description
## Medical Diagnostic Reasoning Diagram: Diabetes Type II Inference Flow
### Overview
This image is a screenshot of a specialized medical software application titled "Medical." It displays a multi-layered diagnostic reasoning diagram that visually maps clinical evidence (tests, symptoms, risk factors) to a final diagnosis of Type II Diabetes. The interface includes an input text panel on the left and a five-layer flowchart on the right, connected by directional arrows.
### Components/Axes
**Application Interface:**
- **Title Bar:** "Medical"
- **Menu Bar:** Options include "Output json," "Read json," "Restart," "Refresh," "Reset background color."
- **Left Panel ("Input text"):** Contains tabs labeled `Input1` through `Input6`. The active tab (`Input2`) shows a blue text area with partially visible patient notes. A tooltip/context menu labeled "Add to first" is visible.
- **Right Panel (Main Diagram):** Organized into five vertical columns labeled `Layer1`, `Layer2`, `Layer3`, `Layer4`, and `Layer5`.
**Diagram Structure & Color Coding:**
- **Layer1 (Evidence Input):** Contains colored boxes representing clinical data.
- **Beige Boxes (Test Results):**
1. "C-peptide release test hints the peak late did not fall back"
2. "Insulin release test hints the peak late did not fall back"
3. "BLOOD Glucose-298"
- **Blue Boxes (Symptoms/Conditions):**
4. "CKD"
5. "progressive short term memory loss"
- **Yellow Boxes (Risk Factors):**
6. "Hypertension"
7. "Dyslipidemia"
8. "Coronary artery disease"
- **Layer2 (Interpretation):** Each Layer1 box connects via a red arrow to a corresponding white box that interprets the evidence.
- **Layer3 (Intermediate Conclusion):** A single gray box labeled "Suspected Diabetes."
- **Layer4 (Primary Diagnosis):** A single gray box labeled "Diabetes."
- **Layer5 (Specific Diagnosis):** A single gray box labeled "Type II Diabetes."
### Detailed Analysis
**Flow and Connections:**
1. **From Layer1 to Layer2:** Each piece of evidence is interpreted.
- C-peptide test → "Related C-peptide peak is more common in patients with type II"
- Insulin test → "Related insulin peak is more common in patients with type II"
- Blood Glucose-298 → "Abnormal random blood glucose is a diagnostic criteria of diabetes"
- CKD → "CKD is a kind of microangiopathy, which is a symptom of diabetes"
- Memory loss → "Progressive short term memory loss is a symptom of diabetes"
- Hypertension → "Hypertension is a risk factor of diabetes"
- Dyslipidemia → "Dyslipidemia is a risk factor of diabetes"
- Coronary artery disease → "Coronary artery disease are risk factors of diabetes"
2. **From Layer2 to Layer3 (Suspected Diabetes):** All Layer2 interpretations **except the first two** (C-peptide and Insulin) have red arrows converging on the "Suspected Diabetes" box. This indicates that symptoms and risk factors collectively raise suspicion.
3. **From Layer3 to Layer4 to Layer5 (Diagnostic Chain):** A linear path: "Suspected Diabetes" → "Diabetes" → "Type II Diabetes."
4. **Direct Path to Specific Diagnosis:** The first two Layer2 boxes (C-peptide and Insulin interpretations) have red lines that **bypass Layers 3 and 4**, connecting directly to the "Type II Diabetes" box in Layer5. This signifies that these specific test results are strong, direct indicators for Type II Diabetes specifically.
**Input Text Fragment (Left Panel):**
Visible text in the `Input2` tab includes:
- "...believe that he is prescribed too many medications, and..."
- "...at when he takes all of them, he is more fatigued and less in..."
- "...s pl... since his last d..."
This suggests the input is a patient history note describing polypharmacy and fatigue, which are clinically relevant to the diagnostic process.
### Key Observations
1. **Hierarchical Reasoning:** The diagram models a diagnostic thought process, moving from raw data (Layer1) to interpretation (Layer2), to a working hypothesis (Layer3), to a general diagnosis (Layer4), and finally to a specific subtype (Layer5).
2. **Evidence Weighting:** The system visually differentiates the strength of evidence. Symptoms and risk factors (blue/yellow) lead to suspicion, while specific metabolic test results (beige) provide a direct path to the final diagnosis.
3. **Clinical Logic:** The interpretations in Layer2 correctly link the evidence to diabetes pathology (e.g., CKD as a microangiopathy, hypertension as a risk factor).
4. **Data Point:** A specific lab value is noted: "BLOOD Glucose-298" (presumably mg/dL), which is a markedly high random glucose level, a key diagnostic criterion.
### Interpretation
This diagram represents a **clinical decision support system** or a **knowledge graph** for diabetes diagnosis. It demonstrates how disparate clinical data points are synthesized through a logical, layered inference engine.
- **What it suggests:** The patient profile likely includes elevated blood glucose, abnormal C-peptide/insulin response patterns, and comorbid conditions (CKD, hypertension, etc.). The system concludes this constellation of findings is most consistent with **Type II Diabetes**.
- **Relationships:** The flowchart explicitly shows that while many factors contribute to a general suspicion of diabetes, the **C-peptide and insulin release test patterns** are treated as highly specific biomarkers for the Type II subtype, warranting a direct diagnostic link.
- **Notable Anomaly/Insight:** The direct connection from the C-peptide/Insulin tests to the final diagnosis, bypassing intermediate steps, highlights a key diagnostic principle: these tests can differentiate Type II from Type I diabetes (where C-peptide is typically low). The system encodes this medical knowledge directly into its visual logic.
- **Purpose:** The tool likely aids in standardizing diagnostic reasoning, ensuring all relevant evidence is considered, and providing a transparent, auditable trail from data to conclusion. The "Output json" menu option suggests the reasoning process can be exported as structured data.
</details>
Figure 8: Demonstration of our annotation tool.
### A.4 Access to DiReCT
Implementation code and annotation tool are available through https://github.com/wbw520/DiReCT. Data will be released through PhysioNet due to safety issues according to the license of MIMIC-IV (PhysioNet Credentialed Health Data License 1.5.0). We will use the same license for DiReCT. The download link will be accessible via GitHub. We confirm that this GitHub link and data link are always accessible. We confirm that we will bear all responsibility in case of violation of rights.
## Appendix B Implementation of Baseline Method
### B.1 Prompt Settings
Table 7: Prompt for narrowing-down module.
| Input Prompt |
| --- |
| Suppose you are one of the greatest AI scientists and medical expert. Let us think step by step. |
| You will review a clinical ’Note’ and your ’Response’ is to diagnose the disease that the patient have for this admission. |
| All possible disease options are in a list structure: {disease_option}. |
| Note that you can only choose one disease from the disease options and directly output the origin name of that disease. |
| Now, start to complete your task. |
| Don’t output any information other than your ’Response’. |
| ’Note’: |
| {note} |
| Your ’Response’: |
Table 8: Prompt for perception module.
| Input Prompt |
| --- |
| Suppose you are one of the greatest AI scientists and medical expert. Let us think step by step. |
| You will review a part of clinical "Note" from a patient. |
| The disease for which the patient was admitted to hospital this time is {disease}. |
| Your task is to extract the original text as confidence "Observations" that lead to {disease}. |
| Here are some premise for the diagnosis of this disease category. You can refer them for your task. Premise are: {premise} |
| Note that you also need to briefly provide the "Reason" for your extraction. |
| Note that both "Observations" and "Reason" should be string. |
| Note that your "Response" should be a list structure as following |
| : [["Observation", "Reason"], ……, ["Observation", "Reason"]] |
| Note that if you can’t find any "Observation" your "Response" should be: []. |
| Now, start to complete your task. |
| Note that you should not output any information other than your "Response". |
| "Note": |
| {note} |
| Note that you should not output any information other than your "Response". |
| Your "Response": |
Table 9: Prompt for reasoning module.
| Input Prompt |
| --- |
| Suppose you are one of the greatest AI scientists and medical expert. Let us think step by step. |
| You will receive a list of "Observations" from a clinical "Note". These "Observations" are possible support to diagnose {disease}. |
| Based on these "Observations", you need to diagnose the "Disease" from the following options: {disease_option}. |
| Here are some golden standards to discriminate diseases. You can refer them for your task. Golden standards are: {premise} |
| Note that you can only choose one "Disease" from the disease options and directly output the name in disease options. |
| Note that you also required to select the "Observations" that satisfy the golden standard to diagnose the "Disease" you choose. |
| Note that you also required to provide the "Reason" for your choice. |
| Note that your "Response" should be a list structure as following |
| :[["Observation", "Reason", "Disease"], ……, ["Observation", "Reason", "Disease"]] |
| Note that if you can’t find any "Observation" to support a disease option, your "Response" should be: None |
| Now, start to complete your task. |
| Note that you should not output any information other than your "Response". |
| "Observations": |
| {observation} |
| Note that you should not output any information other than your "Response". |
| Your "Response": |
In this section, we demonstrate the prompt we used for each module (From Table 7 - 9 for narrowing-down, perception, and reasoning module, respectively).
In Table 7, {disease_option} is the name for all disease categories, and {note} is the content for the whole clinical note. The response for the model is the name of a possible disease $\hat{i}$ .
In Table 8, {disease} is the disease category name predicted in narrowing-down. The content marked blue is the premise, which is only provided during the $k$ setting. In this module, {premise} is offered with all information in the knowledge graph. Different to narrowing-down, {note} is implemented for each clinical data $R=\{r\}$ and the outputs are combined together for $\hat{O}$ and $\hat{E}$ .
In Table 9, {disease} is the disease category name and {disease_option} is consisted by the children nodes $\{d_n\}_n$ . Similarly, the premise on the blue is only available for the $k$ setting. It provides the premise that are criteria for the diagnosis of each children node. {observation} is the extracted $\hat{O}$ in previous step. We provide all the prompts and the complete implementation code on GitHub.
### B.2 Details of Automatic Evaluation
The automatic evaluation is realized by LLama3 8B. We demonstrate the prompt for this implement in Table 10 (for observation) and Table 11 (for rationalization). Note that we do not use few-shot samples for the evaluation of observation. In Table 10, {gt_observation} and {pred_observation} are from model prediction and ground-truth. As this is a simple similarity comparison task to discriminate whether the model finds similar observations to humans, LLama3 itself have such ability. We do not strict to exactly match due to the difference in length of extracted raw text (as long as the observation expresses the same description). In Table 11, {gt_reasoning} and {pred_reasoning} are from model prediction and ground-truth. We require the rationale to be complete (content of the expression can be understood from the rationale alone) and meaningful; therefore, we provide five samples for this evaluation. We also provide all the prompts and the complete implementation code on GitHub.
Table 10: Prompt for evaluation of observation.
| Input Prompt |
| --- |
| Suppose you are one of the greatest AI scientists and medical expert. Let us think step by step. |
| You will receive two "Observations" extracted from a patient’s clinical note. |
| Your task is to discriminate whether they textually description is similar? |
| Note that "Response" should be one selection from "Yes" or "No". |
| Now, start to complete your task. |
| Don’t output any information other than your "Response". |
| "Observation 1": {gt_observation} |
| "Observation 2": {pred_observation} |
| Your "Response": |
Table 11: Prompt for evaluation of rationalization.
| Input Prompt |
| --- |
| Suppose you are one of the greatest AI scientists and medical expert. Let us think step by step. |
| You will receive two "Reasoning" for the explanation of why an observation cause a disease. |
| Your task is to discriminate whether they explain a similar medical diagnosis premise? |
| Note that "Response" should be one selection from "Yes" or "No". |
| Here are some samples: |
| Sample 1: |
| "Reasoning 1": Facial sagging is a classic symptom of stroke |
| "Reasoning 2": Indicates possible facial nerve palsy, a common symptom of stroke |
| "Response": Yes |
| Sample 2: |
| "Reasoning 1": Family history of Diabetes is an important factor |
| "Reasoning 2": Patient’s mother had a history of Diabetes, indicating a possible genetic predisposition to stroke |
| "Response": Yes |
| Sample 3: |
| "Reasoning 1": headache is one of the common symptoms of HTN |
| "Reasoning 2": Possible symptom of HTN |
| "Response": No |
| Sample 4: |
| "Reasoning 1": Acute bleeding is one of the typical symptoms of hemorrhagic stroke |
| "Reasoning 2": The presence of high-density areas on Non-contrast CT Scan is a golden standard for Hemorrhagic Stroke |
| "Response": No |
| Sample 5: |
| "Reasoning 1": Loss of strength on one side of the body, especially when compared to the other side, is a common sign of stroke |
| "Reasoning 2": Supports ischemic stroke diagnosis |
| "Response": No |
| Now, start to complete your task. |
| Don’t output any information other than your "Response". |
| "Reasoning 1": {gt_reasoning} |
| "Reasoning 2": {pred_reasoning} |
| Your "Response": |
### B.3 Prediction Samples
Figure 9 and 10 shows two sample generated by GPT-4. The ground-truth PDD of the input clinical note is Gastroesophageal Reflux Disease (GERD) and Heart Failure (HF). In these figure, purple, orange, and red indicate explanations only in the ground truth, only in prediction, and common in both, respectively; therefore, red is a successful prediction of an explanation, while purple and orange are a false negative and false positive.
<details>
<summary>x8.png Details</summary>
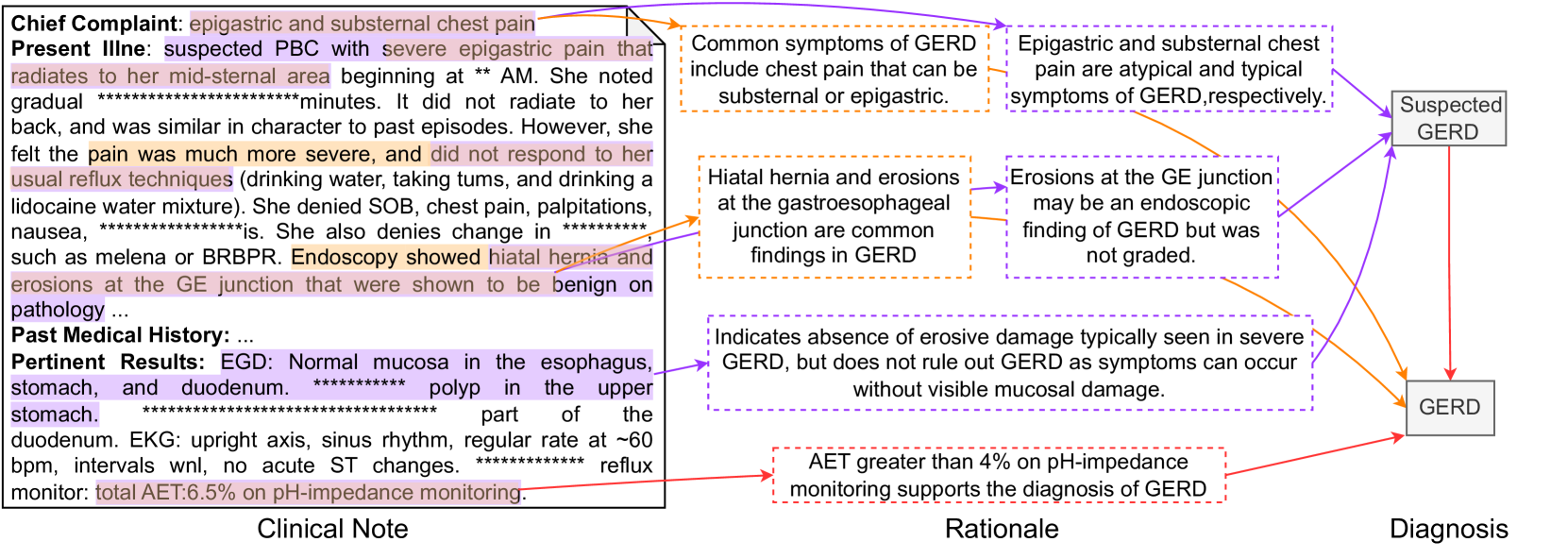
### Visual Description
## Medical Case Analysis Diagram: GERD Diagnostic Pathway
### Overview
This image is a structured medical case analysis diagram that traces the diagnostic reasoning for a patient presenting with chest pain. It visually maps specific findings from a clinical note to their medical rationale, culminating in a diagnosis of GERD (Gastroesophageal Reflux Disease). The diagram is organized into three vertical columns: **Clinical Note** (left), **Rationale** (center), and **Diagnosis** (right). Colored highlights and arrows create explicit links between patient data, medical reasoning, and diagnostic conclusions.
### Components/Axes
The diagram is segmented into three primary regions:
1. **Clinical Note (Left Column):** Contains transcribed excerpts from a patient's medical record.
* **Sections:** Chief Complaint, Present Illness, Past Medical History (truncated), Pertinent Results.
* **Highlighted Text Segments:** Text is highlighted in three colors, each corresponding to a different diagnostic thread:
* **Orange:** Symptoms and findings commonly associated with GERD.
* **Purple:** Atypical symptoms or findings that require specific interpretation.
* **Red:** Objective test results that provide strong diagnostic evidence.
* **Redacted Information:** Several sections of text are obscured with asterisks (`***`), indicating omitted or irrelevant details for this specific analysis.
2. **Rationale (Center Column):** Contains explanatory text boxes that interpret the highlighted clinical findings.
* **Box Style:** Each rationale is enclosed in a dashed border, with the border color matching the highlight color from the Clinical Note (orange, purple, or red).
* **Content:** Each box provides a medical fact or interpretation that connects a specific clinical finding to the diagnosis of GERD.
3. **Diagnosis (Right Column):** Contains the final diagnostic conclusions.
* **Boxes:** Two gray boxes labeled "Suspected GERD" and "GERD".
* **Flow:** Arrows indicate the progression from suspicion to a more definitive diagnosis based on the aggregated evidence.
**Spatial Grounding & Connections:**
* **Arrows:** Colored arrows (orange, purple, red) originate from highlighted text in the Clinical Note, pass through or connect to corresponding rationale boxes, and terminate at the diagnosis boxes.
* **Legend/Color Code:** The color of the highlight, rationale box border, and connecting arrow are consistent for each diagnostic thread:
* **Orange Thread:** Links common GERD symptoms (chest pain) and common endoscopic findings (hiatal hernia, erosions) to both "Suspected GERD" and "GERD".
* **Purple Thread:** Links atypical symptom descriptions and the *absence* of severe erosive damage to "Suspected GERD" and "GERD", with a rationale clarifying that this does not rule out GERD.
* **Red Thread:** Links the objective pH-impedance monitoring result directly to the "GERD" diagnosis box.
### Detailed Analysis
**Clinical Note Transcription (with highlights noted):**
* **Chief Complaint:** `epigastric and substernal chest pain` (Highlighted in **Orange**)
* **Present Illness:** `suspected PBC with severe epigastric pain that radiates to her mid-sternal area` (Highlighted in **Purple**). The note describes pain not responding to usual reflux techniques. It also states: `Endoscopy showed hiatal hernia and erosions at the GE junction that were shown to be benign on pathology` (Highlighted in **Orange**).
* **Pertinent Results:**
* `EGD: Normal mucosa in the esophagus, stomach, and duodenum. ********** polyp in the upper stomach. *************** part of the duodenum.` (Highlighted in **Purple**)
* `EKG: upright axis, sinus rhythm, regular rate at ~60 bpm, intervals wnl, no acute ST changes. *********** reflux monitor: total AET:6.5% on pH-impedance monitoring.` (Highlighted in **Red**)
**Rationale Boxes (Transcribed):**
1. **(Orange Border):** `Common symptoms of GERD include chest pain that can be substernal or epigastric.`
2. **(Orange Border):** `Hiatal hernia and erosions at the gastroesophageal junction are common findings in GERD`
3. **(Purple Border):** `Epigastric and substernal chest pain are atypical and typical symptoms of GERD, respectively.`
4. **(Purple Border):** `Erosions at the GE junction may be an endoscopic finding of GERD but was not graded.`
5. **(Purple Border):** `Indicates absence of erosive damage typically seen in severe GERD, but does not rule out GERD as symptoms can occur without visible mucosal damage.`
6. **(Red Border):** `AET greater than 4% on pH-impedance monitoring supports the diagnosis of GERD`
**Diagnosis Flow:**
* Arrows from the orange and purple rationale threads converge on the **"Suspected GERD"** box.
* All three threads (orange, purple, and red) have arrows pointing to the final **"GERD"** box, indicating a conclusive diagnosis based on the totality of evidence.
### Key Observations
1. **Multi-Evidence Diagnosis:** The diagnosis is not based on a single finding but on a confluence of symptomatic, endoscopic, and objective physiological data.
2. **Handling of Contradictory Evidence:** The diagram explicitly addresses a potential contradiction: the endoscopy showed only mild findings (hiatal hernia, non-graded erosions) and "normal mucosa," which might suggest severe erosive GERD is absent. The purple rationale box clarifies that this absence does not rule out GERD.
3. **Objective vs. Subjective Data:** The strongest, most objective piece of evidence (pH-impedance monitoring with AET 6.5%) is highlighted in red and has a direct, unambiguous arrow to the final "GERD" diagnosis.
4. **Use of Redaction:** The use of `***` focuses the analysis on the medically pertinent information, stripping away irrelevant history or details for this specific diagnostic question.
### Interpretation
This diagram serves as a visual clinical reasoning tool. It demonstrates how a physician synthesizes disparate pieces of patient data to arrive at a diagnosis.
* **What the Data Suggests:** The patient's presentation of epigastric/substernal pain, combined with common endoscopic findings (hiatal hernia) and, most importantly, an Acid Exposure Time (AET) of 6.5% (above the 4% diagnostic threshold), strongly supports a diagnosis of GERD. The diagram successfully argues that even in the absence of severe erosive damage on endoscopy, the symptomatic and pH-impedance evidence is sufficient.
* **Relationship Between Elements:** The arrows create a clear "if-then" logic chain. *If* a patient has symptom X (orange), *then* it is a common symptom of GERD. *If* a test shows result Y (red), *then* it supports the diagnosis. The rationale boxes provide the medical knowledge that justifies these logical links.
* **Notable Pattern:** The pathway highlights a modern diagnostic approach to GERD, which relies heavily on objective physiological testing (pH-impedance monitoring) to confirm or rule out the disease, especially when endoscopic findings are ambiguous or mild. The diagram effectively communicates that GERD is a clinical diagnosis supported by evidence, not solely defined by visible damage to the esophagus.
</details>
Figure 9: An example prediction for a clinical note with PDD of GERD by GPT-4
<details>
<summary>x9.png Details</summary>
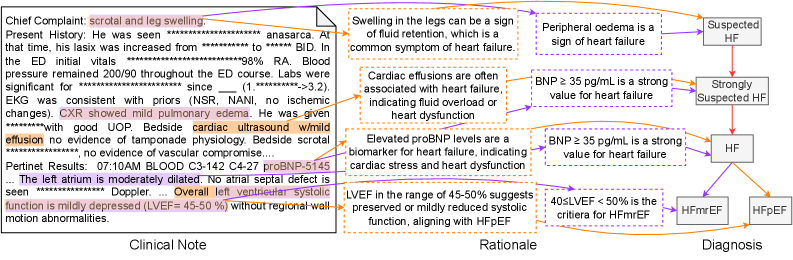
### Visual Description
## Clinical Flowchart: Heart Failure Diagnosis Pathway
### Overview
The image is a structured clinical flowchart that maps a patient's presentation from initial symptoms and clinical findings through a diagnostic rationale to a final classification of heart failure. It is divided into three distinct horizontal sections: **Clinical Note** (left), **Rationale** (center), and **Diagnosis** (right). The flow is indicated by purple arrows connecting text boxes and annotations.
### Components/Axes
The diagram is organized into three primary columns or regions:
1. **Clinical Note (Left Region):** Contains the raw patient data and findings.
2. **Rationale (Center Region):** Contains interpretive statements that link clinical findings to diagnostic criteria.
3. **Diagnosis (Right Region):** Contains the final diagnostic classifications.
**Key Visual Elements:**
* **Text Boxes:** Contain the core information.
* **Purple Arrows:** Indicate the logical flow and relationships between elements.
* **Highlighted Text:** Certain phrases within the Clinical Note are highlighted in red or purple, emphasizing key abnormal findings or values.
* **Annotations:** Small text boxes with arrows point to specific parts of the Clinical Note to provide explanatory context (e.g., "Swelling in the legs can be a sign of fluid retention...").
### Detailed Analysis / Content Details
**1. Clinical Note (Left Region)**
* **Chief Complaint:** `scrotal and leg swelling`
* **Present History:** Patient presented with anasarca. At that time, his lasix was increased from `******` to `******`. In the ED, initial vitals: `HR 98% RA`. Blood pressure remained `200/90` throughout the ED course. Labs significant for `creatinine 1.3 -> 3.2`. EKG was consistent with priors (NSR, NANI) no ischemic changes. He was admitted to medicine service with good UOP. Beside cardiac ultrasound showed `world effusion` no evidence of tamponade physiology. Beside scrotal ultrasound, no evidence of vascular compromise.
* **Perinet Results:** `07/10 AM BLOOD C3-142 C4-27 proBNP 345`
* **Impression:** `The left atrium is mildly dilated. No atrial septal defect is seen. Normal Doppler. Overall left ventricular systolic function is mildly depressed (LVEF 45-50 %) without regional wall abnormalities.`
**2. Rationale (Center Region)**
This section contains five interpretive statements, each linked to specific findings:
* `Swelling in the legs can be a sign of fluid retention, which is a common symptom of heart failure.`
* `Peripheral oedema is a sign of heart failure.`
* `Cardiac effusions are often associated with heart failure, indicating fluid overload or heart dysfunction.`
* `Elevated proBNP levels are a biomarker for heart failure, indicating cardiac stress and heart dysfunction.`
* `LVEF in the range of 45-50% suggests preserved or mildly reduced systolic function, aligning with HFpEF.`
**3. Diagnosis (Right Region)**
This section shows a diagnostic decision tree:
* **Top Box:** `Suspected HF` (connected from "Peripheral oedema is a sign of heart failure").
* **Middle Box:** `Strongly Suspected HF` (connected from two rationales: "BNP ≥ 35 pg/mL is a strong value for heart failure" and "BNP ≥ 35 pg/mL is a strong value for heart failure" - note the duplicate rationale text).
* **Central Box:** `HF` (connected from "Strongly Suspected HF" and the rationale about elevated proBNP).
* **Bottom Branching:** `HF` leads to two final classifications:
* `HFmrEF` (connected via the rationale "40<LVEF <50 % is the criteria for HFmrEF").
* `HFpEF` (connected via the rationale about LVEF 45-50% aligning with HFpEF).
### Key Observations
1. **Data Flow:** The diagram explicitly traces the diagnostic reasoning from a symptom (swelling) through objective findings (imaging, labs) to a specific heart failure classification.
2. **Critical Values:** Key numerical thresholds are highlighted:
* `proBNP 345` (pg/mL) is noted as elevated.
* `LVEF 45-50 %` is the central value for classification.
* The rationale mentions `BNP ≥ 35 pg/mL` as a strong indicator, though the patient's value is given as `proBNP 345`.
3. **Diagnostic Criteria:** The flowchart applies specific criteria:
* `HFmrEF` (Heart Failure with mildly Reduced Ejection Fraction) is defined here as `40 < LVEF < 50 %`.
* `HFpEF` (Heart Failure with preserved Ejection Fraction) is associated with the patient's `LVEF 45-50 %`.
4. **Potential Inconsistency:** The rationale box stating "BNP ≥ 35 pg/mL is a strong value for heart failure" appears twice, pointing to the same "Strongly Suspected HF" box. The patient's lab result is for `proBNP` (value 345), not standard BNP. The rationale text may be a general statement, while the patient's specific value (345) is used to support the "Strongly Suspected" status.
5. **Spatial Layout:** The "Clinical Note" is a dense block of text on the left. The "Rationale" statements are arranged vertically in the center, each with an arrow pointing to a specific part of the Clinical Note. The "Diagnosis" tree is on the right, with a top-down flow from "Suspected" to specific classifications.
### Interpretation
This flowchart serves as a visual clinical decision support tool or an educational diagram illustrating the diagnostic pathway for a specific patient case. It demonstrates how disparate clinical data points—symptoms (edema), imaging (mildly dilated left atrium, reduced LVEF), and biomarkers (elevated proBNP)—are synthesized using established medical rationale to arrive at a nuanced diagnosis.
The data suggests the patient presents with clear signs of heart failure (fluid overload, elevated cardiac biomarker, reduced systolic function). The critical interpretation lies in the LVEF value of 45-50%. This value sits in a borderline zone, leading to the dual classification possibility of **HFmrEF** (by one criterion of 40-50%) and **HFpEF** (as it is at the upper end of that range and "preserved" function is often considered ≥50%). The diagram highlights the importance of precise ejection fraction measurement in sub-classifying heart failure, which has implications for treatment strategies. The inclusion of the proBNP value (345) alongside the general rationale about BNP thresholds shows the application of population-based guidelines to an individual case. The overall message is a structured, evidence-based approach to diagnosing and classifying heart failure from initial presentation.
</details>
Figure 10: An example prediction for a clinical note with PDD of HF by GPT-4
In Figure 9, we can observe that GPT-4 can find the key observation for the diagnosis of GERD, which is consistent with human in both observation and rationale. However, it still lacks the ability to identify all observations and establish accurate relationships for diseases. In Figure 10, the model’s predictions do not align well with those of a human doctor. Key observations, such as the relationships between BNP and LVEF, are incorrectly identified, leading to a final misdiagnosis.
### B.4 Experiments for No Extra Knowledge
Table 12: Prompt for $D^⋆$ setting.
| Input Prompt |
| --- |
| Suppose you are one of the greatest AI scientists and medical expert. Let us think step by step. |
| You will review a clinical ’Note’ and your ’Response’ is to diagnose the disease that the patient have for this admission. |
| All possible disease options are in a list structure: {disease_options}. |
| Note that you can only choose one disease from the disease options and directly output the origin name of that disease. |
| Now, start to complete your task. |
| Don’t output any information other than your ’Response’. |
| ’Note’: |
| {note} |
| Your ’Response’: |
Table 13: Prompt for no knowledge setting.
| Input Prompt |
| --- |
| Suppose you are one of the greatest AI scientists and medical expert. Let us think step by step. |
| You will review a clinical ’Note’ and your ’Response’ is to diagnose the disease that the patient have for this admission. |
| Note that you can only give one disease name and directly output the name of that "Disease". |
| Now, start to complete your task. |
| Don’t output any information other than your ’Response’. |
| ’Note’: |
| {note} |
| Your ’Response’: |
We demonstrate the prompt used for $D^⋆$ and no knowledge settings in Table 12 and Table 13, respectively. {note} is the text of whole clinical note and {note} in Table 12 is the name of all leaf node $D^⋆$ .
### B.5 Experimental Settings
All experiments are implemented with a temperature value of 0. All close sourced models are implemented in a local server with 4 NVIDIA A100 GPU.
## Appendix C Failed Attempts on DiReCT
In this section, we discuss some unsuccessful attempts during the experiments.
Extract observation from the whole clinical note. We try to diagnose the disease and extract observation, and the corresponding rationale using the prompt shown in Table 14. The {note} is offered by the whole content in the clinical note. We find that even though the model can make the correct diagnosis, only a few observations can be extracted (no more than 4), which decreases the completeness and faithfulness.
Table 14: Prompt for extracting observation in one step.
| Input Prompt |
| --- |
| Suppose you are one of the greatest AI scientists and medical expert. Let us think step by step. |
| You will review a clinical ’Note’, and your ’Response’ is to diagnose the disease that the patient has for this admission. |
| All possible disease options are in a list structure: {disease_options}. |
| Note that you can only choose one disease from the disease options and directly output the origin name of that disease. |
| Note that you also need to extract original text as confidence "Observations" that lead to the "Disease" you selected. |
| Note that you should extract all necessary "Observation". |
| Note that you also need to briefly provide the "Reason" for your extraction. |
| Note that both "Observations" and "Reason" should be string. |
| Note that your "Response" should be a list structure as following |
| :[["Observation", "Reason", "Disease"], ……, ["Observation", "Reason", "Disease"]] |
| Now, start to complete your task. |
| Don’t output any information other than your ’Response’. |
| ’Note’ |
| : {note} |
| Your ’Response’: |
End-to-End prediction. We also try to output the whole reasoning process in one step (without iteration) when given observations. We show our prompt in Table 15. We find that using such a prompt model can not correctly recognize the relation between observation, rationale, and diagnosis.
Table 15: Prompt for extracting observation in one step.
| Input Prompt |
| --- |
| Suppose you are one of the greatest AI scientists and medical expert. Let us think step by step. |
| You will receive a list of "Observations" from a clinical "Note" for the diagnosis of stroke. |
| Here is the diagnostic route of stroke in a tree structure: |
| -Suspected Stroke |
| -Hemorrhagic Stroke |
| -Ischemic Stroke |
| Here are some premise for the diagnosis of this disease. You can refer them for your task. Premise are: {premise} |
| Based on these "Observations", starting from the root disease, your target is to diagnose one of the leaf disease. |
| Note that you also required to provide the "Reason" for your reasoning. |
| Note that your "Response" should be a list structure as following |
| :[["Observation", "Reason", "Disease"], ……, ["Observation", "Reason", "Disease"]] |
| Note that if you can’t find any "Observation" to support a disease option, your "Response" should be: None |
| Now, start to complete your task. |
| Note that you should not output any information other than your "Response". |
| "Observations": |
| {observation} |
| Note that you should not output any information other than your "Response". |
| Your "Response": |
## Appendix D Ethical Considerations
Utilizing real-world EHRs, even in a de-identified form, poses inherent risks to patient privacy. Therefore, it is essential to implement rigorous data protection and privacy measures to safeguard sensitive information, in accordance with regulations such as HIPAA. We strictly adhere to the Data Use Agreement of the MIMIC dataset, ensuring that the data is not shared with any third parties. All experiments are implement on a private server. The access to GPT is also a private version.
AI models are susceptible to replicating and even intensifying the biases inherent in their training data. These biases, if not addressed, can have profound implications, particularly in sensitive domains such as healthcare. Unconscious biases in healthcare systems can result in significant disparities in the quality of care and health outcomes among different demographic groups. Therefore, it is imperative to rigorously examine AI models for potential biases and implement robust mechanisms for ongoing monitoring and evaluation. This involves analyzing the model’s performance across various demographic groups, identifying any disparities, and making necessary adjustments to ensure equitable treatment for all. Continual vigilance and proactive measures are essential to mitigate the risk of biased decision-making and to uphold the principles of fairness and justice in AI-driven healthcare solutions.