# ADEPT: Continual Pretraining via Adaptive Expansion and Dynamic Decoupled Tuning
> These authors contributed equally.Corresponding author.
Abstract
Conventional continual pretraining (CPT) for large language model (LLM) domain adaptation often suffers from catastrophic forgetting and limited domain capacity. Existing strategies adopt layer expansion, introducing additional trainable parameters to accommodate new knowledge. However, the uniform expansion and updates still entangle general and domain learning, undermining its effectiveness. Our pilot studies reveal that LLMs exhibit functional specialization, where layers and units differentially encode general-critical capabilities, suggesting that parameter expansion and optimization should be function-aware. We then propose ADEPT, A daptive Expansion and D ynamic D e coupled Tuning for continual p re t raining, a two-stage framework for domain-adaptive CPT. ADEPT first performs General-Competence Guided Selective Layer Expansion, duplicating layers least critical for the general domain to increase representational capacity while minimizing interference with general knowledge. It then applies Adaptive Unit-Wise Decoupled Tuning, disentangling parameter units within expanded layers according to their general-domain importance and assigning asymmetric learning rates to balance knowledge injection and retention. Experiments on mathematical and medical benchmarks show that ADEPT outperforms full-parameter CPT by up to 5.76% on the general domain and 5.58% on the target domain with only 15% of parameters tuned and less than 50% training time. Ablation studies, theoretical analysis, and extended investigations further demonstrate the necessity of targeted expansion and decoupled optimization, providing new principles for efficient and robust domain-adaptive CPT. Our code is open-sourced at https://github.com/PuppyKnightUniversity/ADEPT.
1 Introduction
Large language models (LLMs) have demonstrated remarkable performance across a wide range of general-domain tasks (OpenAI, 2023; Dubey et al., 2024c). However, their deployment in specialized domains, such as mathematics or healthcare, requires targeted adaptation (Ding et al., 2024; Chen et al., 2024; Ahn et al., 2024). Continual pretraining (CPT), which conducts post-pretraining on domain-specific corpora, has emerged as a crucial paradigm for injecting domain knowledge and capabilities into pretrained LLMs (Wu et al., 2024a; Ibrahim et al., 2024; Yıldız et al., 2024).
Despite its promise, CPT faces a persistent challenge: catastrophic forgetting. After pretraining, LLMs already encode substantial general knowledge, leaving limited parameter capacity for integrating new domain-specific information. While domain signals can be forcefully fitted through gradient-based optimization, the aggressive updates on the existing parameters come at the cost of overfitting to the target corpora, which in turn disrupts general abilities and triggers catastrophic forgetting (Liu et al., 2024a; Luo et al., 2025). This tension between new knowledge injection and previous knowledge retention poses a central obstacle to reliable and stable domain adaptation.
To address catastrophic forgetting, some approaches attempt through data-centric strategies, such as data replay or rehearsal (Huang et al., 2024; Zhang et al., 2025). While replay partially preserves prior knowledge, it fails to expand model capacity, leaving the conflict between knowledge injection and retention unresolved. Others focus on increasing capacity via transformer-layer extension (Wu et al., 2024b), yet typically insert new layers uniformly and update all parameters indiscriminately. This expansion strategy neglects the functional specialization within LLMs, where different layers and neurons serve distinct functional roles. Our pilot studies reveal that general-critical layers in LLMs are mainly located in early depths, and functional units within layers contribute unequally to general-domain performance, highlighting functional specialization similar to that found in the human brain (Xu et al., 2025; Zheng et al., 2024; Dai et al., 2022). Consequently, indiscriminate expansion and optimization may overwrite general-critical regions with new knowledge, compromising general competency preservation and leaving forgetting unresolved.
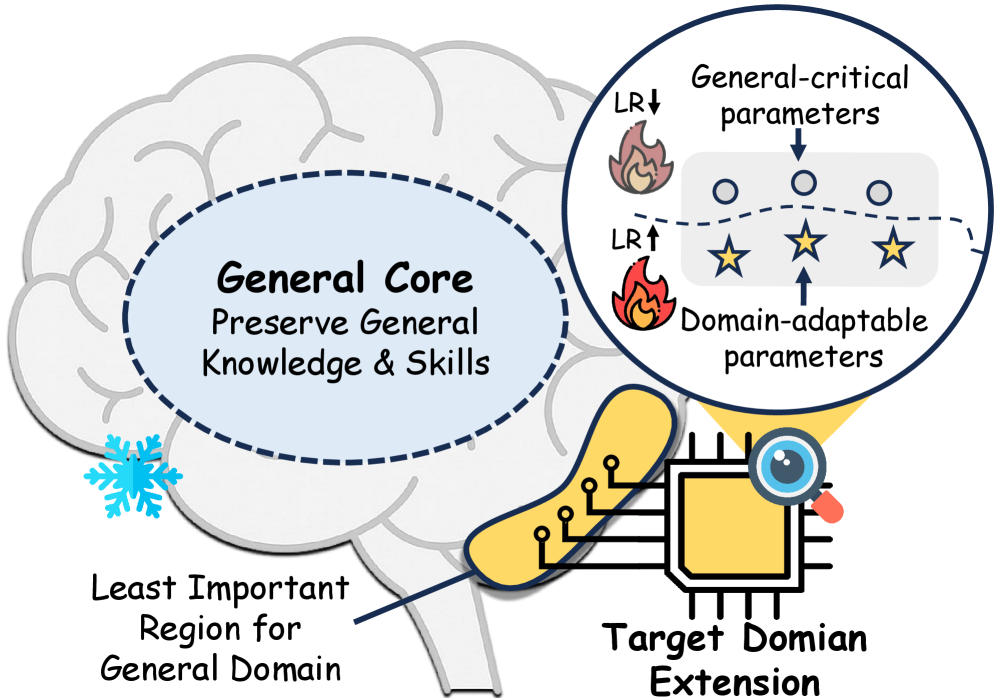
Inspired by the functional specialization perspective, we propose our core insight: effective CPT should expand and update the model adaptively, preserving the regions responsible for the general domain and targeting more adaptable parameters. Specifically, we argue that capacity allocation must be importance-guided, and optimization must be function-decoupled to minimize interference with general competencies. As illustrated in Figure 1, domain-specific extension should be allocated to the regions less constrained by general-domain knowledge and skills, and parameters within these regions should be decoupled and tuned accordingly, preserving general-critical parameters and allowing the rest to be more adaptable to absorb new domain-specific information.
<details>
<summary>x1.png Details</summary>
### Visual Description
## Diagram: Brain with Domain Adaptation
### Overview
The image is a diagram illustrating the concept of domain adaptation in the context of a brain-like structure. It shows a brain with labeled regions representing general knowledge, less important regions, and a target domain extension. A magnified view highlights the distinction between general-critical parameters and domain-adaptable parameters, indicating how learning rates (LR) are adjusted during adaptation.
### Components/Axes
* **Brain:** A simplified representation of a brain, serving as the central metaphor.
* **General Core:** A blue oval within the brain, labeled "General Core" and "Preserve General Knowledge & Skills."
* **Least Important Region for General Domain:** A region at the bottom-left of the brain, marked with a snowflake icon.
* **Target Domain Extension:** A yellow square with integrated circuit-like connections, representing the area for domain-specific learning.
* **Magnified View:** A circular inset showing "General-critical parameters" (represented by circles) and "Domain-adaptable parameters" (represented by stars).
* **Learning Rate (LR) Indicators:** Flames with arrows pointing up (LR↑) and down (LR↓) next to the parameters in the magnified view.
### Detailed Analysis
* **General Core:** Located in the center of the brain, this area is responsible for preserving general knowledge and skills.
* **Least Important Region:** Situated at the bottom-left, this region is deemed less crucial for general domain tasks, indicated by a blue snowflake.
* **Target Domain Extension:** A yellow square at the bottom-right, connected to the brain via a yellow extension, represents the area where domain-specific learning occurs. It has a magnifying glass icon, suggesting closer inspection.
* **Magnified View Details:**
* **General-critical parameters:** Represented by circles, these parameters are associated with a decreased learning rate (LR↓), indicated by a flame icon with a downward arrow.
* **Domain-adaptable parameters:** Represented by stars, these parameters are associated with an increased learning rate (LR↑), indicated by a flame icon with an upward arrow.
* The parameters are separated by a dashed line.
### Key Observations
* The diagram emphasizes the distinction between general knowledge and domain-specific learning.
* Learning rates are adjusted differently for general-critical and domain-adaptable parameters.
* The "Least Important Region" suggests a potential area for modification or pruning during domain adaptation.
### Interpretation
The diagram illustrates a domain adaptation strategy where the brain (or a neural network) retains its core general knowledge while adapting to a specific target domain. The "General Core" represents the pre-trained knowledge, while the "Target Domain Extension" signifies the area where new, domain-specific knowledge is acquired. The magnified view highlights the key mechanism: adjusting learning rates. General-critical parameters, essential for general knowledge, have their learning rates reduced to prevent catastrophic forgetting. Domain-adaptable parameters, crucial for the new domain, have their learning rates increased to facilitate learning. The "Least Important Region" might represent parameters that can be pruned or repurposed without significantly affecting general performance. This approach aims to balance knowledge retention and adaptation, enabling the system to perform well in both general and specific domains.
</details>
Figure 1: Illustration of the core idea of ADEPT. Target domain extension are applied on the least important region for general domain, minimizing catastrophic forgetting. Asymmetric learning rates are applied to parameter subsets for targeted knowledge injection.
Building on this insight, we propose A daptive Expansion and D ynamic D e coupled Tuning for continual p re- t raining (ADEPT), a framework for domain-adaptive continual pretraining. ADEPT comprises two stages: General-Competence Guided Selective Layer Expansion, which identifies and duplicates layers least critical for the general domain, allocating additional capacity precisely where interference with general capabilities is minimized, thereby preventing catastrophic forgetting. Adaptive Unit-Wise Decoupled Tuning, which disentangles the parameters within the expanded layers based on their importance to the general domain. Asymmetric learning rates are then applied on their subsets, ensuring that general-critical parameters are preserved while more adaptable parameters can fully absorb domain-specific knowledge. Extensive experiments on mathematical and medicine domains demonstrate that ADEPT enables efficient and robust domain knowledge injection, while substantially alleviating catastrophic forgetting. Specifically, compared to full-parameter CPT, ADEPT achieves up to 5.58% accuracy gain on target-domain benchmarks, and up to 5.76% gain on the general domain, confirming both effective knowledge acquisition and strong retention of general competencies. Furthermore, ADEPT attains these improvements with only 15% of parameters tuned, and reduces training time relative to other baselines greatly, highlighting its efficiency. Ablation studies and theoretical analysis further validate the designs of ADEPT.
To summarize, our contributions are threefold:
1. Insightfully, we highlight the importance of considering functional specialization in LLMs for continual pretraining through empirical experiments and theoretical analysis, advocating for targeted layer expansion and decoupled training as a principled solution to domain adaptation.
1. Technically, we propose ADEPT, a framework that consists of General-Competence Guided Selective Layer Expansion and Adaptive Unit-Wise Decoupled Tuning, enabling adaptive and effective domain knowledge integration while minimizing catastrophic forgetting.
1. Empirically, we conduct extensive experiments on both mathematical and medical domains, demonstrating that ADEPT consistently outperforms baselines in domain performance while preserving general competencies.
<details>
<summary>x2.png Details</summary>
### Visual Description
## Heatmaps: Qwen3 Model Layer Analysis
### Overview
The image presents three heatmaps comparing the activity across different layers and components of Qwen3 models with varying sizes (1.7B, 4B, and 8B parameters). The heatmaps visualize the relative activation levels within each layer for different modules, providing insights into how the model's processing changes with scale.
### Components/Axes
* **Titles:**
* Top-left: "Qwen3-1.7B-Base"
* Top-center: "Qwen3-4B-Base"
* Top-right: "Qwen3-8B-Base"
* **Y-axis:** "Layer"
* Leftmost heatmap: Ranges from 0 to 27, incrementing by 1.
* Center heatmap: Ranges from 0 to 34, incrementing by 2.
* Rightmost heatmap: Ranges from 0 to 34, incrementing by 2.
* **X-axis:** (Modules, from left to right)
* "mlp.down_proj"
* "mlp.up_proj"
* "self_atten.o_proj"
* "mlp.gate_proj"
* "self_atten.v_proj"
* "self_atten.q_proj"
* "self_atten.k_proj"
* "post_attention_layernorm"
* "input_layernorm"
* "self_atten.k_norm"
* "self_atten.a_norm"
* **Color Legend:** Located on the right side of the rightmost heatmap. The color gradient ranges from light green (0) to dark blue (1), representing the activation level.
### Detailed Analysis
**Qwen3-1.7B-Base (Left Heatmap):**
* The heatmap shows activity concentrated in the lower layers (0-10).
* **mlp.down_proj:** High activation in layers 0-9, decreasing gradually.
* **mlp.up_proj:** Similar to mlp.down_proj, high activation in layers 0-9.
* **self_atten.o_proj:** Moderate activation in layers 0-5.
* **mlp.gate_proj:** Low to moderate activation in layers 0-5.
* **self_atten.v_proj:** Very low activation across all layers.
* **self_atten.q_proj:** Very low activation across all layers.
* **self_atten.k_proj:** Very low activation across all layers.
* **post_attention_layernorm:** Very low activation across all layers.
* **input_layernorm:** Very low activation across all layers.
* **self_atten.k_norm:** Very low activation across all layers.
* **self_atten.a_norm:** Very low activation across all layers.
**Qwen3-4B-Base (Center Heatmap):**
* Activity is more spread out across the layers compared to the 1.7B model.
* **mlp.down_proj:** High activation in layers 0-15, then decreases.
* **mlp.up_proj:** High activation in layers 0-15, then decreases.
* **self_atten.o_proj:** Moderate activation in layers 0-10.
* **mlp.gate_proj:** Low to moderate activation in layers 0-10.
* **self_atten.v_proj:** Very low activation across all layers.
* **self_atten.q_proj:** Very low activation across all layers.
* **self_atten.k_proj:** Very low activation across all layers.
* **post_attention_layernorm:** Very low activation across all layers.
* **input_layernorm:** Very low activation across all layers.
* **self_atten.k_norm:** Very low activation across all layers.
* **self_atten.a_norm:** Very low activation across all layers.
**Qwen3-8B-Base (Right Heatmap):**
* The activity pattern is similar to the 4B model, but with slightly higher activation in the lower layers.
* **mlp.down_proj:** High activation in layers 0-15, then decreases.
* **mlp.up_proj:** High activation in layers 0-15, then decreases.
* **self_atten.o_proj:** Moderate activation in layers 0-10.
* **mlp.gate_proj:** Low to moderate activation in layers 0-10.
* **self_atten.v_proj:** Very low activation across all layers.
* **self_atten.q_proj:** Very low activation across all layers.
* **self_atten.k_proj:** Very low activation across all layers.
* **post_attention_layernorm:** Very low activation across all layers.
* **input_layernorm:** Very low activation across all layers.
* **self_atten.k_norm:** Very low activation across all layers.
* **self_atten.a_norm:** Very low activation across all layers.
### Key Observations
* The mlp.down_proj and mlp.up_proj modules show the highest activation levels across all three models, particularly in the lower layers.
* The self_atten.o_proj and mlp.gate_proj modules exhibit moderate activation, also concentrated in the lower layers.
* The remaining modules (self_atten.v_proj, self_atten.q_proj, self_atten.k_proj, post_attention_layernorm, input_layernorm, self_atten.k_norm, self_atten.a_norm) show very low activation across all layers and models.
* As the model size increases (1.7B to 4B to 8B), the activity tends to spread out more across the layers, suggesting that larger models utilize more layers for processing.
### Interpretation
The heatmaps provide a visual representation of the internal workings of the Qwen3 models. The concentration of activity in the mlp.down_proj and mlp.up_proj modules suggests that these modules play a crucial role in the model's processing, especially in the initial layers. The low activation of other modules might indicate that they are either less important or that their activity is more distributed and less concentrated in specific layers.
The trend of activity spreading out across more layers as the model size increases suggests that larger models are able to distribute the processing load more effectively, potentially leading to improved performance. The consistent patterns across the different model sizes also indicate that the overall architecture and processing flow remain similar, with the larger models simply scaling up the existing structure.
</details>
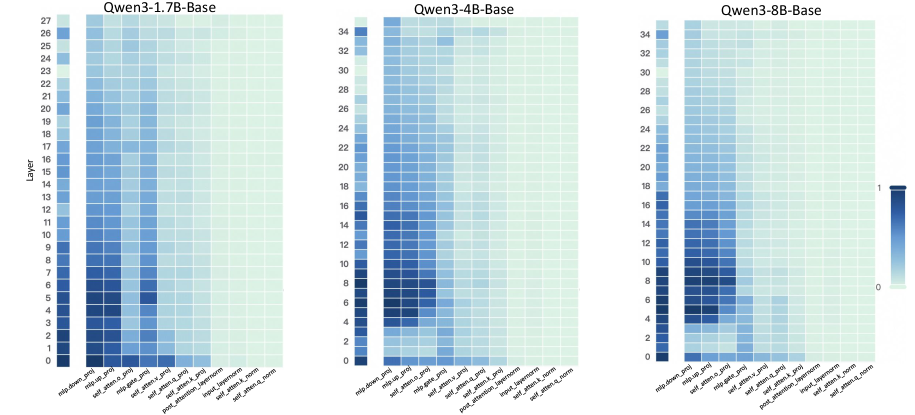
Figure 2: Layer- and unit-level importance distribution of the Qwen3 family. The vertical axis corresponds to different layers, while the horizontal axis denotes parameter units within each layer. Deeper blue indicates higher importance for preserving general-domain competencies.
2 Pilot Study: Probing Parameter Importance
2.1 Experimental Setup for Importance Probing
To investigate the functional specialization of LLMs and understand how different parameters contribute to preserving general-domain knowledge during CPT, we conduct importance probing on multiple backbone models, including Qwen3-Base (1.7B, 4B, 8B) (Yang et al., 2025a) and LLaMA3-8B (Dubey et al., 2024b). Our analyses focus on probing general-knowledge-critical parameters rather than domain-specific ones. The rationale is that successful CPT must inject new, domain-specific knowledge without inducing catastrophic forgetting. This necessitates identifying and preserving the model’s core parameters that are crucial for its general-domain competencies. By contrast, domain knowledge can then be effectively allocated to less critical parameters, without risking the erosion of pre-existing knowledge and skills. To support this analysis, we construct a General Competence Detection Corpus containing broad world knowledge and instruction-following tasks in both English and Chinese, which servers as the probing ground to reflect a model’s general competencies. Details of its construction are provided in Appendix B.3.
2.2 Layer-Level Importance Probing
Our first research question is: How do different layers contribute to preserving general knowledge? To answer this, we measure the importance of each transformer layer by the model’s degradation in general-domain performance when that layer is ablated. Formally, given the General Competence Detection Corpus $\mathcal{D}_{\text{probe}}$ , we first compute the baseline next-token prediction loss of the pretrained LLM $M_{0}$ :
$$
\mathcal{L}_{\text{base}}=\frac{1}{|\mathcal{D}_{\text{probe}}|}\sum_{x\in\mathcal{D}_{\text{probe}}}\ell\big(M_{0}(x),x\big), \tag{1}
$$
where $\ell(·)$ denotes the standard next-token prediction loss in CPT. For each transformer layer $l∈\{1,...,L\}$ , we mask its output via a residual bypass and recompute the loss:
$$
\hat{\mathcal{L}}^{(l)}=\frac{1}{|\mathcal{D}_{\text{probe}}|}\sum_{x\in\mathcal{D}_{\text{probe}}}\ell\big(M_{0}^{(-l)}(x),x\big), \tag{2}
$$
where $M_{0}^{(-l)}$ denotes the model with the $l$ -th layer masked. The importance of layer $l$ is defined as the loss increase relative to the baseline:
$$
I_{\text{layer}}^{(l)}=\hat{\mathcal{L}}^{(l)}-\mathcal{L}_{\text{base}}. \tag{3}
$$
A larger $I_{\text{layer}}^{(l)}$ indicates that layer $l$ plays a more critical role in preserving general knowledge. Figure 2 (left-hand bars) reports the layer-level importance distributions of the Qwen3 family (results for LLaMA3-8B provided in Appendix D). We find that general-knowledge-critical layers are concentrated in the early layers, with importance gradually decreasing toward later layers. This uneven distribution suggests that uniformly expanding layers across the entire depth would be suboptimal. Since some layers are tightly coupled with general knowledge while others are more flexible, uniform expansion not only risks representational interference in critical layers but also allocates parametric budget where it is too constrained to be leveraged for domain learning. In contrast, identifying more adaptable layers with minimal impact on general knowledge and allocating expansion there for knowledge injection is a superior strategy. This leads to our first key observation:
Observation I: Layers exhibit heterogeneous importance for preserving general competencies, which motivates a selective expansion strategy that targets layers less constrained by general abilities yet more adaptable for domain adaptation.
2.3 Unit-Level Importance Probing
Building on the layer-level exploration, our next research question is: How do parameter units within each layer contribute to preserving general knowledge? To answer this, we partition each transformer layer into functional units (e.g., attention projections, MLP components, and normalization) and assess their relative contributions to preserving general competencies. The detailed partitioning scheme is provided in Appendix C. This granularity provides a more fine-grained perspective than layer-level probing, while avoiding the prohibitive cost of neuron-level analysis. Formally, for each parameter $\theta_{j}$ in a unit $U$ , we estimate its importance using a first-order Taylor approximation:
$$
I_{j}=\theta_{j}\cdot\nabla_{\theta_{j}}\mathcal{L}, \tag{4}
$$
where $\mathcal{L}$ is the autoregressive training loss. The importance of unit $U$ is then defined as the average importance of its parameters:
$$
I_{\text{unit}}=\frac{1}{|U|}\sum_{j\in U}I_{j}. \tag{5}
$$
A higher $I_{\text{unit}}$ indicates that the unit plays a more critical role in preserving general competencies. Figure 2 (right-hand heatmaps) illustrates the unit-level importance distributions of the Qwen3 family (results for LLaMA3-8B provided in Appendix D). We observe that importance is unevenly distributed across modules within a layer, with some units contributing more to general competencies and others more flexible. This finding suggests that treating all parameter units equally would be suboptimal, as a single update rule cannot simultaneously protect critical units and fully train adaptable ones, risking either damaging previous knowledge or failing to sufficiently learn new knowledge. This motivates us to pursue unit-level decoupling, where training can selectively protect critical units while enabling less general-relevant units to absorb new knowledge without constraint. This leads to our second key observation:
Observation II: Parameter units within each layer exhibit heterogeneous importance, which motivates unit-level decoupling that selectively protects critical units while enabling more adaptable ones to sufficiently absorb domain knowledge.
Summary. Building on the above observations, we propose ADEPT, a continual pretraining framework designed to enable effective domain knowledge injection while preserving general competencies. Inspired by the uneven importance distribution of layers (Observation I), ADEPT selectively expands layers less constrained by general abilities but more receptive to domain adaptation, thereby introducing fresh parameter capacity rather than uniformly expanding layers as in LLaMA-Pro (Wu et al., 2024b). Guided by the heterogeneous importance of parameter units within layers (Observation II), ADEPT further performs unit-level decoupling on the expanded layers, protecting critical units while enabling adaptable ones to specialize in domain knowledge.
<details>
<summary>x3.png Details</summary>
### Visual Description
## Diagram: General-Competence Guided Selective Layer Expansion and Adaptive Unit-Wise Decoupled Tuning
### Overview
The image presents a two-stage diagram illustrating a method for selective layer expansion and adaptive unit-wise decoupled tuning in a neural network. Stage 1 focuses on identifying important layers and expanding them, while Stage 2 focuses on decoupling and tuning individual units.
### Components/Axes
**Stage 1: General-Competence Guided Selective Layer Expansion**
* **Step 1: General-Competence Aware Layer Importance Probing:**
* **Parameters:** Represented by a brain icon.
* **LLM:** A cartoon avatar wearing a VR headset.
* **General-Competence Detection Corpus:** Represented by a document icon with scribbles.
* **Probing Iteration:** Denoted by 'i'.
* **Layers:** L1, L2, Li, LM, represented as stacked blocks. One layer (Li) is shown as "Deactivated" (greyed out), while another is "Activated".
* **ΔLi:** Change in loss.
* **L:** Loss.
* **Step 2: Selective Expansion via Identity Copy:**
* **General-Competence Importance:** Y-axis of a bar graph.
* **ΔLi:** Change in loss.
* **Select Lowest-K Layers:** Text label.
* **Layer Index:** X-axis of the bar graph.
* **Copy:** Arrows indicating copying of layers.
* **Selective Expansion:** Expanded layers.
* **L(M+k):** Label indicating the copied layer.
**Stage 2: Adaptive Unit-Wise Decoupled Tuning**
* **Step 1: Unit-wise Neuron Decoupling:**
* **General-Competence Detection Corpus:** Represented by a document icon with scribbles.
* **Domain-adaptive Units:** Units highlighted with a star icon.
* **General-critical Units:** Units highlighted with a circle icon.
* **Calculate Unit-wise Importance:** I(wij) = |wij∇wijL|
* Two 3D surface plots, one labeled "Domain-adaptive Units" (red/orange) and the other "General-critical Units" (purple/blue).
* **Step 2: Dynamic Learning Rate Adaptation:**
* **Pretrain Dataset:** Represented by a document icon.
* **Decoupled Tuning:** Text label.
* **Higher Learning Rate:** Indicated by an upward-pointing arrow and a flame icon.
* **Lower Learning Rate:** Indicated by a downward-pointing arrow and a flame icon.
* **Update:** Text label with an arrow.
* Two 3D surface plots, one showing "Higher Learning Rate" (red/orange) and the other "Lower Learning Rate" (purple/blue).
**Legend (Right Side)**
* **Trainable:** Flame icon.
* **Frozen:** Snowflake icon.
* **Identity Copy:** Dashed arrow.
* **Update Flow:** Dotted arrow.
* **Forward Flow:** Solid arrow.
* **Probing:** Magnifying glass icon.
* **Domain Units:** Star icon.
* **General Units:** Circle icon.
* **Original Layer:** Light brown rectangle.
* **Masked Layer:** Grey rectangle.
* **Expanded Layer:** Red rectangle.
* **L:** Next Token Prediction Loss.
### Detailed Analysis
**Stage 1:**
* The process starts with "Parameters" and "General-Competence Detection Corpus" feeding into an LLM.
* The LLM probes the layers (L1 to LM) to determine their importance.
* The probing iteration is denoted by 'i'.
* Layers are either "Deactivated" or "Activated" based on the probing.
* The "Selective Expansion via Identity Copy" step involves selecting the lowest-K layers based on their importance (ΔLi).
* These selected layers are then copied and expanded.
**Stage 2:**
* The "Unit-wise Neuron Decoupling" step involves calculating the importance of individual units within the layers.
* The formula for calculating unit-wise importance is given as I(wij) = |wij∇wijL|.
* The "Dynamic Learning Rate Adaptation" step adjusts the learning rate for each unit based on its importance.
* Units with higher importance receive a higher learning rate, while units with lower importance receive a lower learning rate.
### Key Observations
* The diagram highlights a two-stage process for optimizing neural networks.
* Stage 1 focuses on identifying and expanding important layers, while Stage 2 focuses on decoupling and tuning individual units.
* The use of visual cues, such as icons and arrows, helps to illustrate the flow of information and the different steps involved in the process.
* The 3D surface plots in Stage 2 provide a visual representation of the learning rate adaptation process.
### Interpretation
The diagram illustrates a method for improving the efficiency and effectiveness of neural network training. By selectively expanding important layers and adaptively tuning individual units, the network can learn more quickly and achieve better performance. The method leverages the concept of "General-Competence" to guide the selection and tuning process, ensuring that the network is optimized for a specific task or domain. The use of decoupled tuning allows for more fine-grained control over the learning process, enabling the network to adapt to the specific characteristics of the data. The diagram suggests a sophisticated approach to neural network optimization that combines layer-level and unit-level techniques.
</details>
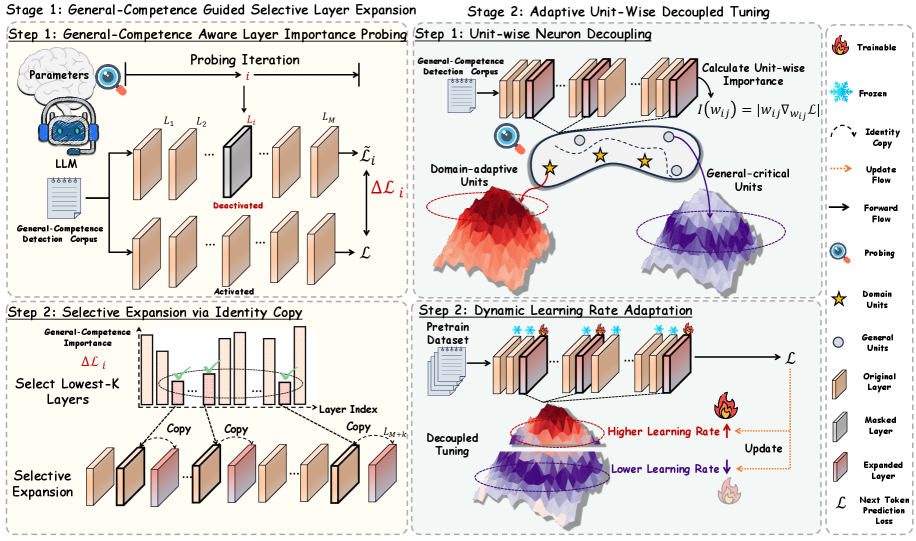
Figure 3: Illustration of ADEPT.
3 Methodology
As illustrated in Figure 3, ADEPT includes two stages:
- # Stage 1: General-Competence Guided Selective Layer Expansion. adaptively selects and duplicates layers that minimally affect general competencies while being more adaptable to domain-specific knowledge, thereby introducing fresh representational capacity for domain adaptation.
- # Stage 2: Adaptive Unit-Wise Decoupled Tuning. further decouples units within the expanded layers and apply learning-rate–driven adaptive tuning according to their importance to the general domain, ensuring knowledge injection while preserving general competencies.
3.1 General-Competence Guided Selective Layer Expansion
This stage aims to selectively expand model parameters in a way that introduces fresh representational capacity for domain adaptation while preserving general-domain competencies. To this end, we first estimate the contribution of each transformer layer to preserving general knowledge through General-Competence Aware Layer Importance Probing, and then perform Selective Parameter Expansion via Identity Copy to duplicate layers that are least critical for general abilities yet more adaptable to domain-specific knowledge.
General-Competence Aware Layer Importance Probing. To guide selective expansion, we leverage the layer importance scores $I_{\text{layer}}^{(l)}$ defined as Eq. 3. Intuitively, $I_{\text{layer}}^{(l)}$ quantifies how much the $l$ -th layer contributes to preserving general-domain knowledge. Layers with lower scores are deemed less critical for general competencies and are thus selected for expansion, as they can accommodate domain-specific adaptation with minimal risk of catastrophic forgetting.
Selective Parameter Expansion via Identity Copy. Based on the importance scores $I_{\text{layer}}^{(l)}$ , we sort layers by ascending importance and select the $k$ least-important ones for general competence:
$$
\mathcal{S}_{k}=\operatorname*{arg\,min}_{\begin{subarray}{c}\mathcal{S}\subseteq\{1,\ldots,L\}\\
|\mathcal{S}|=k\end{subarray}}\sum_{l\in\mathcal{S}}I_{\text{layer}}^{(l)}. \tag{6}
$$
We denote the selected set $\mathcal{S}_{k}$ as the Domain-Adaptable Layers. For each selected layer $l∈\mathcal{S}_{k}$ , we create a parallel copy by directly duplicating its parameters without re-initialization ( $\tilde{\Theta}^{(l)}=\Theta^{(l)}$ ). To preserve stability, we follow the Function Preserving Initialization (FPI) principle (Chen et al., 2015), ensuring that the expanded model $M_{1}$ produces identical outputs as the original model $M_{0}$ at initialization. Concretely, in the duplicated branch, we set the output projections of both attention and feed-forward sublayers to zero ( $W_{\text{MHSA}}^{\text{out}}=0,\,W_{\text{FFN}}^{\text{out}}=0$ ), so the forward computation remains unchanged ( $M_{1}(x)=M_{0}(x),\,∀ x$ ). The duplicated layers thus provide fresh representational capacity that can specialize for domain signals with minimal risk of eroding general-knowledge-critical parameters in the original pathway. As formally established in Appendix F.1, expanding the layers with the lowest general-competence importance provably minimizes the risk of forgetting. Intuitively, this strategy ensures that new capacity is added where interference with general abilities is weakest, yielding the most favorable trade-off between domain adaptation and knowledge retention.
3.2 Adaptive Unit-Wise Decoupled Tuning
This stage aims to further reduce catastrophic forgetting and enable fine-grained control over parameters within the expanded layers. To achieve this, we first decouple each expanded layer into semantic units and evaluate their importance using gradient-based estimation (Unit-wise Neuron Decoupling), and then dynamically adjust learning rates for different units according to their importance scores during training (Dynamic Learning Rate Adaptation).
Unit-wise Neuron Decoupling. Guided by the heterogeneous importance of parameter units within layers, we performs unit-level decoupling on the expanded layers. Following the probing analysis in Section 2.3, we quantify unit importance $I_{\text{unit}}$ using gradient sensitivity signals (cf. Eq. 5), which aggregate the first-order contributions of parameters $\theta_{j}$ to the training loss $\mathcal{L}$ via $∇_{\theta_{j}}\mathcal{L}$ . A higher $I_{\text{unit}}$ indicates greater contribution to general competencies and thus warrants more conservative updates, whereas less important units are encouraged to adapt more aggressively to domain-specific signals.
Dynamic Learning Rate Adaptation. Based on the unit importance $I_{\text{unit}}$ in Eq. 5, we assign adaptive learning rates to different units within the expanded layers:
$$
\text{lr}_{U}=2\cdot(1-I_{\text{unit}})\cdot\text{lr}_{\text{base}}, \tag{7}
$$
where $\text{lr}_{\text{base}}$ is the base learning rate, and the coefficient $2$ normalizes the global scale to keep the effective average approximately unchanged. Units more important for general knowledge (higher $I_{\text{unit}}$ ) receive smaller learning rates to reduce overwriting, while less important units are encouraged to adapt more aggressively to domain-specific data. Training proceeds with the standard autoregressive objective: $\mathcal{L}=-\sum_{t=1}^{T}\log P(x_{t}\mid x_{<t};\Theta)$ . Since the importance of units may change as training progresses, we periodically recompute $I_{\text{unit}}$ and update learning rates accordingly, ensuring dynamic adaptation throughout learning. The full training procedure is provided in Appendix L. Appendix F.2 further shows that allocating learning rates inversely to unit importance minimizes an upper bound on general-domain forgetting. In essence, this design formalizes the intuition that highly general-critical units should be preserved via conservative updates, while less critical yet more adaptable ones can update more aggressively to absorb domain-specific information.
Table 1: Performance comparison across Mathematical and Medical domains. Bold numbers indicate the best performance, and underlined numbers denote the second best.
| Method | Mathematics | Medical | | | | | | | | |
| --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- |
| General | Domain | General | Domain | | | | | | | |
| MMLU | CMMLU | GSM8K | ARC-Easy | ARC-Challenge | MMLU | CMMLU | MedQA | MMCU-Medical | CMB | |
| Qwen3-1.7B-Base | | | | | | | | | | |
| Vanilla | 62.57 | 66.86 | 57.62 | 81.44 | 51.19 | 62.57 | 66.86 | 48.39 | 69.17 | 63.67 |
| PT-Full | 60.07 | 62.84 | 51.86 | 81.24 | 49.65 | 59.44 | 62.84 | 48.45 | 67.45 | 62.77 |
| Replay | 60.69 | 63.52 | 54.74 | 81.01 | 49.73 | 60.52 | 63.85 | 49.00 | 67.32 | 62.20 |
| Llama-Pro | 61.54 | 63.40 | 60.03 | 81.08 | 49.80 | 59.80 | 65.51 | 50.43 | 66.51 | 63.54 |
| PT-LoRA | 60.07 | 62.69 | 59.50 | 80.22 | 49.34 | 57.31 | 59.68 | 47.29 | 61.55 | 57.60 |
| TaSL | 60.34 | 62.95 | 59.07 | 79.76 | 48.89 | 62.48 | 66.14 | 47.06 | 67.62 | 61.15 |
| ADEPT | 62.62 | 67.06 | 70.51 | 82.48 | 52.62 | 62.80 | 66.89 | 50.75 | 71.98 | 65.43 |
| Qwen3-4B-Base | | | | | | | | | | |
| Vanilla | 73.19 | 77.92 | 69.07 | 85.52 | 59.13 | 73.19 | 77.92 | 62.77 | 82.44 | 78.92 |
| PT-Full | 70.33 | 73.07 | 60.96 | 85.31 | 57.59 | 69.48 | 72.77 | 62.84 | 81.34 | 76.88 |
| Replay | 70.46 | 73.72 | 63.91 | 85.06 | 57.68 | 70.74 | 73.81 | 63.55 | 80.60 | 76.74 |
| Llama-Pro | 72.42 | 77.39 | 73.16 | 85.14 | 57.76 | 72.28 | 77.28 | 62.53 | 81.20 | 78.12 |
| PT-LoRA | 70.20 | 72.90 | 71.34 | 84.18 | 57.25 | 72.73 | 76.78 | 61.59 | 80.49 | 76.92 |
| TaSL | 70.50 | 73.20 | 70.84 | 83.68 | 56.75 | 73.03 | 77.08 | 60.99 | 79.20 | 77.08 |
| ADEPT | 73.21 | 78.30 | 76.19 | 88.44 | 60.98 | 72.95 | 78.77 | 64.49 | 84.58 | 79.87 |
| Qwen3-8B-Base | | | | | | | | | | |
| Vanilla | 76.94 | 82.09 | 69.98 | 87.12 | 64.25 | 76.94 | 82.09 | 66.30 | 86.45 | 81.67 |
| PT-Full | 74.90 | 78.49 | 80.21 | 85.90 | 61.77 | 74.06 | 78.82 | 67.24 | 87.69 | 85.27 |
| Replay | 75.19 | 78.92 | 81.12 | 85.98 | 62.37 | 74.51 | 78.86 | 68.89 | 86.66 | 84.73 |
| Llama-Pro | 76.16 | 81.42 | 80.97 | 86.62 | 63.91 | 76.58 | 81.69 | 66.77 | 87.19 | 83.76 |
| PT-LoRA | 75.66 | 80.81 | 82.87 | 86.36 | 62.46 | 76.60 | 81.57 | 67.01 | 86.70 | 83.04 |
| TaSL | 76.63 | 80.37 | 80.54 | 84.81 | 59.09 | 76.42 | 81.86 | 66.51 | 86.20 | 82.54 |
| ADEPT | 76.80 | 82.11 | 83.87 | 89.29 | 64.51 | 76.77 | 82.11 | 69.24 | 89.84 | 85.80 |
| Llama3-8B-Base | | | | | | | | | | |
| Vanilla | 65.33 | 50.83 | 36.84 | 84.18 | 54.01 | 65.33 | 50.83 | 58.91 | 46.29 | 35.61 |
| PT-Full | 61.62 | 46.21 | 49.73 | 84.01 | 53.52 | 59.15 | 51.39 | 59.23 | 66.58 | 61.65 |
| Replay | 62.00 | 53.31 | 49.51 | 82.49 | 54.18 | 59.98 | 54.52 | 59.07 | 65.84 | 61.71 |
| Llama-Pro | 64.53 | 50.26 | 48.29 | 83.29 | 53.07 | 64.19 | 50.59 | 59.94 | 53.96 | 47.05 |
| PT-LoRA | 64.86 | 49.82 | 48.82 | 83.80 | 54.01 | 64.34 | 50.13 | 58.84 | 56.05 | 48.22 |
| TaSL | 65.16 | 50.11 | 35.43 | 83.29 | 53.51 | 64.64 | 50.43 | 55.55 | 58.34 | 47.69 |
| ADEPT | 65.35 | 51.90 | 50.57 | 84.96 | 55.52 | 65.17 | 51.92 | 61.17 | 67.03 | 61.78 |
4 Experiment
4.1 Experimental Setup
Datasets. We evaluate ADEPT across two domains, Mathematics and Medicine. For the mathematical domain, we use OpenWebMath (Paster et al., 2023), together with AceReason-Math (Chen et al., 2025), concatenated into the continual pretraining corpora. For the medical domain, we adopt the multilingual MMedC corpus (Qiu et al., 2024), together with IndustryIns and MMedBench, forming the medical pretraining corpora. Dataset statistics are provided in Appendix B.1 and Appendix B.2. In addition, for detecting general-knowledge-critical regions, we construct a General Competence Detection Corpus, following the same setting as in Section 2 and described in Appendix B.3.
Baselines. We compare ADEPT with a broad range of baselines from four perspectives:
- Full-parameter tuning. PT-Full directly updates all model parameters on the target corpora.
- Replay-based tuning. Replay mitigates catastrophic forgetting by mixing general-domain data into the training process (Que et al., 2024).
- Architecture expansion. LLaMA-Pro (Wu et al., 2024b) expands the model by uniformly inserting new layers into each transformer block while freezing the original weights. Only the newly introduced parameters are trained, enabling structural growth while preserving prior knowledge.
- Parameter-efficient tuning. PT-LoRA performs CPT using Low-Rank Adaptation (Hu et al., 2022), updating only a small set of task-adaptive parameters. TaSL (Feng et al., 2024a) extends PT-LoRA to a multi-task regime by decoupling LoRA matrices across transformer layers, allowing different subsets of parameters to specialize for different tasks.
See Appendix B.6 for implementation details of all baselines.
Backbone Models. To assess the generality of our method, we instantiate ADEPT on multiple backbone models, including Qwen3-Base (1.7B, 4B, 8B) (Yang et al., 2025a) and LLaMA3.1-8B-Base (Dubey et al., 2024b), covering a wide range of parameter scales and architectural variants.
Evaluation Metrics and Strategy. We adopt multiple-choice question answering accuracy as the primary evaluation metric across all tasks (see Appendix B.9 for further details). For the Mathematics domain, we evaluate on GSM8K (Cobbe et al., 2021), ARC-Easy (Clark et al., 2018), and ARC-Challenge (Clark et al., 2018), which collectively span a wide range of reasoning difficulties. For the Medical domain, we use MedQA (Jin et al., 2021), MMCU-Medical (Zeng, 2023), and CMB (Wang et al., 2023b), covering diverse medical subjects and varying levels of complexity. Among them, MedQA is an English benchmark, while MMCU-Medical and CMB are in Chinese. To assess the model’s ability to retain general-domain knowledge during continual pretraining, we additionally evaluate on MMLU (Hendrycks et al., 2020) and CMMLU (Li et al., 2023), two broad-coverage benchmarks for general knowledge and reasoning in English and Chinese, respectively.
4.2 Experimental Results
Performance Comparison. As shown in Table 1, ADEPT consistently outperforms all CPT baselines across both mathematical and medical domains, confirming its effectiveness in domain-specific knowledge acquisition while substantially alleviating catastrophic forgetting. Concretely, ADEPT achieves substantial domain-specific improvements. Across all backbones and domain benchmarks, ADEPT consistently surpasses baselines, achieving the strongest performance. For instance, on Qwen3-1.7B-Base, ADEPT boosts GSM8K accuracy from 57.62% to 70.51% $\uparrow$ , bringing a large gain that highlights its advantage on enhancing LLMs’ complex reasoning. Similarly, on LLaMA3-8B-Base, it drastically improves CMB accuracy improves from 35.61% to 61.78% $\uparrow$ , underscoring the strong enhancement of medical-domain capabilities. On average, ADEPT achieves up to 5.58% gains over full-parameter CPT on target-domain benchmarks, confirming its advantage in domain knowledge acquisition. Furthermore, ADEPT demonstrates clear advantages in mitigating catastrophic forgetting. Whereas most baselines suffer noticeable degradation on general benchmarks such as MMLU and CMMLU, ADEPT preserves the pretrained LLMs’ general-domain competencies, and in some cases even surpasses the vanilla backbone. Notably, with Qwen3-4B under medical CPT, ADEPT improves CMMLU accuracy from 77.92% to 78.77% $\uparrow$ . It also results in an average performance increase of 5.76% on general benchmarks over full-parameter CPT. We attribute this to the disentanglement of domain-specific and general parameters, which prevents harmful representational interference during adaptation, ensuring that learning specialized knowledge does not corrupt the model’s foundational abilities. Instead, this focused learning process appears to refine the model’s overall competencies, leading to synergistic improvements on general-domain tasks. In summary, ADEPT offers a robust solution for CPT achieving superior domain adaptation while effectively preserving general knowledge.
Table 2: Ablation study on ADEPT in Medical domain. Bold numbers indicate the best performance, and underlined numbers denote the second best.
| Method | Qwen3-1.7B-Base | Llama3-8B-Base | | | | | | | | |
| --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- |
| MMLU | CMMLU | MedQA | MMCU-Medical | CMB | MMLU | CMMLU | MedQA | MMCU-Medical | CMB | |
| ADEPT | 62.80 | 66.89 | 50.75 | 70.98 | 65.43 | 65.17 | 51.92 | 61.17 | 61.78 | 67.03 |
| w/o Stage-1 | 57.31 | 59.68 | 47.29 | 61.55 | 57.60 | 57.88 | 50.76 | 58.32 | 53.32 | 60.32 |
| w/o Stage-2 | 61.56 | 64.33 | 49.23 | 66.19 | 64.36 | 64.34 | 50.74 | 59.60 | 50.68 | 57.36 |
| Uniform Expansion | 59.80 | 65.51 | 50.43 | 66.51 | 63.54 | 64.19 | 50.59 | 59.94 | 47.05 | 53.96 |
Ablation Study. To investigate the effectiveness of each component in ADEPT, we conduct ablation experiments in the medical domain using two representative backbones, Qwen3-1.7B and Llama3-8B. In w/o Stage-1, we remove the General-Competence Guided Selective Layer Expansion and directly apply Adaptive Unit-Wise Decoupled Tuning on the $k$ Domain-Adaptable Layers without introducing any new parameters. In w/o Stage-2, we discard the dynamic decoupled tuning stage and instead directly fine-tune the expanded layers from Stage-1. In Uniform Expansion, we replace importance-guided expansion with uniformly inserted layers followed by fine-tuning, which is equivalent to the strategy adopted in LLaMA-Pro. As shown in Table 2, removing either Stage-1 or Stage-2 leads to clear degradation in both general and domain-specific performance, confirming that both adaptive expansion and decoupled tuning are indispensable. In particular, eliminating Stage-1 results in the largest performance drop, suggesting that adaptive capacity allocation is crucial for enabling effective domain adaptation without sacrificing general-domain competencies. Meanwhile, replacing importance-guided expansion with uniform expansion yields inferior results, underscoring the advantage of expanding only the most domain-adaptable layers.
<details>
<summary>x4.png Details</summary>
### Visual Description
## Density Contour Plots: Comparing Text Embeddings
### Overview
The image presents three sets of density contour plots, each comparing the distribution of "General Text" and "Medical Text" embeddings in a two-dimensional space. The plots are arranged horizontally, labeled "Vanilla," "W/o stage1," and "ADEPT." Each plot includes a 2D contour plot, a density plot above it representing the distribution along the horizontal axis (dim 1), and a density plot to the right representing the distribution along the vertical axis (dim 2). The plots aim to visualize how different embedding methods separate general and medical text.
### Components/Axes
* **Titles:** "Vanilla," "W/o stage1," "ADEPT" (placed above each respective plot).
* **Legends:** Each plot has a legend in the top-left corner indicating:
* "General Text" (represented by the color blue)
* "Medical Text" (represented by the color red)
* **Axes Labels:**
* Horizontal axis (dim 1): Ranges from approximately -75 to 75 for "Vanilla", -100 to 100 for "W/o stage1", and -100 to 100 for "ADEPT".
* Vertical axis (dim 2): Ranges from approximately -75 to 100 for "Vanilla", -60 to 60 for "W/o stage1", and -60 to 60 for "ADEPT".
* **Axis Markers:**
* Vanilla: dim 1: -50, 0, 50. dim 2: -75, -50, -25, 0, 25, 50, 75, 100
* W/o stage1: dim 1: -100, -50, 0, 50, 100. dim 2: -60, -40, -20, 0, 20, 40, 60
* ADEPT: dim 1: -100, -50, 0, 50. dim 2: -60, -40, -20, 0, 20, 40, 60
### Detailed Analysis
**1. Vanilla:**
* **2D Contour Plot:**
* "General Text" (blue): The contours form a cluster centered around dim 1 = -25 and dim 2 = 25. The distribution is somewhat elongated along a diagonal.
* "Medical Text" (red): The contours form a cluster centered around dim 1 = 25 and dim 2 = -10. The distribution is more spread out compared to "General Text."
* **Horizontal Density Plot (dim 1):**
* "General Text" (blue): Shows a single peak around -25.
* "Medical Text" (red): Shows a single peak around 25.
* **Vertical Density Plot (dim 2):**
* "General Text" (blue): Shows a single peak around 25.
* "Medical Text" (red): Shows a single peak around -10.
**2. W/o stage1:**
* **2D Contour Plot:**
* "General Text" (blue): The contours are more dispersed, with a primary cluster around dim 1 = -25 and dim 2 = 10, and a secondary cluster around dim 1 = 25 and dim 2 = -20.
* "Medical Text" (red): The contours are also dispersed, with a primary cluster around dim 1 = 25 and dim 2 = 10, and a secondary cluster around dim 1 = -25 and dim 2 = -20. There is significant overlap between the two distributions.
* **Horizontal Density Plot (dim 1):**
* "General Text" (blue): Shows two peaks, one around -25 and another around 25.
* "Medical Text" (red): Shows two peaks, one around -25 and another around 25.
* **Vertical Density Plot (dim 2):**
* "General Text" (blue): Shows two peaks, one around 10 and another around -20.
* "Medical Text" (red): Shows two peaks, one around 10 and another around -20.
**3. ADEPT:**
* **2D Contour Plot:**
* "General Text" (blue): The contours form a cluster centered around dim 1 = -40 and dim 2 = 20.
* "Medical Text" (red): The contours form a cluster centered around dim 1 = 40 and dim 2 = -20. The separation between the two distributions is more pronounced than in the other two plots.
* **Horizontal Density Plot (dim 1):**
* "General Text" (blue): Shows a single peak around -40.
* "Medical Text" (red): Shows a single peak around 40.
* **Vertical Density Plot (dim 2):**
* "General Text" (blue): Shows a single peak around 20.
* "Medical Text" (red): Shows a single peak around -20.
### Key Observations
* The "Vanilla" plot shows some separation between "General Text" and "Medical Text," but there is still some overlap.
* The "W/o stage1" plot shows significant overlap between the two distributions, indicating poor separation.
* The "ADEPT" plot shows the best separation between "General Text" and "Medical Text," with distinct clusters for each.
### Interpretation
The plots visualize the effectiveness of different embedding methods in distinguishing between general and medical text. The "ADEPT" method appears to be the most effective, as it produces the most distinct clusters for the two types of text. Removing "stage1" results in the worst separation. The "Vanilla" method provides an intermediate level of separation. This suggests that the "ADEPT" method is better at capturing the semantic differences between general and medical text, which could be beneficial for various natural language processing tasks in the medical domain. The overlap in "W/o stage1" suggests that stage 1 is critical for separating the two types of text.
</details>
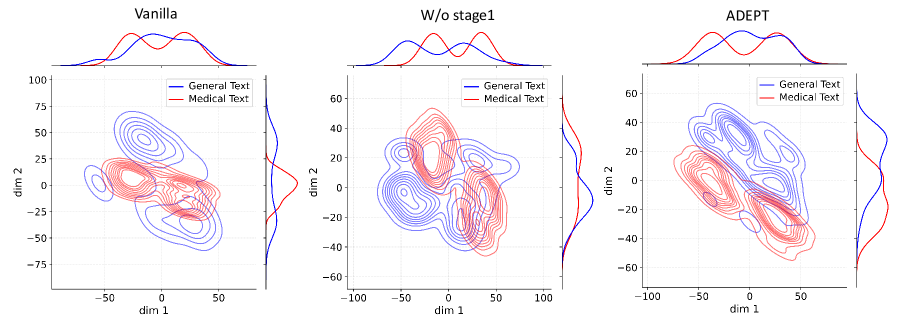
Figure 4: Activation distribution analysis of Qwen3-8B.
Decoupling Effectiveness on Expanded Parameters. We visualize cross-domain activations using Kernel Density Estimation (KDE) (Silverman, 2018), sampling 500 instances from both Medical and General corpora. For the original Qwen3-8B-Base (left in Figure 4), the most domain-adaptable layer (lowest $I_{\text{layer}}$ ) still shows heavy overlap between general and medical activations, evidencing strong parameter coupling. Direct decoupling without expansion (w/o Stage-1, middle) on the same layer fails to reduce this entanglement, confirming that pretrained parameters are inherently difficult to separate. In contrast, after expansion (right), the duplicated layers serve as a “blank slate,” yielding clearly separated activations across domains. Additional analyses on more backbones are provided in Appendix C.1, where we observe that this trend consistently holds across nearly all evaluated LLMs, further validating the generality of our approach.
<details>
<summary>figures/wordcloud_med.jpg Details</summary>
### Visual Description
## Word Cloud: Medical Research Terms
### Overview
The image is a word cloud, visually representing the frequency of different terms related to medical research. The size of each word corresponds to its frequency or importance within the context from which the words were extracted. The words are arranged randomly and colored in shades of green and blue.
### Components/Axes
* **Words:** The primary components are the individual words themselves, such as "Health," "Patient," "Drug," "Treatment," "Disease," etc.
* **Size:** The size of each word indicates its relative frequency or importance. Larger words appear more frequently.
* **Color:** The words are colored in shades of green and blue.
### Detailed Analysis or ### Content Details
Here's a breakdown of the words and their relative prominence based on size:
* **Largest Words:**
* "Health" (Green): One of the most prominent words, suggesting a strong focus on general health-related topics.
* "Patient" (Green): Indicates a significant emphasis on patient-related research.
* "Treatment" (Blue): Highlights the importance of treatment methodologies.
* "Drug" (Blue): Suggests a focus on pharmaceutical interventions.
* **Large Words:**
* "Care" (Blue): Indicates a focus on healthcare services.
* "Hospital" (Green): Suggests research related to hospital settings.
* "Disease" (Green): Indicates a focus on specific diseases.
* "Medication" (Green): Highlights the importance of pharmaceutical interventions.
* "Heart" (Blue): Suggests a focus on cardiovascular health.
* "Diabetes" (Blue): Indicates a focus on diabetes research.
* "Symptoms" (Blue): Highlights the importance of symptom identification.
* "Therapy" (Blue): Suggests a focus on therapeutic interventions.
* "Brain" (Blue): Indicates a focus on neurological research.
* "Tumor" (Green): Highlights the importance of cancer research.
* "Prescription" (Blue): Suggests a focus on pharmaceutical interventions.
* "Randomized" (Blue): Indicates a focus on randomized controlled trials.
* **Medium-Sized Words:**
* "Dental" (Green): Indicates a focus on dental health.
* "Medical" (Green): Suggests research related to medical settings.
* "RNA" (Green): Highlights the importance of RNA research.
* "Medicine" (Yellow): Indicates a focus on medical settings.
* "Results" (Green): Highlights the importance of research outcomes.
* "MRI" (Green): Suggests research related to medical imaging.
* **Smaller Words:**
* "FDA" (Blue): Indicates a focus on regulatory compliance.
* "GDPR" (Green): Indicates a focus on data privacy.
* "Research" (Green): Highlights the importance of research activities.
* "Arthritis" (Green): Suggests research related to arthritis.
* "Concentrations" (Green): Indicates a focus on chemical concentrations.
* "Cases" (Blue): Indicates a focus on case studies.
* "Illness" (Green): Suggests research related to illness.
* "Protein" (Green): Highlights the importance of protein research.
* "LOS" (Green): Length of Stay
* "Best" (Green): Indicates a focus on best practices.
* "MD" (Blue): Indicates a focus on medical doctors.
* "DICE" (Blue): Indicates a focus on dice research.
* "Cancer" (Green): Suggests research related to cancer.
* "Healthy" (Green): Indicates a focus on healthy lifestyles.
* "Investigator" (Green): Highlights the importance of research investigators.
* "CDC" (Green): Indicates a focus on the Centers for Disease Control and Prevention.
* "Warning" (Green): Highlights the importance of safety warnings.
* "Data" (Blue): Indicates a focus on data analysis.
* "Diagnosed" (Green): Highlights the importance of diagnosis.
* "Drugs" (Green): Suggests a focus on pharmaceutical interventions.
* "Stay" (Green): Indicates a focus on patient stay.
* "Study" (Green): Highlights the importance of research studies.
* "Significant" (Green): Indicates a focus on significant findings.
* "Effects" (Green): Highlights the importance of research effects.
* "Mental" (Blue): Indicates a focus on mental health.
* "Addiction" (Green): Suggests research related to addiction.
* "Diseases" (Green): Indicates a focus on specific diseases.
* "Time" (Green): Highlights the importance of time-related factors.
* "Vaccine" (Blue): Suggests research related to vaccines.
* "Hospitals" (Green): Indicates a focus on hospital settings.
* "Photograph" (Green): Highlights the importance of photographic documentation.
* "Effect" (Green): Highlights the importance of research effects.
* "Role" (Green): Indicates a focus on roles in healthcare.
* "Blood" (Green): Indicates a focus on blood-related research.
* "News" (Green): Highlights the importance of news dissemination.
* "Serious" (Green): Indicates a focus on serious conditions.
* "Researchers" (Green): Highlights the importance of research personnel.
* "Ways" (Green): Indicates a focus on different approaches.
* "Lot" (Green): Indicates a focus on lot sizes.
* "Of" (Green): Indicates a focus on relationships.
* "You" (Green): Indicates a focus on patient involvement.
* "Cell" (Green): Indicates a focus on cellular research.
* "Leading" (Green): Indicates a focus on leading research.
* "MS" (Blue): Indicates a focus on multiple sclerosis.
* "Kidney" (Green): Indicates a focus on kidney health.
* "Emergency" (Green): Indicates a focus on emergency care.
* "Dementia" (Green): Indicates a focus on dementia research.
* "Ildenafil" (Yellow): Indicates a focus on ildenafil research.
* "Diagnosis" (Green): Highlights the importance of diagnosis.
* "Izaion" (Green): Indicates a focus on ionization research.
* "UE" (Green): Indicates a focus on user experience.
* "Po" (Green): Indicates a focus on patient outcomes.
* "FS" (Green): Indicates a focus on financial services.
* "Most" (Green): Indicates a focus on most common conditions.
* "Doctor" (Green): Indicates a focus on doctor involvement.
* "Pain" (Green): Indicates a focus on pain management.
* "Ynec" (Yellow): Indicates a focus on gynecological research.
### Key Observations
* The word cloud emphasizes the importance of "Health," "Patient," "Treatment," and "Drug" in medical research.
* There is a significant focus on specific diseases like "Diabetes" and "Cancer."
* The inclusion of terms like "FDA" and "GDPR" suggests an awareness of regulatory and privacy considerations.
### Interpretation
The word cloud provides a snapshot of the key themes and areas of focus within medical research. The prominence of terms like "Health," "Patient," and "Treatment" underscores the patient-centric nature of the field. The presence of disease-specific terms indicates targeted research efforts, while the inclusion of regulatory terms highlights the importance of ethical and legal considerations. The word cloud suggests a field that is both broad in scope and highly specialized, with a strong emphasis on improving patient outcomes and advancing medical knowledge.
</details>
(a) Token distributions shift in Medical
<details>
<summary>figures/wordcloud_math.jpg Details</summary>

### Visual Description
## Word Cloud: Research Topics
### Overview
The image is a word cloud, visually representing the frequency of different terms. The size of each word corresponds to its frequency or importance. The words are related to research, academia, and scientific topics.
### Components/Axes
There are no axes in a word cloud. The key component is the collection of words themselves, with their relative sizes indicating frequency.
### Detailed Analysis or ### Content Details
Here's a transcription of the words, grouped loosely by location and size (largest to smallest):
* **Left Side:**
* algorithm
* students
* paper
* user
* talk
* scripts
* Oct
* problem
* sense
* udos
* de
* **Center:**
* mathematics
* space
* effects
* following
* editor
* physics
* answer
* previous
* quantum
* parameters
* model
* bit
* reaction
* identity
* theorem
* isation
* **Right Side:**
* question
* PhD
* results
* Rig
* value
* open
* profit
* error
* features
* code
* al
* ule
* to
* A
* chner
* case
* **Other words:**
* PE
* phenotype
* ob
* variance
* scatter
* et
* F
* Tony
* Sized
* ja
* ric
* ormal
* esium
* pton
* papers
* $$
* AG
* sh
* connected
* quant
* new
* ex
* CM
* the
* FT
* ATE
* he
* groups
* {
* }
* |
* b1
* 2
* qu
* $?$
### Key Observations
* The largest words include "algorithm," "mathematics," "space," "effects," "following," "editor," "physics," "answer," "previous," "quantum," "question," "PhD," and "results."
* There are some symbols and abbreviations scattered throughout, such as "$", "ex", "PE", "CM", "FT", "ATE", "AG", "sh", "b1", "2", "{", "}", "|", and "?".
* The words are arranged in a seemingly random fashion, typical of word clouds.
### Interpretation
The word cloud provides a visual summary of the topics most frequently discussed or emphasized in a given context (likely a collection of research papers, articles, or discussions). The prominence of terms like "algorithm," "mathematics," "physics," and "quantum" suggests a focus on quantitative and scientific research. The presence of words like "question," "results," and "PhD" indicates an academic or research-oriented environment. The cloud highlights the key themes and areas of interest within the source material.
</details>
(b) Token distributions shift in Mathematical
Figure 5: Token distribution shifts across domains. Word cloud visualizations of shifted tokens reveal that ADEPT achieves highly focused alignment, with most changes concentrated on domain-specific terminology.
Token Distribution Shift Analysis. To assess how ADEPT injects domain knowledge while preserving general competencies, we analyze token-level shifts between the base and continually pretrained models. Following Lin et al. (2024), tokens are categorized as unshifted, marginal, or shifted. Only a small proportion of tokens shift, while most remain unchanged, indicating stable adaptation. In the medical domain, merely 2.18% shift (vs. 5.61% under full pretraining), largely medical terms such as “prescription,” “diagnosis,” and “therapy” (Figure 5(a)). In the mathematical domain, only 1.24% shift, mainly scientific terms such as “theorem” and “equation” (Figure 5(b)). Further details and analyses are provided in Appendix I. These results demonstrate that ADEPT achieves precise and economical domain knowledge injection while minimizing perturbation to general competence.
Extended Investigations and Key Insights. We further investigate several design choices of ADEPT in appendix: In Appendix E, we investigate alternative strategies for probing layer importance and observe the consistency of different measurement methods, offering insight into how importance estimation affects adaptation outcomes. Appendix G explores the effect of expanding different numbers of layers and reveals how the number of expansion layers should be selected under different circumstances and the potential reasons behind this. Appendix H shows that even with relatively low-quality importance detection corpus from pretrain data, our approach maintains strong generalization across domains, suggesting the robustness of ADEPT. Appendix J demonstrates our insights into the potential for merging expanded layers that are independently trained on different domains, offering an intriguing direction for achieving multi-domain adaptation with minimal catastrophic forgetting. In addition, Appendix B.8 analyzes the training efficiency of ADEPT, showing that our selective updating design substantially accelerates convergence compared to baselines.
5 Conclusions and Future Works
We present ADEPT, a framework for LLM continual pretraining for domain adaptation that effectively tackles catastrophic forgetting, leveraging functional specialization in LLMs. By selectively expanding layers less critical to the general domain and adaptively updating decoupled parameter units, ADEPT minimizes catastrophic forgetting while efficiently incorporating domain-specific expertise. Our experiments show significant improvements in both domain performance and general knowledge retention compared to baselines. Future work could focus on refining the decoupled tuning mechanism, designing more sophisticated learning rate strategies beyond linear mapping to allow for more precise adjustments. Another direction is to explore better dynamic and real-time methods for measuring parameter importance during training.
6 Ethics Statement
All datasets used for training and evaluation in this study are publicly available versions obtained from the Hugging Face platform. The datasets have been curated, cleaned, and de-identified by their respective data providers prior to release. No patient personal information or identifiable medical data is present. Consequently, the research does not involve human subjects, and there are no related concerns regarding privacy, confidentiality, or legal liability. And for full transparency, we report all aspects of large language model (LLM) involvement in the Appendix K.
We strictly adhered to the usage and redistribution licenses provided by the original dataset authors and hosting platforms. Our research poses no risk of harm to individuals or groups and does not contain any potentially harmful insights, models, or applications. Additionally, there are no conflicts of interest or sponsorship concerns associated with this work. We are committed to research integrity and ethical standards consistent with the ICLR Code of Ethics.
7 Reproducibility Statement
We actively support the spirit of openness and reproducibility advocated by ICLR. To ensure the reproducibility of our research, we have taken the following measures:
1. Disclosure of Base Models: All base models used in our experiments are explicitly identified and described in the main text. This allows readers to directly reference and obtain these models.
1. Datasets and Experimental Details: All experiments are conducted on publicly available datasets from the Hugging Face platform. In Appendix B, we provide a comprehensive description of our experimental implementation, including dataset sources, browser links, and detailed data processing procedures. We also detail the experimental setup, such as training duration, hardware environment (e.g., GPU type), and configuration of hyperparameters, including LoRA_rank, number of extended layers, batch_size, and max_length. These details facilitate transparent verification and replication of our results.
1. Open-Source Code Release: To further support reproducibility, we release all training and evaluation code in an anonymous repository (https://anonymous.4open.science/status/ADEPT-F2E3). The repository contains clear instructions on installation, data downloading, preprocessing, and experimentation, allowing interested researchers to replicate our results with minimal effort.
References
- Aghajanyan et al. (2021) Armen Aghajanyan, Sonal Gupta, and Luke Zettlemoyer. Intrinsic dimensionality explains the effectiveness of language model fine-tuning. In Chengqing Zong, Fei Xia, Wenjie Li, and Roberto Navigli (eds.), Proceedings of the 59th Annual Meeting of the Association for Computational Linguistics and the 11th International Joint Conference on Natural Language Processing, ACL/IJCNLP 2021, (Volume 1: Long Papers), Virtual Event, August 1-6, 2021, pp. 7319–7328. Association for Computational Linguistics, 2021. doi: 10.18653/V1/2021.ACL-LONG.568. URL https://doi.org/10.18653/v1/2021.acl-long.568.
- Ahn et al. (2024) Janice Ahn, Rishu Verma, Renze Lou, Di Liu, Rui Zhang, and Wenpeng Yin. Large language models for mathematical reasoning: Progresses and challenges. In Proceedings of the 18th Conference of the European Chapter of the Association for Computational Linguistics: Student Research Workshop, pp. 225–237, 2024.
- Arbel et al. (2024) Iftach Arbel, Yehonathan Refael, and Ofir Lindenbaum. Transformllm: Adapting large language models via llm-transformed reading comprehension text. arXiv preprint arXiv:2410.21479, 2024.
- Chen et al. (2024) Junying Chen, Zhenyang Cai, Ke Ji, Xidong Wang, Wanlong Liu, Rongsheng Wang, Jianye Hou, and Benyou Wang. Huatuogpt-o1, towards medical complex reasoning with llms. arXiv preprint arXiv:2412.18925, 2024.
- Chen et al. (2015) Tianqi Chen, Ian Goodfellow, and Jonathon Shlens. Net2net: Accelerating learning via knowledge transfer. arXiv preprint arXiv:1511.05641, 2015.
- Chen et al. (2025) Yang Chen, Zhuolin Yang, Zihan Liu, Chankyu Lee, Peng Xu, Mohammad Shoeybi, Bryan Catanzaro, and Wei Ping. Acereason-nemotron: Advancing math and code reasoning through reinforcement learning. arXiv preprint arXiv:2505.16400, 2025.
- Cheng et al. (2023) Daixuan Cheng, Shaohan Huang, and Furu Wei. Adapting large language models via reading comprehension. In The Twelfth International Conference on Learning Representations, 2023.
- Clark et al. (2019) Kevin Clark, Urvashi Khandelwal, Omer Levy, and Christopher D. Manning. What does BERT look at? an analysis of bert’s attention. In Tal Linzen, Grzegorz Chrupala, Yonatan Belinkov, and Dieuwke Hupkes (eds.), Proceedings of the 2019 ACL Workshop BlackboxNLP: Analyzing and Interpreting Neural Networks for NLP, BlackboxNLP@ACL 2019, Florence, Italy, August 1, 2019, pp. 276–286. Association for Computational Linguistics, 2019. doi: 10.18653/V1/W19-4828. URL https://doi.org/10.18653/v1/W19-4828.
- Clark et al. (2018) Peter Clark, Isaac Cowhey, Oren Etzioni, Tushar Khot, Ashish Sabharwal, Carissa Schoenick, and Oyvind Tafjord. Think you have solved question answering? try arc, the ai2 reasoning challenge. arXiv preprint arXiv:1803.05457, 2018.
- Cobbe et al. (2021) Karl Cobbe, Vineet Kosaraju, Mohammad Bavarian, Mark Chen, Heewoo Jun, Lukasz Kaiser, Matthias Plappert, Jerry Tworek, Jacob Hilton, Reiichiro Nakano, et al. Training verifiers to solve math word problems. arXiv preprint arXiv:2110.14168, 2021.
- Dai et al. (2022) Damai Dai, Li Dong, Yaru Hao, Zhifang Sui, Baobao Chang, and Furu Wei. Knowledge neurons in pretrained transformers. In Proceedings of the 60th Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers), pp. 8493–8502, 2022.
- Ding et al. (2024) Hongxin Ding, Yue Fang, Runchuan Zhu, Xinke Jiang, Jinyang Zhang, Yongxin Xu, Xu Chu, Junfeng Zhao, and Yasha Wang. 3ds: Decomposed difficulty data selection’s case study on llm medical domain adaptation. arXiv preprint arXiv:2410.10901, 2024.
- Ding et al. (2025) Hongxin Ding, Baixiang Huang, Yue Fang, Weibin Liao, Xinke Jiang, Zheng Li, Junfeng Zhao, and Yasha Wang. Promed: Shapley information gain guided reinforcement learning for proactive medical llms. arXiv preprint arXiv:2508.13514, 2025.
- Dubey et al. (2024a) Abhimanyu Dubey, Abhinav Jauhri, Abhinav Pandey, Abhishek Kadian, Ahmad Al-Dahle, Aiesha Letman, Akhil Mathur, Alan Schelten, Amy Yang, Angela Fan, Anirudh Goyal, Anthony Hartshorn, Aobo Yang, Archi Mitra, Archie Sravankumar, Artem Korenev, Arthur Hinsvark, Arun Rao, Aston Zhang, Aurélien Rodriguez, Austen Gregerson, Ava Spataru, Baptiste Rozière, Bethany Biron, Binh Tang, Bobbie Chern, Charlotte Caucheteux, Chaya Nayak, Chloe Bi, Chris Marra, Chris McConnell, Christian Keller, Christophe Touret, Chunyang Wu, Corinne Wong, Cristian Canton Ferrer, Cyrus Nikolaidis, Damien Allonsius, Daniel Song, Danielle Pintz, Danny Livshits, David Esiobu, Dhruv Choudhary, Dhruv Mahajan, Diego Garcia-Olano, Diego Perino, Dieuwke Hupkes, Egor Lakomkin, Ehab AlBadawy, Elina Lobanova, Emily Dinan, Eric Michael Smith, Filip Radenovic, Frank Zhang, Gabriel Synnaeve, Gabrielle Lee, Georgia Lewis Anderson, Graeme Nail, Grégoire Mialon, Guan Pang, Guillem Cucurell, Hailey Nguyen, Hannah Korevaar, Hu Xu, Hugo Touvron, Iliyan Zarov, Imanol Arrieta Ibarra, Isabel M. Kloumann, Ishan Misra, Ivan Evtimov, Jade Copet, Jaewon Lee, Jan Geffert, Jana Vranes, Jason Park, Jay Mahadeokar, Jeet Shah, Jelmer van der Linde, Jennifer Billock, Jenny Hong, Jenya Lee, Jeremy Fu, Jianfeng Chi, Jianyu Huang, Jiawen Liu, Jie Wang, Jiecao Yu, Joanna Bitton, Joe Spisak, Jongsoo Park, Joseph Rocca, Joshua Johnstun, Joshua Saxe, Junteng Jia, Kalyan Vasuden Alwala, Kartikeya Upasani, Kate Plawiak, Ke Li, Kenneth Heafield, Kevin Stone, and et al. The llama 3 herd of models. CoRR, abs/2407.21783, 2024a. doi: 10.48550/ARXIV.2407.21783. URL https://doi.org/10.48550/arXiv.2407.21783.
- Dubey et al. (2024b) Abhimanyu Dubey, Abhinav Jauhri, Abhinav Pandey, Abhishek Kadian, Ahmad Al-Dahle, Aiesha Letman, Akhil Mathur, Alan Schelten, Amy Yang, Angela Fan, et al. The llama 3 herd of models. arXiv e-prints, pp. arXiv–2407, 2024b.
- Dubey et al. (2024c) Abhimanyu Dubey, Abhinav Jauhri, Abhinav Pandey, Abhishek Kadian, Ahmad Al-Dahle, Aiesha Letman, Akhil Mathur, Alan Schelten, Amy Yang, Angela Fan, et al. The llama 3 herd of models. arXiv e-prints, pp. arXiv–2407, 2024c.
- Fang et al. (2025) Yue Fang, Yuxin Guo, Jiaran Gao, Hongxin Ding, Xinke Jiang, Weibin Liao, Yongxin Xu, Yinghao Zhu, Zhibang Yang, Liantao Ma, et al. Toward better ehr reasoning in llms: Reinforcement learning with expert attention guidance. arXiv preprint arXiv:2508.13579, 2025.
- Feng et al. (2024a) Yujie Feng, Xu Chu, Yongxin Xu, Guangyuan Shi, Bo Liu, and Xiao-Ming Wu. Tasl: Continual dialog state tracking via task skill localization and consolidation. arXiv preprint arXiv:2408.09857, 2024a.
- Feng et al. (2024b) Yujie Feng, Xu Chu, Yongxin Xu, Guangyuan Shi, Bo Liu, and Xiao-Ming Wu. Tasl: Continual dialog state tracking via task skill localization and consolidation. In Lun-Wei Ku, Andre Martins, and Vivek Srikumar (eds.), Proceedings of the 62nd Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers), ACL 2024, Bangkok, Thailand, August 11-16, 2024, pp. 1266–1279. Association for Computational Linguistics, 2024b. doi: 10.18653/V1/2024.ACL-LONG.69. URL https://doi.org/10.18653/v1/2024.acl-long.69.
- Hendrycks et al. (2020) Dan Hendrycks, Collin Burns, Steven Basart, Andy Zou, Mantas Mazeika, Dawn Song, and Jacob Steinhardt. Measuring massive multitask language understanding. arXiv preprint arXiv:2009.03300, 2020.
- Hewitt & Manning (2019) John Hewitt and Christopher D. Manning. A structural probe for finding syntax in word representations. In Jill Burstein, Christy Doran, and Thamar Solorio (eds.), Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, NAACL-HLT 2019, Minneapolis, MN, USA, June 2-7, 2019, Volume 1 (Long and Short Papers), pp. 4129–4138. Association for Computational Linguistics, 2019. doi: 10.18653/V1/N19-1419. URL https://doi.org/10.18653/v1/n19-1419.
- Hu et al. (2022) Edward J Hu, Yelong Shen, Phillip Wallis, Zeyuan Allen-Zhu, Yuanzhi Li, Shean Wang, Lu Wang, Weizhu Chen, et al. Lora: Low-rank adaptation of large language models. ICLR, 1(2):3, 2022.
- Huang et al. (2024) Jianheng Huang, Leyang Cui, Ante Wang, Chengyi Yang, Xinting Liao, Linfeng Song, Junfeng Yao, and Jinsong Su. Mitigating catastrophic forgetting in large language models with self-synthesized rehearsal. In Proceedings of the 62nd Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers), pp. 1416–1428, 2024.
- Ibrahim et al. (2024) Adam Ibrahim, Benjamin Thérien, Kshitij Gupta, Mats L Richter, Quentin Anthony, Timothée Lesort, Eugene Belilovsky, and Irina Rish. Simple and scalable strategies to continually pre-train large language models. arXiv preprint arXiv:2403.08763, 2024.
- Jacot et al. (2018) Arthur Jacot, Clément Hongler, and Franck Gabriel. Neural tangent kernel: Convergence and generalization in neural networks. In Samy Bengio, Hanna M. Wallach, Hugo Larochelle, Kristen Grauman, Nicolò Cesa-Bianchi, and Roman Garnett (eds.), Advances in Neural Information Processing Systems 31: Annual Conference on Neural Information Processing Systems 2018, NeurIPS 2018, December 3-8, 2018, Montréal, Canada, pp. 8580–8589, 2018. URL https://proceedings.neurips.cc/paper/2018/hash/5a4be1fa34e62bb8a6ec6b91d2462f5a-Abstract.html.
- Jiang et al. (2024) Xinke Jiang, Yue Fang, Rihong Qiu, Haoyu Zhang, Yongxin Xu, Hao Chen, Wentao Zhang, Ruizhe Zhang, Yuchen Fang, Xu Chu, et al. Tc-rag: turing-complete rag’s case study on medical llm systems. arXiv preprint arXiv:2408.09199, 2024.
- Jiang et al. (2025) Xinke Jiang, Ruizhe Zhang, Yongxin Xu, Rihong Qiu, Yue Fang, Zhiyuan Wang, Jinyi Tang, Hongxin Ding, Xu Chu, Junfeng Zhao, et al. Hykge: A hypothesis knowledge graph enhanced rag framework for accurate and reliable medical llms responses. In Proceedings of the 63rd Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers), pp. 11836–11856, 2025.
- Jin et al. (2021) Di Jin, Eileen Pan, Nassim Oufattole, Wei-Hung Weng, Hanyi Fang, and Peter Szolovits. What disease does this patient have? a large-scale open domain question answering dataset from medical exams. Applied Sciences, 11(14):6421, 2021.
- Ke et al. (2023) Zixuan Ke, Yijia Shao, Haowei Lin, Tatsuya Konishi, Gyuhak Kim, and Bing Liu. Continual pre-training of language models. In The Eleventh International Conference on Learning Representations, ICLR 2023, Kigali, Rwanda, May 1-5, 2023. OpenReview.net, 2023. URL https://openreview.net/forum?id=m_GDIItaI3o.
- Kirkpatrick et al. (2017) James Kirkpatrick, Razvan Pascanu, Neil Rabinowitz, Joel Veness, Guillaume Desjardins, Andrei A Rusu, Kieran Milan, John Quan, Tiago Ramalho, Agnieszka Grabska-Barwinska, et al. Overcoming catastrophic forgetting in neural networks. Proceedings of the national academy of sciences, 114(13):3521–3526, 2017.
- Li et al. (2023) Haonan Li, Yixuan Zhang, Fajri Koto, Yifei Yang, Hai Zhao, Yeyun Gong, Nan Duan, and Timothy Baldwin. Cmmlu: Measuring massive multitask language understanding in chinese. arXiv preprint arXiv:2306.09212, 2023.
- Lin et al. (2024) Bill Yuchen Lin, Abhilasha Ravichander, Ximing Lu, Nouha Dziri, Melanie Sclar, Khyathi Raghavi Chandu, Chandra Bhagavatula, and Yejin Choi. The unlocking spell on base llms: Rethinking alignment via in-context learning. In The Twelfth International Conference on Learning Representations, ICLR 2024, Vienna, Austria, May 7-11, 2024. OpenReview.net, 2024. URL https://openreview.net/forum?id=wxJ0eXwwda.
- Liu et al. (2024a) Chengyuan Liu, Yangyang Kang, Shihang Wang, Lizhi Qing, Fubang Zhao, Chao Wu, Changlong Sun, Kun Kuang, and Fei Wu. More than catastrophic forgetting: Integrating general capabilities for domain-specific LLMs. In Yaser Al-Onaizan, Mohit Bansal, and Yun-Nung Chen (eds.), Proceedings of the 2024 Conference on Empirical Methods in Natural Language Processing, pp. 7531–7548, Miami, Florida, USA, November 2024a. Association for Computational Linguistics. doi: 10.18653/v1/2024.emnlp-main.429. URL https://aclanthology.org/2024.emnlp-main.429/.
- Liu et al. (2024b) Shih-Yang Liu, Chien-Yi Wang, Hongxu Yin, Pavlo Molchanov, Yu-Chiang Frank Wang, Kwang-Ting Cheng, and Min-Hung Chen. Dora: Weight-decomposed low-rank adaptation. In International Conference on Machine Learning, pp. 32100–32121. PMLR, 2024b.
- Luo et al. (2025) Yun Luo, Zhen Yang, Fandong Meng, Yafu Li, Jie Zhou, and Yue Zhang. An empirical study of catastrophic forgetting in large language models during continual fine-tuning. IEEE Transactions on Audio, Speech and Language Processing, 2025.
- OpenAI (2023) OpenAI. Gpt-4 technical report. ArXiv, abs/2303.08774, 2023.
- Paster et al. (2023) Keiran Paster, Marco Dos Santos, Zhangir Azerbayev, and Jimmy Ba. Openwebmath: An open dataset of high-quality mathematical web text. arXiv preprint arXiv:2310.06786, 2023.
- Peng et al. (2025) Ru Peng, Kexin Yang, Yawen Zeng, Junyang Lin, Dayiheng Liu, and Junbo Zhao. Dataman: Data manager for pre-training large language models. In The Thirteenth International Conference on Learning Representations, ICLR 2025, Singapore, April 24-28, 2025. OpenReview.net, 2025. URL https://openreview.net/forum?id=eNbA8Fqir4.
- Qiu et al. (2024) Pengcheng Qiu, Chaoyi Wu, Xiaoman Zhang, Weixiong Lin, Haicheng Wang, Ya Zhang, Yanfeng Wang, and Weidi Xie. Towards building multilingual language model for medicine. Nature Communications, 15(1):8384, 2024.
- Que et al. (2024) Haoran Que, Jiaheng Liu, Ge Zhang, Chenchen Zhang, Xingwei Qu, Yinghao Ma, Feiyu Duan, Zhiqi Bai, Jiakai Wang, Yuanxing Zhang, et al. D-cpt law: Domain-specific continual pre-training scaling law for large language models. Advances in Neural Information Processing Systems, 37:90318–90354, 2024.
- Silverman (2018) Bernard W Silverman. Density estimation for statistics and data analysis. Routledge, 2018.
- Song et al. (2023) Yifan Song, Peiyi Wang, Weimin Xiong, Dawei Zhu, Tianyu Liu, Zhifang Sui, and Sujian Li. Infocl: Alleviating catastrophic forgetting in continual text classification from an information theoretic perspective. In Findings of the Association for Computational Linguistics: EMNLP 2023, pp. 14557–14570, 2023.
- Wang et al. (2023a) Xiao Wang, Tianze Chen, Qiming Ge, Han Xia, Rong Bao, Rui Zheng, Qi Zhang, Tao Gui, and Xuan-Jing Huang. Orthogonal subspace learning for language model continual learning. In Findings of the Association for Computational Linguistics: EMNLP 2023, pp. 10658–10671, 2023a.
- Wang et al. (2023b) Xidong Wang, Guiming Hardy Chen, Dingjie Song, Zhiyi Zhang, Zhihong Chen, Qingying Xiao, Feng Jiang, Jianquan Li, Xiang Wan, Benyou Wang, et al. Cmb: A comprehensive medical benchmark in chinese. arXiv preprint arXiv:2308.08833, 2023b.
- Wu et al. (2024a) C Wu, W Lin, X Zhang, Y Zhang, W Xie, and Y Wang. Pmc-llama: toward building open-source language models for medicine. Journal of the American Medical Informatics Association: JAMIA, pp. ocae045–ocae045, 2024a.
- Wu et al. (2024b) Chengyue Wu, Yukang Gan, Yixiao Ge, Zeyu Lu, Jiahao Wang, Ye Feng, Ying Shan, and Ping Luo. Llama pro: Progressive llama with block expansion. arXiv preprint arXiv:2401.02415, 2024b.
- Xiong et al. (2023) Weimin Xiong, Yifan Song, Peiyi Wang, and Sujian Li. Rationale-enhanced language models are better continual relation learners. In Proceedings of the 2023 Conference on Empirical Methods in Natural Language Processing, pp. 15489–15497, 2023.
- Xu et al. (2025) Yongxin Xu, Ruizhe Zhang, Xinke Jiang, Yujie Feng, Yuzhen Xiao, Xinyu Ma, Runchuan Zhu, Xu Chu, Junfeng Zhao, and Yasha Wang. Parenting: Optimizing knowledge selection of retrieval-augmented language models with parameter decoupling and tailored tuning. In Wanxiang Che, Joyce Nabende, Ekaterina Shutova, and Mohammad Taher Pilehvar (eds.), Proceedings of the 63rd Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers), pp. 11643–11662, Vienna, Austria, July 2025. Association for Computational Linguistics. ISBN 979-8-89176-251-0. doi: 10.18653/v1/2025.acl-long.571. URL https://aclanthology.org/2025.acl-long.571/.
- Yang et al. (2025a) An Yang, Anfeng Li, Baosong Yang, Beichen Zhang, Binyuan Hui, Bo Zheng, Bowen Yu, Chang Gao, Chengen Huang, Chenxu Lv, et al. Qwen3 technical report. arXiv preprint arXiv:2505.09388, 2025a.
- Yang et al. (2025b) Zhibang Yang, Xinke Jiang, Rihong Qiu, Ruiqing Li, Yihang Zhang, Yue Fang, Yongxin Xu, Hongxin Ding, Xu Chu, Junfeng Zhao, et al. Dfams: Dynamic-flow guided federated alignment based multi-prototype search. arXiv preprint arXiv:2508.20353, 2025b.
- Yang et al. (2024) Zitong Yang, Neil Band, Shuangping Li, Emmanuel Candes, and Tatsunori Hashimoto. Synthetic continued pretraining. arXiv preprint arXiv:2409.07431, 2024.
- Yıldız et al. (2024) Çağatay Yıldız, Nishaanth Kanna Ravichandran, Nitin Sharma, Matthias Bethge, and Beyza Ermis. Investigating continual pretraining in large language models: Insights and implications. arXiv preprint arXiv:2402.17400, 2024.
- Zeng (2023) Hui Zeng. Measuring massive multitask chinese understanding. arXiv preprint arXiv:2304.12986, 2023.
- Zhang et al. (2023) Hongbo Zhang, Junying Chen, Feng Jiang, Fei Yu, Zhihong Chen, Jianquan Li, Guiming Chen, Xiangbo Wu, Zhiyi Zhang, Qingying Xiao, et al. Huatuogpt, towards taming language model to be a doctor. arXiv preprint arXiv:2305.15075, 2023.
- Zhang et al. (2025) Yunan Zhang, Shuoran Jiang, Mengchen Zhao, Yuefeng Li, Yang Fan, Xiangping Wu, and Qingcai Chen. Gere: Towards efficient anti-forgetting in continual learning of llm via general samples replay. arXiv preprint arXiv:2508.04676, 2025.
- Zheng et al. (2024) Zifan Zheng, Yezhaohui Wang, Yuxin Huang, Shichao Song, Mingchuan Yang, Bo Tang, Feiyu Xiong, and Zhiyu Li. Attention heads of large language models: A survey. arXiv preprint arXiv:2409.03752, 2024.
Appendix A Related Work
A.1 Continual Pretraining for LLMs
Continual pretraining updates pretrained LLMs with new corpora to equip them with new knowledge and capabilities. Data-centric approaches adopt data replay to mitigate catastrophic forgetting (Huang et al., 2024; Zhang et al., 2025; Xiong et al., 2023; Song et al., 2023), or utilize data construction strategies to synthesize training corpora (Yang et al., 2024; Arbel et al., 2024). However, these methods make no changes to the model or training procedure, failing to effective inject new knowledge due to capacity saturation and only partially alleviating forgetting. Another line of works focus on adjusting model architecture and training strategy. LoRA (Hu et al., 2022) improve efficiency for fine-tuning by adapting low-rank updates on top of frozen backbones, but their limited adjustments to LLMs can not effectively address continual pretraining for deep domain adaptation. LLaMA-Pro (Wu et al., 2024b) expands model blocks and tunes the added parameters on new corpora, improving knowledge injection and mitigating forgetting compared to vanilla CPT. Yet existing expansion policies insert layers uniformly across depths and treat all expanded parameters indiscriminately during optimization, leaving open how to place capacity where domain signals concentrate and update it without disturbing general knowledge. Classical continual-learning regularizers (Kirkpatrick et al., 2017) constrain updates on weights deemed important to previous tasks, but they do not guide where capacity allocation nor how to target LLM domain adaptation learning.
A.2 Functional Specialization in LLMs
Growing evidence indicates that, akin to human brains, LLMs exhibit functional specialization, where different regions such as layers, attention heads and neurons play distinct roles. A series of causal and studies show that factual knowledge are predominantly stored in FFN layers (Dai et al., 2022), and attention heads usually play specialized roles for certain functions (Zheng et al., 2024), suggesting that knowledge and skills are unevenly distributed in LLMs. Inspired by this specialization, several methods have tried to decouple functional modules during training. For instance, Parenting (Xu et al., 2025) separates the subspaces responsible for evidence-following and noise-robustness in retrieval-augmented generation, and optimizes them with tailored objectives to improve performance under noisy retrieval. Similarly, TaSL (Feng et al., 2024a) addresses multi-task adaptation by disentangling LoRA parameters from different tasks and merging them in a weighted manner, which helps reduce interference. Other works on orthogonal (Wang et al., 2023a) or decomposed LoRA (Liu et al., 2024b) further reflects the idea that training different parameter subspaces separately improves robustness and transfer. Despite these advances, prior work does not address CPT, where the tension between knowledge injection and retention needs to be tackled. To our knowledge, our work is the first to explicitly leverage functional specialization during CPT to simultaneously improve domain performance and alleviate catastrophic forgetting.
A.3 Domain Adaptation via Other Paradigms.
Beyond continual pretraining, prior studies have explored other paradigms for adapting large language models to specific domains. Retrieval-augmented generation approaches, such as HyKGE (Jiang et al., 2025), TC-RAG (Jiang et al., 2024), and DFAMS (Yang et al., 2025b), enhance domain adaptation through external retrieval and knowledge grounding. Supervised fine-tuning methods, including 3DS (Ding et al., 2024) and HuatuoGPT (Zhang et al., 2023), align general models to domain distributions via instruction tuning on curated data. Reinforcement learning approaches, such as ProMed (Ding et al., 2025) and EAG-RL (Fang et al., 2025), further refine reasoning and decision-making through reward optimization. While these directions provide complementary insights into domain adaptation, they are beyond the scope of this work, which focuses on continual pretraining as a self-contained and scalable solution.
Appendix B Data Recipe and Experiment Settings
To demonstrate the applicability and generalizability of our approach, we conducted domain-adaptive continual pretraining experiments on two distinct and highly significant domains: Mathematics and Medicine, both of which play crucial roles in the advancement of artificial intelligence and the applications of LLM. The mathematical domain often poses challenges that emphasize a model’s reasoning and computational abilities, while the medical domain predominantly requires a deep understanding and memorization of medical concepts. From a cognitive perspective, we believe that the capabilities that need to be infused into the model differ significantly between these two domains, which further demonstrates the generalisability of our approach.
The continual pretraining process leverages both pretraining datasets for foundational knowledge and supervised fine-tuning (SFT) datasets for task-specific optimization (Cheng et al., 2023). Below, we detail the data composition and processing details. All data used will be processed into the format of pre-training data.
B.1 Medical Pretrain Data Source
Our medicine datasets are divided into pre-training data, designed to provide extensive general knowledge, and supervised fine-tuning (SFT) data, which refine the model’s understanding for specific instructions in the medicine domain (will be converted to pretrain data format when training).
- Pre-training data: we utilize English and Chinese portions of MMedC dataset, a multilingual medical dataset, furnishing a total of 14.3 billion tokens.
- Instrution tuning data: we incorporate two supervised datasets:
1. IndustryIns, contributing 1.6 billion tokens from instruction-based examples
2. MMedBench, with 18 million tokens focused on medical reasoning tasks.
Table 3: Overview of medicine Datasets. This table summarizes medicine-specific pre-training and SFT datasets, including their language coverage, dataset links, and used token counts. For MMedC, we only use the English and Chinese parts and we only use the Health-Medicine subset.
| Dataset Name | Dataset Type | Language | Dataset Link | #Token Used |
| --- | --- | --- | --- | --- |
| MMedC | Pre-training | Multilingual | Henrychur/MMedC | 14.3B |
| IndustryIns | SFT | Chinese and English | BAAI/IndustryInstruction | 1.6B |
| MMedBench | SFT | Chinese and English | Henrychur/MMedBench | 18M |
B.2 Mathematics Pretrain Data Source
Mathematics pretrain datasets include both pre-training and fine-tuning data (will be converted to pretrain data format when training), structured similarly to the medicine datasets.
- Pre-training data: we use the Open-Web-Math (Paster et al., 2023) dataset, containing a diverse set of general mathematics knowledge amounting to 14.7 billion tokens.
- For Instruction-tuning data: we use the AceReason-Math (Chen et al., 2025), contributing 102 million tokens, with a strong emphasis on chain-of-thought reasoning and problem-solving.
Table 4: Overview of Mathematics Datasets. This table includes the pre-training and SFT datasets for mathematical reasoning, highlighting their contents, links, and used token counts.
| Dataset Name | Dataset Type | Language | Dataset Link | #Used Token |
| --- | --- | --- | --- | --- |
| Open-Web-Math | Pre-training | English | open-web-math/open-web-math | 14.7B |
| AceReason-Math | SFT | English | nvidia/AceReason-Math | 102M |
B.3 General Competence Detection Corpus
To accurately probe which parameters are critical for preserving general knowledge during continual pretraining, we construct a General Importance Detection Corpus. This corpus is designed to capture both broad world knowledge and instruction-following capability in English and Chinese. Specifically, we include the development splits of two widely recognized multi-task benchmarks, MMLU_dev and CMMLU_dev to capture general knowledge without data leakage.
MMLU and CMMLU are formatted as multiple-choice question answering tasks with explicit prompts and ground-truth answers. For these, we compute gradient-based importance only on the target answer tokens to avoid biases from prompt formatting, thereby capturing each parameter group’s contribution to accuracy.
To clarify how gradient signals are obtained, we illustrate two examples. In SFT-style corpora (e.g., MMLU, CMMLU), only the ground-truth answer token contributes to gradient computation, ensuring clean signals for decision-making importance. In PT-style corpora (e.g., FineWeb_Edu), all tokens contribute under the causal LM objective, providing dense gradients that reflect general modeling capacity. Examples are shown in Example 1 and Example 2.
Table 5: General Competence Detection Corpus. #Examples means the number of examples we used.
| Dataset | Language | #Examples | Hugging Face Link |
| --- | --- | --- | --- |
| MMLU_dev | English | 285 | cais/mmlu |
| CMMLU_dev | Chinese | 295 | haonan-li/cmmlu |
The statistics of the selected datasets are summarized in Table 5.
Example 1
Gradient Flow in SFT Data for Importance Estimation Input Prompt: Question: Find all $c∈\mathbb{Z}_{3}$ such that $\mathbb{Z}_{3}[x]/(x^{2}+c)$ is a field. A. 0 B. 1 C. 2 D. 3 Answer: B Explanation: In this SFT setup, only the target answer token (e.g., B) is used to compute gradients for parameter importance. The input question and options are excluded from gradient computation to avoid encoding biases from instruction formatting. By focusing gradient signals solely on the correct answer token, we measure how each parameter contributes to decision-making accuracy under structured knowledge tasks, while preventing overfitting to input patterns and ensuring clean separation between training and probing data.
Example 2
Gradient Flow in PT Data for Importance Estimation Context (Compute Gradient): The heart is a muscular organ responsible for pumping blood throughout the body. It consists of four chambers: the left and right atria, and the left and right ventricles. Oxygen-poor blood enters the right atrium, then flows to the right ventricle, which pumps it to the lungs. After oxygenation, blood returns to the left atrium, moves to the left ventricle, and is finally pumped into the aorta for systemic circulation. This process is regulated by electrical signals originating in the sinoatrial node. These signals ensure synchronized contraction and efficient blood flow. Explanation: In PT-style training, parameter importance is computed using causal language modeling loss across the entire sequence. Every token — both context and continuation — contributes to the gradient signal. This captures how parameters support general language modeling over natural text distributions. Unlike SFT, there is no explicit input/output separation; instead, each token is predicted from its prefix, making the gradient flow dense and continuous. This allows us to assess parameter sensitivity in open-ended, domain-relevant pre-training scenarios such as those provided by FineWeb_Edu.
B.4 Data Processing
To generate training corpus in pretrain format, SFT data is structured by concatenating questions, chain-of-thought (CoT) reasoning, and final answers for each instance. This ensures that the model is optimized for multi-step reasoning tasks common in medicine applications. We take Example 3 as an example.
Example 3
Problem: On Liar Island, half the people lie only on Wednesday, Friday, and Saturday, while the other half lie only on Tuesday, Thursday, and Sunday. One day, everyone on the island says: “I will tell the truth tomorrow.” What day is it? (2021 Xin Xiwang Bei Competition, Grade 2, Preliminary Math Exam) Analysis: We examine the truth-telling patterns over the week: •
First group (lies on Wed, Fri, Sat): Truth pattern across 7 days: True, True, False, True, False, False, True. •
Second group (lies on Tue, Thu, Sun): Truth pattern: True, False, True, False, True, True, False. Now evaluate each option: Option A (Tuesday): If today is Tuesday, the first group tells the truth today, so their statement “I will tell the truth tomorrow” implies they should tell the truth on Wednesday. But they lie on Wednesday — contradiction. The second group lies today, so their statement is false, meaning they will not tell the truth tomorrow (i.e., lie on Wednesday). But they actually tell the truth on Wednesday — also a contradiction. So A is invalid. Option B (Wednesday): First group lies today; their statement is false → they will not tell the truth tomorrow (i.e., lie on Thursday). But they tell the truth on Thursday — contradiction. Second group tells the truth today → they should tell the truth on Thursday. But they lie on Thursday — contradiction. So B is invalid. Option C (Friday): First group lies today → statement is false → they will not tell the truth tomorrow (i.e., lie on Saturday). They do lie on Saturday — consistent. Second group tells the truth today → they will tell the truth on Saturday. They do tell the truth on Saturday — consistent. So C is correct. Option D (Saturday): First group lies today → should lie on Sunday. But they tell the truth on Sunday — contradiction. Second group tells the truth today → should tell the truth on Sunday. But they lie on Sunday — contradiction. So D is invalid. Option E (Sunday): First group tells the truth today → should tell the truth on Monday. They do — consistent. Second group lies today → their statement is false → they will not tell the truth on Monday (i.e., lie). But they tell the truth on Monday — contradiction. So E is invalid. Therefore, the correct answer is C (Friday).
[This example demonstrates how structured SFT data — consisting of a standalone problem (in blue), detailed step-by-step analysis (in green) and a short answer (in red) — is concatenated into a single coherent narrative. In PT-style training, such concatenation enables models to learn implicit reasoning patterns from natural language flow, bridging supervised fine-tuning signals with pre-training objectives.]
To handle input sequences that exceed the maximum context length of 4096 tokens imposed by transformer-based models, we apply a sliding window segmentation strategy with overlap, following the approach used in DataMan (Peng et al., 2025). For any sequence longer than 4096 tokens, we split it into multiple segments, each of length at most 4096, using a sliding window with a stride of 3072 tokens and an overlap of 1024 tokens (i.e., 1/4 of the window size). This ensures that consecutive segments share contextual information when training in the same or adjacent batches, preserving semantic continuity and high data utilization rate across boundaries.
Formally, given a token sequence $D=[t_{1},t_{2},...,t_{L}]$ of length $L>4096$ , we generate $K=\left\lceil\frac{L-1024}{3072}\right\rceil$ segments. The $k$ -th segment is defined as $S_{k}=D[\ell_{k}:r_{k}]$ , where $\ell_{k}=(k-1)· 3072+1$ and $r_{k}=\min(\ell_{k}+4097,L)$ . The overlapping region between $S_{k}$ and $S_{k+1}$ consists of the last 1024 tokens of $S_{k}$ , which are identical to the first 1024 tokens of $S_{k+1}$ .
This method prevents information loss due to truncation and allows the model to learn from continuous context during training. The 1024-token overlap helps maintain coherence at segment boundaries, which is crucial for tasks requiring long-range understanding, while keeping computational overhead manageable.
B.5 Final Data Organization Scheme
Our final training data is organized as follows:
1. English pre-training corpus
1. Chinese pre-training corpus (if have)
1. English supervised fine-tuning (SFT) corpus
1. Chinese SFT corpus (if have)
This organization is motivated by several key points in Qwen3 Technical Report (Yang et al., 2025a) and Llama3 Technical Report (Dubey et al., 2024a). First, we follow the principle that high-quality data (SFT data in our work) should be used after extensive pre-training on large-scale general corpora, allowing the model to first acquire broad knowledge and language structure, and then specialize on more curated tasks and instructions.
What’s more, according to the technical reports, it is further beneficial to place the same language’s data together during training—this maximizes the coherence within each mini-batch and reduces unintended cross-lingual transfer until later stages. Most LLMs are dominated by English corpora in their pre-training phase, supporting the choice of placing English data first. Finally, during later training stages, continued training and decay are performed on SFT examples, which aligns with established recipes for improving supervised task performance.
B.6 Compared Methods.
- Full-parameter tuning. PT-Full directly updates all model parameters on the target corpus, serving as the most straightforward yet commonly used baseline for continual pretraining.
- Replay-based tuning. Replay mitigates catastrophic forgetting by mixing general-domain data into the continual pretraining process (Que et al., 2024), thereby preserving part of the original knowledge distribution while adapting to the new domain. Following (Zhang et al., 2025), based on the data from Data Recipe, we randomly sampled totally 1.91B data from FinewebEdu and FinewebEdu-Chinese at a ratio of 7:3, and randomly shuffled them into the domain-specific data, helping the model better recall general domain knowledge.
- Architecture expansion. LLaMA-Pro (Wu et al., 2024b) expands the model by uniformly inserting new layers into each transformer block while freezing the original weights. Only the newly introduced parameters are trained, enabling structural growth while preserving prior knowledge.
- Parameter-efficient tuning. PT-LoRA performs continual pretraining using Low-Rank Adaptation (Hu et al., 2022), updating only a small set of task-adaptive parameters. TaSL (Feng et al., 2024a) extends PT-LoRA to a multi-task regime by decoupling LoRA matrices across transformer layers, allowing different subsets of parameters to specialize for different tasks. This enables more fine-grained adaptation to domain-specific signals. We used the DEV sets of MMLU and CMMLU to assess general capabilities, and their mathematics and medical subsets to specifically evaluate mathematical and medical competencies, respectively. Taking the medical domain as an example, we treat the original model as one equipped with a LoRA module initialized to all zeros. The final LoRA module is then obtained by merging the domain-specific LoRA with the original (empty) LoRA using TaSL.
B.7 Experimental Implementation.
We conduct our all pre-training experiments on the Qwen3-1.7B-Base/Qwen3-4B-Base/Qwen3-8B-Base/Llama3-8B model with the following hyperparameter configuration. Training is performed for 3 epochs using a batch size of 512 (8 NVIDIA H800 GPUs) and a maximum sequence length of 4096 tokens. We utilize a cosine learning rate scheduler with an initial learning rate of 3.0e-5 and a warmup ratio of 0.03. Optimization is performed in bf16 precision.
For methods requiring block expansion, we expand 4 layers; for methods based on LoRA, we set the LoRA rank to 256 to ensure the number of trainable parameters is roughly comparable between the two approaches. For the medicine injection into Llama models, which have poor Chinese support, we expand 8 layers for block expansion methods and set the LoRA rank to 512 for LoRA-based methods.
For our ADEPT, we calculate the importance score and update learning rate per 500 iterations. (It does not affect the impact of warmup, decay scheduler on the learning rate, but only performs a reallocation.)
B.8 Efficiency Analysis of ADEPT for Medical Applications
Table 6: Training Time Comparison in the Medical Domain. We select representative baselines including full-parameter (PT-Full) training, PT-Lora, and Llama Pro to validate the effectiveness of our method. The bold entries denote the optimal results.
| | Qwen3-1.7B | Qwen3-4B | Qwen3-8B | Llama3-8B |
| --- | --- | --- | --- | --- |
| PT-Full | 2 days, 17h | 5 days, 14h | 8 days, 9h | 7 days, 22h |
| ADEPT | 1 day, 9h | 2 days, 11h | 3 days, 15h | 3 days, 19h |
| PT-Lora | 3 days, 0h | 6 days, 4h | 8 days, 23h | 8 days, 2h |
| Llama Pro | 2 days, 1h | 3 days, 14h | 5 days, 8h | 4 days, 21h |
As shown in the Table 6, our ADEPT approach achieves the fastest training time across all tested model sizes, with Llama Pro being the next most efficient competitor. The substantial efficiency gain of our method is mainly attributed to its design: ADEPT only updates a small subset of parameters, primarily located in the deeper layers of the network. This structure allows the backward computation graph to terminate earlier, significantly reducing the overall training time.
B.9 Evaluation Setting
We evaluate the performance of large language models on multiple-choice question answering tasks using accuracy as the primary metric. For a given question with $N$ candidate options (typically $N=4$ , labeled A, B, C, D), the model’s prediction is determined by computing the sequence-level likelihood of each option when appended to the question stem.
Specifically, let $Q$ denote the input question and $O_{i}$ represent the $i$ -th answer option (e.g., A. True, B. False). The model computes the conditional probability of the full sequence $Q\parallel O_{i}$ (i.e., the concatenation of the question and the $i$ -th option) under the causal language modeling objective. We calculate the average negative log-likelihood (or perplexity, PPL) of the tokens in $O_{i}$ given $Q$ :
$$
\text{PPL}(O_{i}\mid Q)=\exp\left(-\frac{1}{|O_{i}|}\sum_{t=1}^{|O_{i}|}\log P(o_{t}\mid Q,o_{1},\dots,o_{t-1})\right) \tag{8}
$$
The model selects the option with the lowest perplexity as its predicted answer:
$$
\hat{y}=\arg\min_{O_{i}\in\{\text{A},\text{B},\text{C},\text{D}\}}\text{PPL}(O_{i}\mid Q) \tag{9}
$$
This method, often referred to as perplexity-based decoding, does not require fine-tuning or additional parameters and is widely used for evaluation of base models. It leverages the pre-training objective directly by predicting the next token, making it particularly suitable for evaluating general knowledge in base LLMs.
Finally, accuracy is defined as the percentage of questions for which the model’s predicted answer matches the ground-truth label:
$$
\text{Accuracy}=\frac{1}{M}\sum_{j=1}^{M}\mathbb{I}(\hat{y}_{j}=y_{j}) \tag{10}
$$
where $M$ is the total number of test questions, $\hat{y}_{j}$ is the model’s prediction on the $j$ -th question, $y_{j}$ is the true label, and $\mathbb{I}(·)$ is the indicator function.
For our experiments, we evaluate all model checkpoints using the lm_harness https://github.com/EleutherAI/lm-evaluation-harness framework. For the Mathematics domain, we adopt the default configurations of GSM8K_cot, ARC-Easy, and ARC-Challenge. For the Medical domain, we design custom configuration files for MedQA, MMCU-Medical, and CMB, following the official evaluation protocols of MMLU and CMMLU. For the General domain, we directly evaluate on MMLU and CMMLU. In all cases, we use 5-shot prompts and greedy decoding (temperature = 0) for inference. This standardized evaluation protocol ensures fair comparison across models and tasks.
Appendix C Model Parameter Group
To enable efficient and semantically meaningful parameter decoupling during fine-tuning, we partition the model parameters into modular units based on their functional roles within the transformer architecture. Given the substantial number of model parameters, extremely fine-grained control at the neuron level—as used in methods like DAS (Ke et al., 2023) —is computationally prohibitive and contradicts the goal of parameter-efficient adaptation. Moreover, such fine granularity often leads to training instability due to noisy importance estimation.
On the other hand, treating an entire layer as a single unit (e.g., standard LoRA) is too coarse and lacks semantic discrimination. While TaSL (Feng et al., 2024b) proposes decomposing LoRA into LoRA_A and LoRA_B, this approach is specific to low-rank adapters and does not generalize well to full-layer decomposition.
To strike a balance between granularity and efficiency, we introduce a semantic-aware module partitioning strategy, which divides each transformer layer into multiple functional units according to their architectural semantics. This design allows us to manipulate parameters at a meaningful intermediate level—finer than whole layers, but coarser than individual neurons—achieving a practical trade-off between controllability and computational feasibility.
Table 7 presents the detailed parameter grouping scheme used in this work, exemplified on the LLaMA architecture.
Table 7: Model Parameter Grouping Scheme
| Parameter Type | Parameter Name | Description |
| --- | --- | --- |
| Attention | self_attn.q_proj.weight | Query projection weight; maps input to query space |
| self_attn.k_proj.weight | Key projection weight; maps input to key space | |
| self_attn.v_proj.weight | Value projection weight; maps input to value space | |
| self_attn.o_proj.weight | Output projection weight; projects attention output back to target dimension | |
| MLP | mlp.gate_proj.weight | Gating projection weight; controls information flow in SwiGLU activation |
| mlp.up_proj.weight | Up-projection weight; maps features to higher-dimensional intermediate space | |
| mlp.down_proj.weight | Down-projection weight; projects features back to original dimension | |
| LayerNorm | input_layernorm.weight | Input layer normalization weight; normalizes input before attention |
| post_attention_layernorm.weight | Normalization weight after attention; stabilizes post-attention outputs | |
As shown in Table 7, each transformer layer is decomposed into three primary functional modules: Attention, MLP, and LayerNorm. Within each module, parameters are grouped by their semantic role:
- The Attention module includes all four linear projections ( $Q$ , $K$ , $V$ , $O$ ), which collectively handle context modeling through self-attention.
- The MLP module contains the up, gate, and down projection layers, responsible for non-linear feature transformation.
- The LayerNorm components are kept separate due to their distinct role in stabilizing activations and gradient flow.
This grouping enables targeted manipulation of specific sub-functions (e.g., disabling attention outputs or freezing normalization statistics) while maintaining training stability and interpretability.
C.1 Compatibility between Layer Expansion and Decoupling
First, we would like to share our understanding of the Compatibility between Layer Expansion and Decoupling:
1. Although layer expansion can minimize changes to the original parameter space, this alone makes it difficult to fully prevent model drift during long-term pre-training. Parameter decoupling offers a more fine-grained means of controlling this phenomenon.
2. Since our models are pre-trained on a large corpus, their parameter space is inherently uncontrollable, making thorough decoupling of the original model parameters challenging. In contrast, the newly expanded parameters initially contribute nothing to the model’s output. As we continue domain-specific training in the medical field, gradually decoupling these new parameters is more conducive to achieving complete decoupling.
<details>
<summary>figures/1.7B_Base_27_layers.jpg Details</summary>
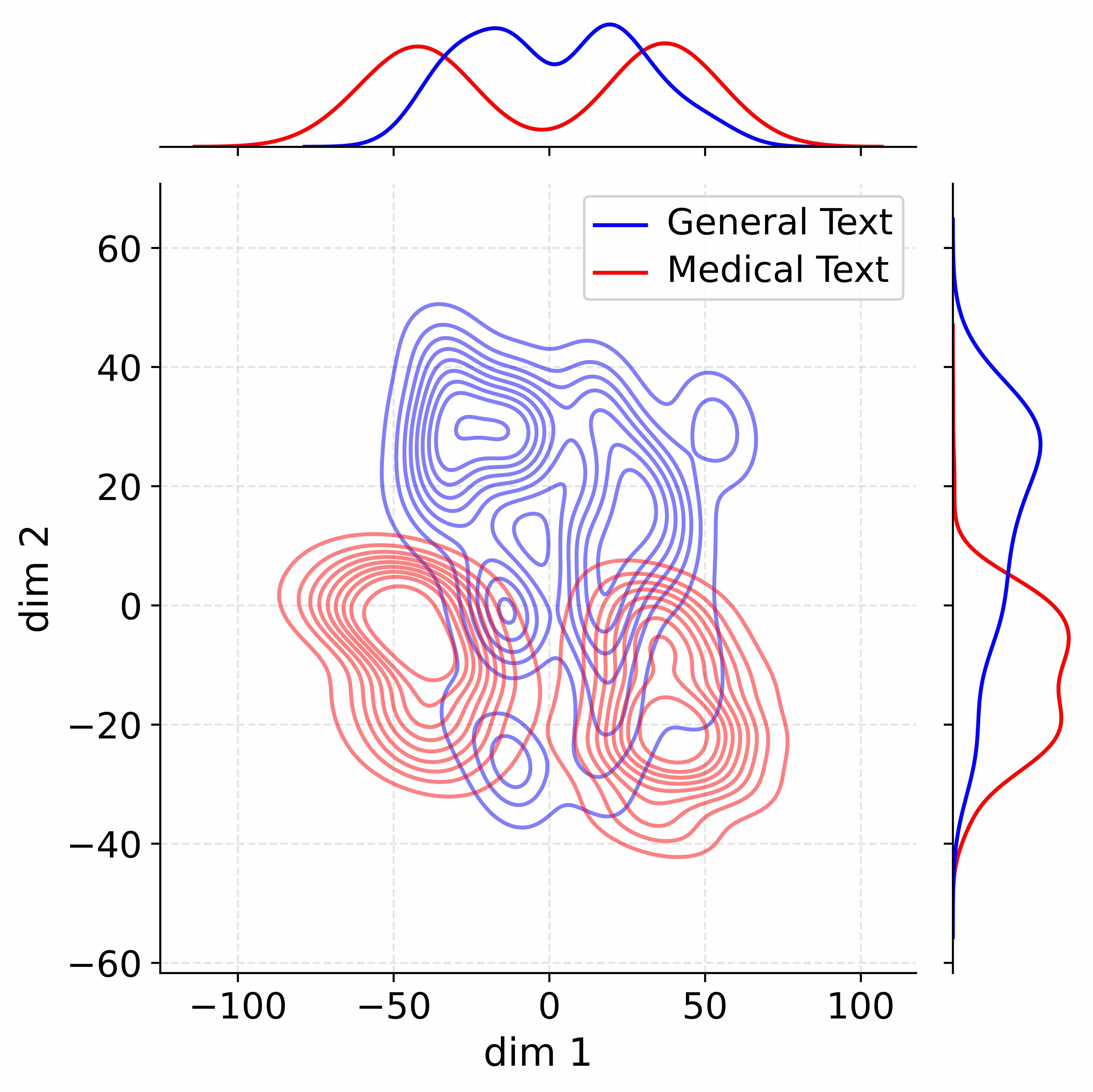
### Visual Description
## Density Contour Plot: General Text vs. Medical Text
### Overview
The image is a density contour plot comparing "General Text" and "Medical Text" across two dimensions (dim 1 and dim 2). It includes marginal density plots along the top (dim 1) and right side (dim 2). The plot visualizes the distribution and concentration of data points for each text type.
### Components/Axes
* **Main Plot:** Density contour plot showing the distribution of "General Text" (blue) and "Medical Text" (red) in a two-dimensional space.
* **X-axis (dim 1):** Ranges from -100 to 100, with tick marks at -100, -50, 0, 50, and 100.
* **Y-axis (dim 2):** Ranges from -60 to 60, with tick marks at -60, -40, -20, 0, 20, 40, and 60.
* **Top Marginal Plot:** Density plot showing the distribution of "General Text" (blue) and "Medical Text" (red) along the x-axis (dim 1).
* **Right Marginal Plot:** Density plot showing the distribution of "General Text" (blue) and "Medical Text" (red) along the y-axis (dim 2).
* **Legend:** Located in the top-center of the plot.
* Blue line: "General Text"
* Red line: "Medical Text"
### Detailed Analysis
**1. Main Density Contour Plot:**
* **General Text (Blue):**
* The highest density area is centered around x = -40 and y = 30.
* The contours extend from approximately x = -80 to x = 40 and y = 0 to y = 50.
* A secondary, less dense area is located around x = 30 and y = 10.
* **Medical Text (Red):**
* The highest density area is centered around x = -40 and y = -10.
* The contours extend from approximately x = -80 to x = 40 and y = -40 to y = 10.
* A secondary, less dense area is located around x = 30 and y = -20.
**2. Top Marginal Density Plot (dim 1):**
* **General Text (Blue):**
* Shows two peaks, one around x = -40 and another around x = 30.
* **Medical Text (Red):**
* Shows two peaks, one around x = -40 and another around x = 30. The peak at x=-40 is smaller than the peak at x=30.
**3. Right Marginal Density Plot (dim 2):**
* **General Text (Blue):**
* Shows a single peak around y = 30.
* **Medical Text (Red):**
* Shows a single peak around y = -10.
### Key Observations
* The "General Text" distribution is shifted towards higher values of dim 2 compared to "Medical Text".
* Both "General Text" and "Medical Text" show a bimodal distribution along dim 1, with peaks around -40 and 30.
* The marginal distributions confirm the shift in dim 2 and the bimodal nature of dim 1.
### Interpretation
The density contour plot suggests that "General Text" and "Medical Text" have distinct distributions in the two-dimensional space defined by dim 1 and dim 2. "General Text" tends to have higher values along dim 2 compared to "Medical Text". The bimodal distribution along dim 1 for both text types indicates the presence of two distinct clusters or sub-categories within each text type. The plot could represent a feature space derived from text embeddings, where dim 1 and dim 2 are principal components or other relevant dimensions. The separation between the distributions suggests that these dimensions are useful for distinguishing between "General Text" and "Medical Text".
</details>
(a) Vanilla
<details>
<summary>figures/1.7B_only_grad_27_layers.jpg Details</summary>
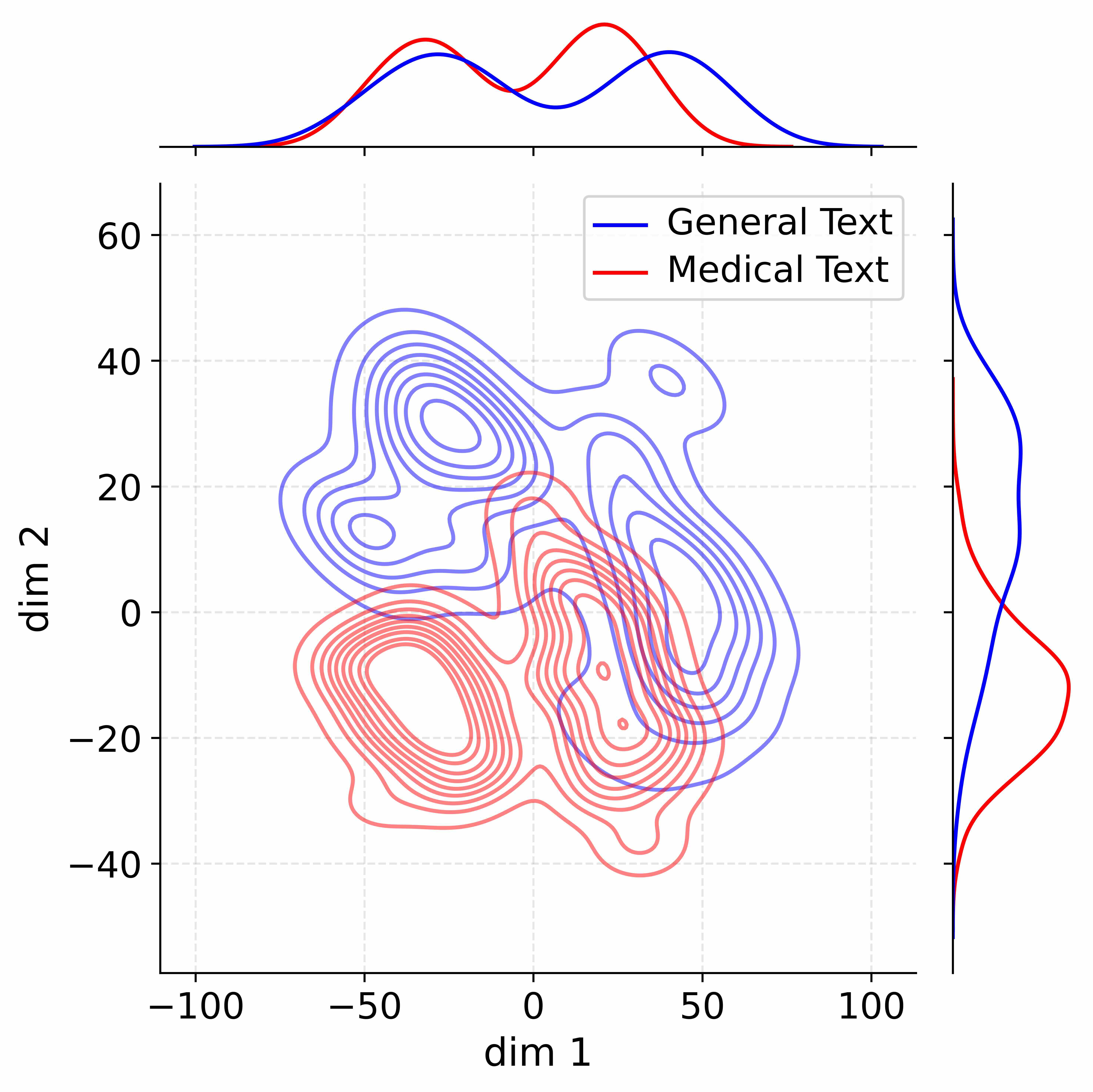
### Visual Description
## Contour Plot: Density Distribution of Text Types
### Overview
The image is a contour plot showing the density distribution of two types of text, "General Text" and "Medical Text", across two dimensions, "dim 1" and "dim 2". Marginal density plots are shown along the top and right axes.
### Components/Axes
* **Main Plot:** A 2D contour plot with "dim 1" on the x-axis and "dim 2" on the y-axis.
* X-axis ("dim 1"): Ranges from -100 to 100, with tick marks at -100, -50, 0, 50, and 100.
* Y-axis ("dim 2"): Ranges from -40 to 60, with tick marks at -40, -20, 0, 20, 40, and 60.
* Gridlines: Light gray dashed lines.
* **Top Marginal Plot:** A 1D density plot showing the distribution of "dim 1" for both text types.
* **Right Marginal Plot:** A 1D density plot showing the distribution of "dim 2" for both text types.
* **Legend:** Located in the top-center of the plot.
* "General Text": Represented by a blue line.
* "Medical Text": Represented by a red line.
### Detailed Analysis
* **General Text (Blue):**
* In the main plot, the blue contours indicate a higher density region centered approximately around dim1 = -30 and dim2 = 30. There's a secondary, less dense region around dim1 = 40 and dim2 = 20.
* Top marginal plot: The blue line shows two peaks, one around -30 and another around 40.
* Right marginal plot: The blue line shows a single peak around dim2 = 30.
* **Medical Text (Red):**
* In the main plot, the red contours indicate a higher density region centered approximately around dim1 = -20 and dim2 = -10.
* Top marginal plot: The red line shows a single peak around -20.
* Right marginal plot: The red line shows a single peak around dim2 = -10.
### Key Observations
* The "General Text" distribution is bimodal along "dim 1", suggesting two distinct clusters or sub-categories within the general text data.
* The "Medical Text" distribution is unimodal in both dimensions, indicating a more concentrated cluster.
* There is some overlap between the two distributions, particularly in the region around dim1 = -20 to 20 and dim2 = 0 to 20.
### Interpretation
The plot visualizes the distribution of "General Text" and "Medical Text" in a two-dimensional space defined by "dim 1" and "dim 2". The separation between the blue and red contours suggests that these two types of text occupy different regions in this space, implying that "dim 1" and "dim 2" are able to differentiate between them. The bimodal distribution of "General Text" along "dim 1" could indicate that this category is composed of two distinct sub-categories, while "Medical Text" appears to be more homogeneous. The overlap between the distributions suggests that there are some instances where the two types of text are similar or indistinguishable based on these two dimensions.
</details>
(b) W/o stage1
<details>
<summary>figures/1.7B_final_Layer31_true.jpg Details</summary>
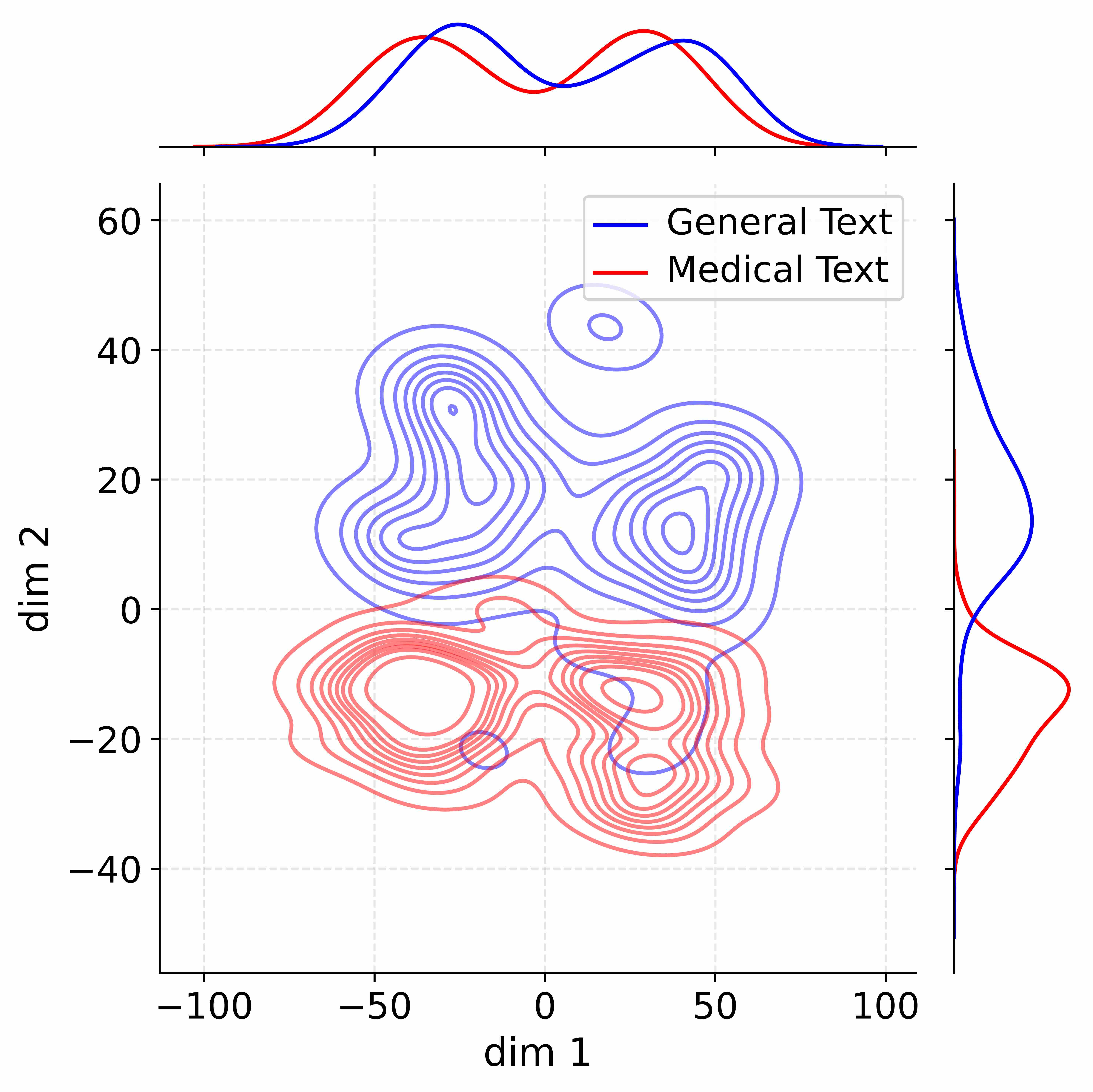
### Visual Description
## Density Plot: General Text vs. Medical Text
### Overview
The image is a density plot comparing "General Text" and "Medical Text" across two dimensions, labeled "dim 1" and "dim 2". The plot shows the distribution of data points for each type of text, with contour lines indicating areas of higher density. Marginal density plots are shown along the top and right edges of the main plot.
### Components/Axes
* **Main Plot:**
* X-axis: "dim 1", ranging from -100 to 100, with tick marks at -100, -50, 0, 50, and 100.
* Y-axis: "dim 2", ranging from -40 to 60, with tick marks at -40, -20, 0, 20, 40, and 60.
* Gridlines: Light gray dashed lines at each tick mark on both axes.
* **Top Marginal Plot:**
* Shows the density distribution of "dim 1" for both "General Text" (blue) and "Medical Text" (red).
* **Right Marginal Plot:**
* Shows the density distribution of "dim 2" for both "General Text" (blue) and "Medical Text" (red).
* **Legend (Top-Left):**
* "General Text" - Blue line
* "Medical Text" - Red line
### Detailed Analysis
* **General Text (Blue):**
* In the main plot, the density contours for "General Text" are concentrated in the upper-left quadrant, with a primary cluster around dim1 = -40 and dim2 = 20, and a secondary cluster around dim1 = 0 and dim2 = 45.
* The top marginal plot shows two peaks for "General Text" along "dim 1", one around -40 and another around 0.
* The right marginal plot shows a single peak for "General Text" along "dim 2", centered around 30.
* **Medical Text (Red):**
* In the main plot, the density contours for "Medical Text" are concentrated in the lower-right quadrant, with a primary cluster around dim1 = 20 and dim2 = -20.
* The top marginal plot shows a single peak for "Medical Text" along "dim 1", centered around 20.
* The right marginal plot shows a single peak for "Medical Text" along "dim 2", centered around -10.
### Key Observations
* The distributions of "General Text" and "Medical Text" are clearly separated in the two-dimensional space.
* "General Text" tends to have higher values for "dim 2" compared to "Medical Text".
* "Medical Text" tends to have higher values for "dim 1" compared to "General Text".
* The marginal distributions confirm the separation observed in the main plot.
### Interpretation
The density plot suggests that "General Text" and "Medical Text" can be distinguished based on their values along "dim 1" and "dim 2". The separation of the density contours indicates that these two dimensions capture some underlying characteristics that differentiate the two types of text. The clustering of "General Text" in the upper-left quadrant and "Medical Text" in the lower-right quadrant suggests that these dimensions might represent different aspects of the text, such as topic, style, or complexity. Further analysis would be needed to determine the specific meaning of "dim 1" and "dim 2" and how they relate to the characteristics of "General Text" and "Medical Text".
</details>
(c) ADEPT
Figure 6: Kernel Density Estimation of activations for Qwen3-1.7B-Base under different configurations. Our layer extension strategy enables effective parameter decoupling. Expanded layers: 22, 23, 25, and 27.
To examine the effectiveness of our layer extension strategy, we conduct activation distribution analysis across multiple backbones. For each model, we first identify the most domain-adaptable layer (i.e., the layer with the lowest $I_{\text{layer}}$ ). We then randomly sample 500 instances from both the Medical and General corpora, compute activations at the selected layer, and visualize their distributions using Kernel Density Estimation (KDE). The following three configurations are compared: (1) the original base model, where we visualize the most domain-adaptable layer; (2) direct decoupling without expansion (w/o Stage-1), where we visualize the same most domain-adaptable layer; (3) our method with expanded layers, where we visualize the newly created expanded layer (copied from the most domain-adaptable layer).
Figure 6 presents the results from three different model configurations, providing compelling evidence for the advantages of our proposed approach.
Figure 6 a) shows the activation distribution in layer 27 of the original Qwen3-1.7B-Base model. The substantial overlap between general and medical text distributions indicates strong parameter coupling, which is an expected consequence of mixed-domain pretraining. This coupling makes it challenging to achieve clean separation of domain-specific functionalities through conventional fine-tuning approaches. However, the divergence between the peak values in the general domain and the medical domain also indicates the potential for decoupling.
This coupling phenomenon persisted in our ablation studies with only the decoupling method in Figure 6 b). Despite our attempts to decouple the medical and general modules when training, the model’s activation distributions remained largely entangled (the graph still shows substantial overlap), failing to achieve distinct separation between domains. This observation further supports our argument that pre-existing parameter coupling from mixed-domain pretraining creates inherent challenges for direct decoupling approaches.
In contrast, Figure 6 c) demonstrates the activation distribution in layer 31 of our extended model, where we first expanded the model by copying parameters from layer 27 and then applied decoupling training. The clear separation between general and medical text distributions suggests successful parameter decoupling. This superior decoupling effect can be attributed to our “blank slate” approach: the extended layers, while initialized with copied parameters, provide a fresh parameter space that hasn’t been constrained by mixed-domain pretraining. During decoupling training, these extended layers can adapt more freely to domain-specific patterns through gradient descent and importance-based learning rate adjustments.
To validate our hypothesis, we also examine the effect of applying in Qwen3-4B-Base (Figure 7), Qwen3-8B-Base (Figure 8), Llama3-8B (Figure 9). The results indicate limited separation between domains, which supports our argument that the entangled parameters from mixed-domain pretraining are challenging to decouple through training alone.
These findings demonstrate that our layer extension strategy provides a more effective pathway for parameter decoupling compared to direct decoupling training. By creating a new parameter space through layer extension, we avoid the constraints of pre-existing parameter coupling, allowing for cleaner separation of domain-specific functionalities during subsequent training. This approach offers a promising direction for developing more modular and domain-adaptable language models.
<details>
<summary>figures/4B_Base_Layer35.jpg Details</summary>
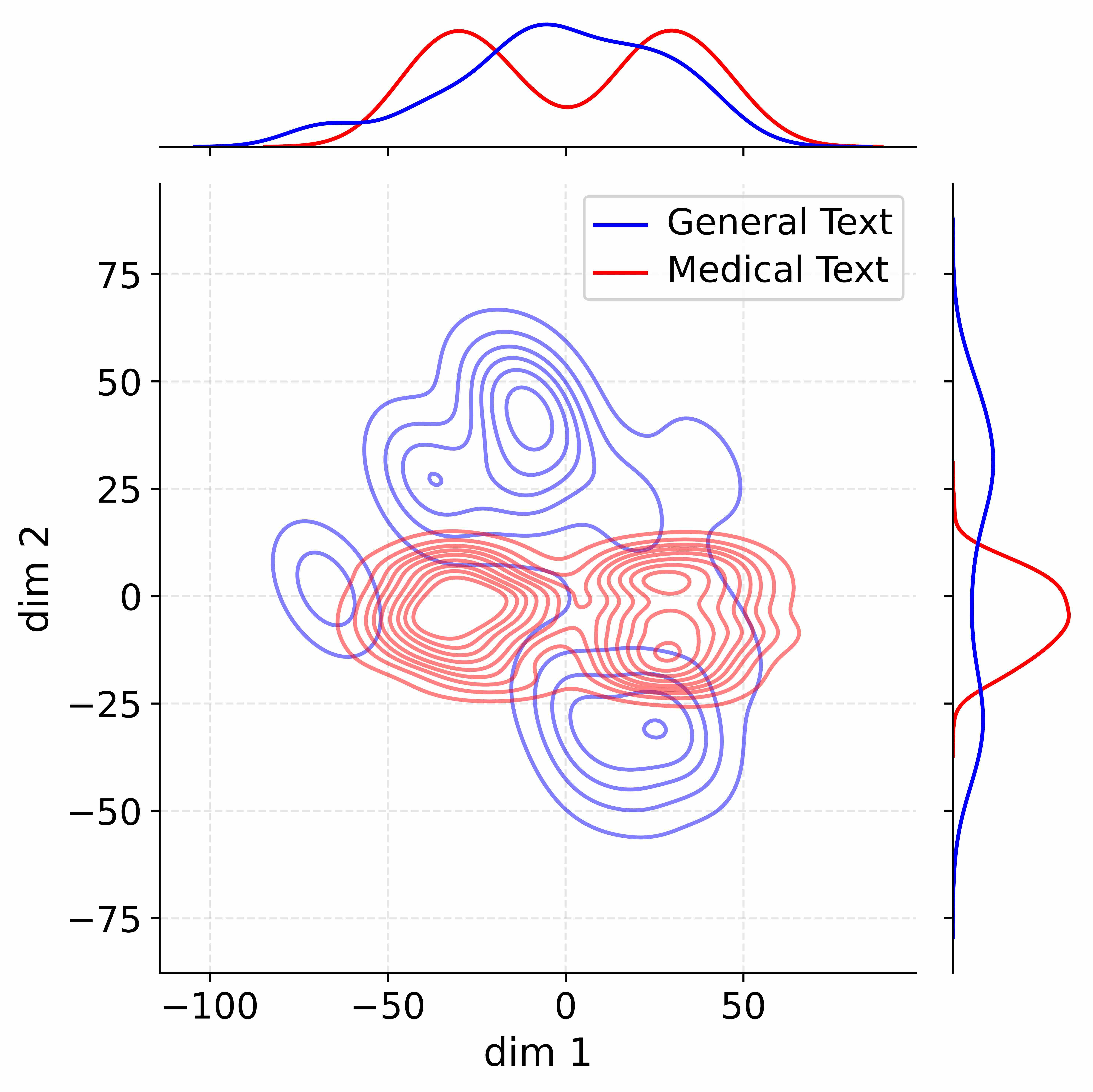
### Visual Description
## Density Plot: General Text vs. Medical Text
### Overview
The image is a density plot comparing the distribution of "General Text" and "Medical Text" across two dimensions (dim 1 and dim 2). It includes density contours in the main plot area and marginal density plots along the top and right edges.
### Components/Axes
* **Main Plot:**
* X-axis: "dim 1", ranging from -100 to 50.
* Y-axis: "dim 2", ranging from -75 to 75.
* Gridlines: Light gray, dashed lines at intervals of 25 on both axes.
* Contours:
* Blue contours represent "General Text".
* Red contours represent "Medical Text".
* **Top Marginal Plot:**
* X-axis: Aligned with the main plot's x-axis ("dim 1").
* Y-axis: Density of the data along dim 1.
* Curves:
* Blue curve represents "General Text".
* Red curve represents "Medical Text".
* **Right Marginal Plot:**
* X-axis: Density of the data along dim 2.
* Y-axis: Aligned with the main plot's y-axis ("dim 2").
* Curves:
* Blue curve represents "General Text".
* Red curve represents "Medical Text".
* **Legend:** Located at the top-center of the plot.
* Blue line: "General Text"
* Red line: "Medical Text"
### Detailed Analysis
* **General Text (Blue):**
* Main Plot: The density contours are concentrated in two main regions: one around (-50, 25) and another around (0, 50). There's also a smaller concentration around (25, -25).
* Top Marginal Plot: The blue curve shows two peaks, one around -50 and another around 0.
* Right Marginal Plot: The blue curve shows a peak around 25.
* **Medical Text (Red):**
* Main Plot: The density contours are concentrated in a region around (0, 0), extending towards (50, 0) and (-50, 0).
* Top Marginal Plot: The red curve shows a peak around 0.
* Right Marginal Plot: The red curve shows a peak around 0.
### Key Observations
* The "General Text" distribution is more spread out and has two distinct clusters along dim 1.
* The "Medical Text" distribution is more concentrated around the origin (0, 0) in both dimensions.
* There is some overlap between the two distributions, particularly in the region around (0, 0).
### Interpretation
The density plot suggests that "General Text" and "Medical Text" have different characteristics when projected onto these two dimensions. "General Text" exhibits a bimodal distribution along dim 1, indicating two distinct patterns or clusters within the data. "Medical Text," on the other hand, is more centrally located, suggesting a more uniform or concentrated pattern. The overlap between the two distributions indicates some degree of similarity or shared characteristics, but overall, the two types of text appear to be distinguishable based on these dimensions. The marginal distributions provide further insight into the distribution of each text type along each individual dimension.
</details>
(a) Vanilla
<details>
<summary>figures/4B_only_grad_Layer35.jpg Details</summary>
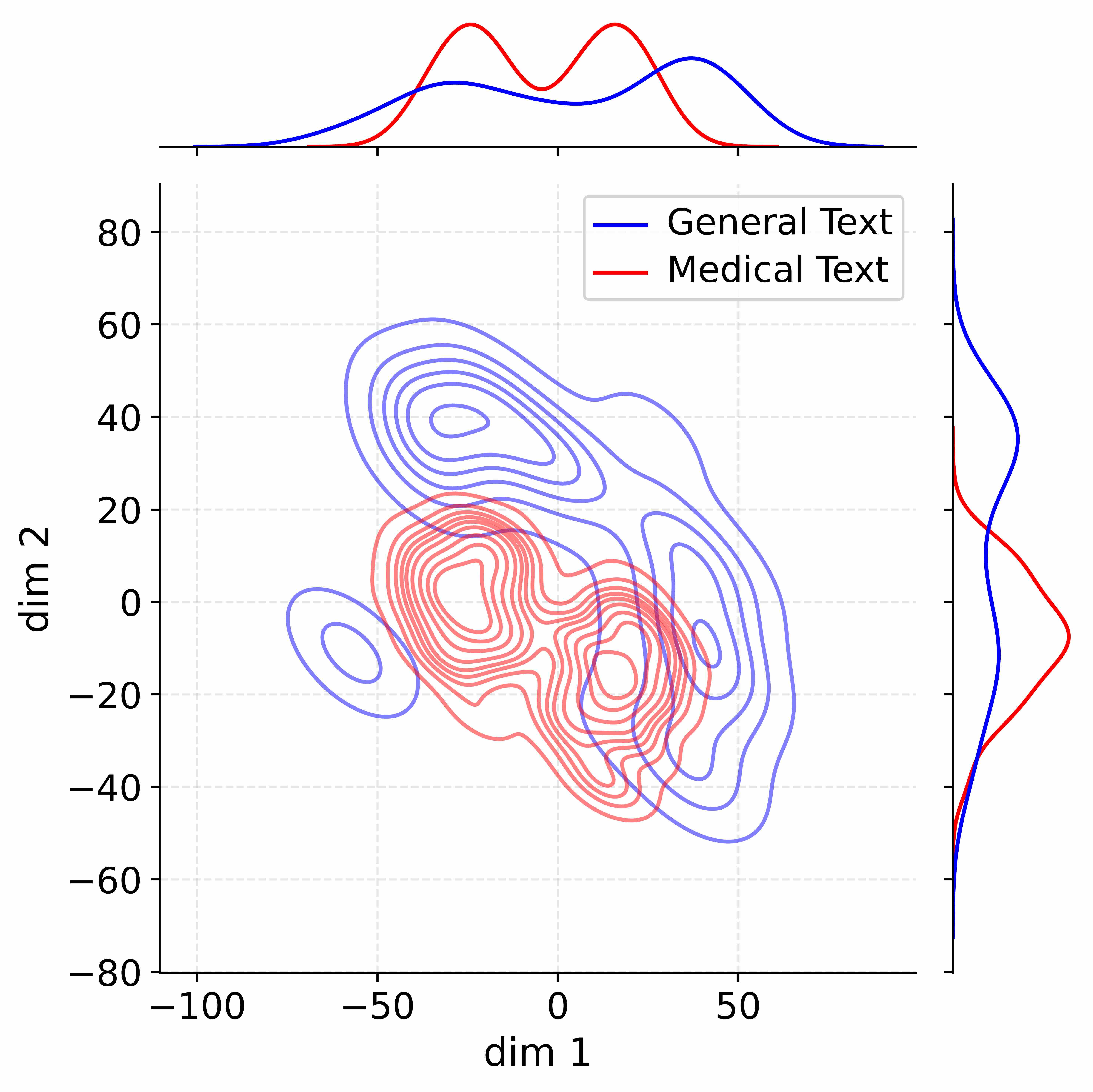
### Visual Description
## Density Plot: General Text vs. Medical Text
### Overview
The image is a density plot comparing "General Text" and "Medical Text" across two dimensions (dim 1 and dim 2). It includes a 2D contour plot in the center, with marginal density plots along the top (dim 1) and right side (dim 2). The plot uses blue for "General Text" and red for "Medical Text".
### Components/Axes
* **X-axis (dim 1):** Ranges from -100 to 50, with tick marks at -100, -50, 0, and 50.
* **Y-axis (dim 2):** Ranges from -80 to 80, with tick marks at -80, -60, -40, -20, 0, 20, 40, 60, and 80.
* **Legend (top-center):**
* Blue line: "General Text"
* Red line: "Medical Text"
* **Top Marginal Plot:** Shows the density distribution of dim 1 for both "General Text" (blue) and "Medical Text" (red).
* **Right Marginal Plot:** Shows the density distribution of dim 2 for both "General Text" (blue) and "Medical Text" (red).
* **Grid:** Light gray dashed lines provide visual structure.
### Detailed Analysis
* **General Text (Blue):**
* **2D Contour:** The highest density area is centered around dim 1 = -50 and dim 2 = 40. There's a secondary density area around dim 1 = 50 and dim 2 = 0.
* **Top Marginal Plot (dim 1):** The blue line shows a bimodal distribution, with peaks around -50 and 50.
* **Right Marginal Plot (dim 2):** The blue line shows a unimodal distribution, with a peak around 40.
* **Medical Text (Red):**
* **2D Contour:** The highest density area is centered around dim 1 = 0 and dim 2 = 0.
* **Top Marginal Plot (dim 1):** The red line shows a bimodal distribution, with peaks around -10 and 20.
* **Right Marginal Plot (dim 2):** The red line shows a unimodal distribution, with a peak around 0.
### Key Observations
* "General Text" has higher density in the upper-left and upper-right quadrants, while "Medical Text" is concentrated around the origin.
* The marginal distributions confirm these observations, showing distinct peaks for each type of text along both dimensions.
* The "General Text" distribution along dim 1 is bimodal, suggesting two distinct clusters within the general text data.
* The "Medical Text" distribution along dim 1 is also bimodal, but the peaks are closer together compared to "General Text".
### Interpretation
The density plot visualizes the distribution of "General Text" and "Medical Text" in a two-dimensional space. The separation of the density contours suggests that these two types of text occupy different regions in this space, indicating that they can be distinguished based on the features represented by dim 1 and dim 2. The bimodal distributions along dim 1 for both text types suggest that there are sub-clusters within each category. The "General Text" clusters are more separated than the "Medical Text" clusters, implying greater variability within the "General Text" data.
</details>
(b) W/o stage1
<details>
<summary>figures/4B_Layer39.jpg Details</summary>
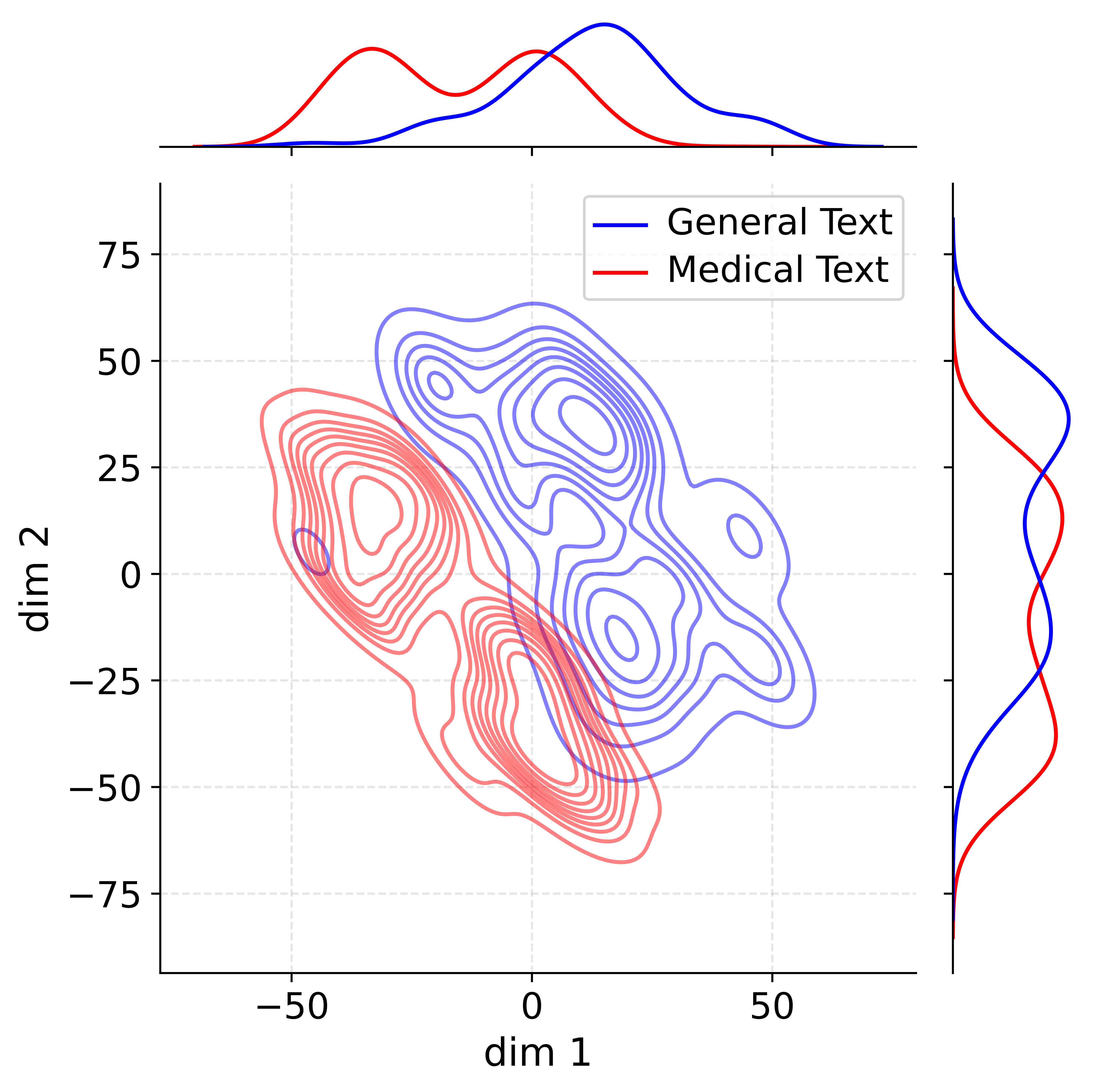
### Visual Description
## Contour Plot: Distribution of Text Types in Two Dimensions
### Overview
The image is a contour plot showing the distribution of two types of text, "General Text" and "Medical Text," across two dimensions, labeled "dim 1" and "dim 2." Density plots are shown along the top and right margins, representing the marginal distributions of each text type along each dimension.
### Components/Axes
* **Main Plot:** A 2D contour plot with "dim 1" on the x-axis and "dim 2" on the y-axis.
* X-axis ("dim 1") ranges from approximately -75 to 75, with labeled ticks at -50, 0, and 50.
* Y-axis ("dim 2") ranges from approximately -75 to 75, with labeled ticks at -75, -50, -25, 0, 25, 50, and 75.
* The plot has a light gray grid.
* **Top Density Plot:** Shows the distribution of "dim 1" for both "General Text" (blue) and "Medical Text" (red).
* **Right Density Plot:** Shows the distribution of "dim 2" for both "General Text" (blue) and "Medical Text" (red).
* **Legend:** Located in the top-center of the plot.
* "General Text" is represented by a blue line.
* "Medical Text" is represented by a red line.
### Detailed Analysis
* **General Text (Blue):**
* In the main plot, the blue contours are centered around dim1 = 20 and dim2 = 30. The contours are elongated diagonally.
* Top density plot: The blue line shows a distribution with a peak around dim1 = 20.
* Right density plot: The blue line shows a distribution with a peak around dim2 = 30.
* **Medical Text (Red):**
* In the main plot, the red contours are centered around dim1 = -30 and dim2 = -20. The contours are elongated diagonally.
* Top density plot: The red line shows a distribution with a peak around dim1 = -30.
* Right density plot: The red line shows a distribution with a peak around dim2 = -20.
### Key Observations
* The "General Text" and "Medical Text" distributions are clearly separated in the 2D space.
* The marginal distributions in the top and right plots confirm the separation observed in the main plot.
* The distributions are somewhat elongated, suggesting a correlation between "dim 1" and "dim 2" for both text types.
### Interpretation
The plot visualizes the distribution of "General Text" and "Medical Text" across two dimensions. The clear separation between the two distributions suggests that "dim 1" and "dim 2" are effective at distinguishing between these two types of text. The marginal distributions provide further insight into how each dimension contributes to this separation. The elongation of the contours suggests that the two dimensions are correlated within each text type. This could imply that the underlying features represented by "dim 1" and "dim 2" are related for both general and medical text.
</details>
(c) ADEPT
Figure 7: Visualization of activation distributions for Qwen3-4B-Base model configurations showing the effectiveness of our layer extension strategy for parameter decoupling. We expand the layer 28, 30, 31, 35 of Qwen3-4B-Base.
<details>
<summary>figures/8B_Base_Layer30.jpg Details</summary>
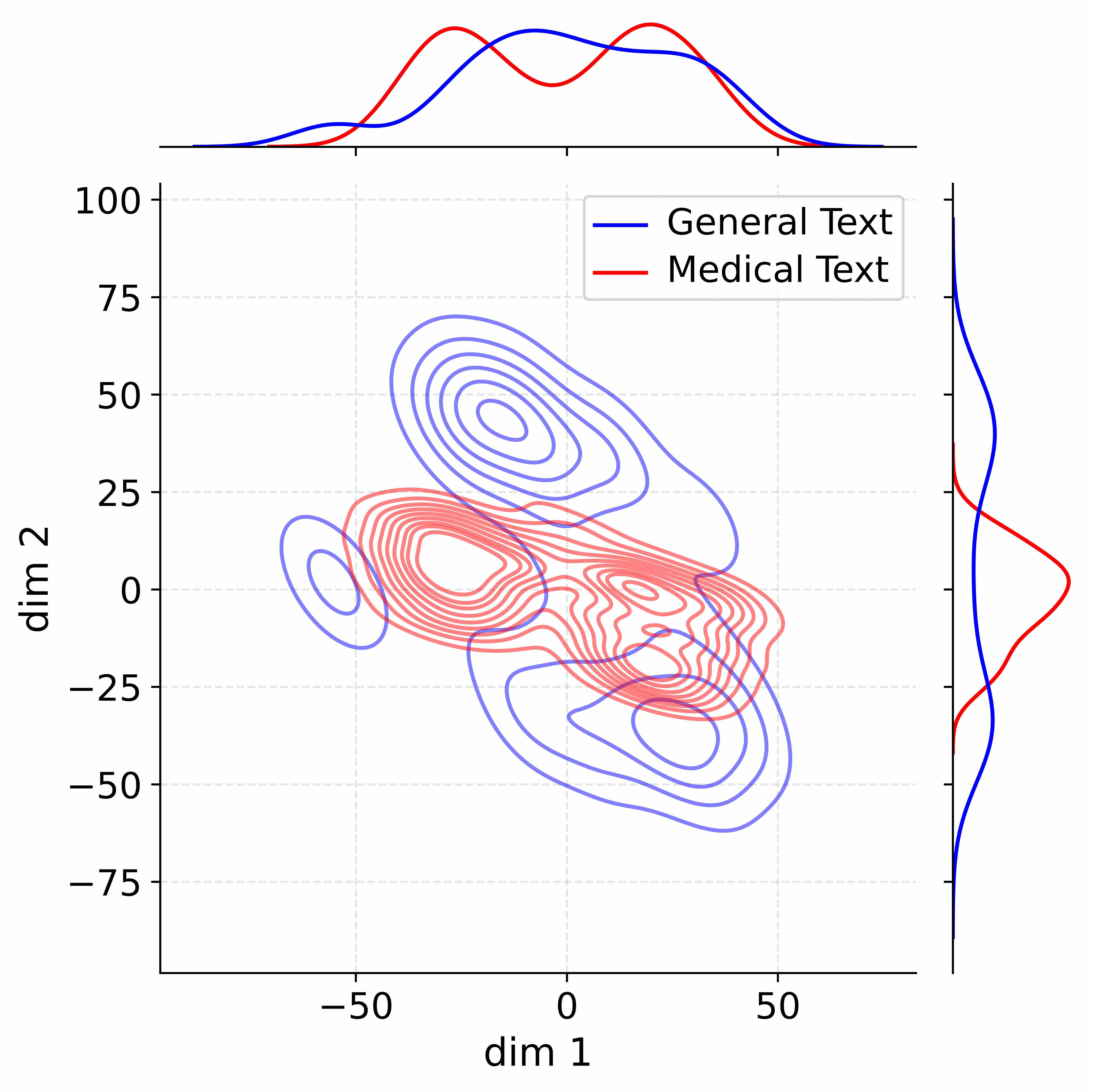
### Visual Description
## Contour Plot: General Text vs. Medical Text
### Overview
The image is a contour plot comparing the distribution of "General Text" and "Medical Text" across two dimensions (dim 1 and dim 2). The plot includes marginal distributions for each dimension along the top and right edges.
### Components/Axes
* **Main Plot:** A 2D contour plot with "dim 1" on the x-axis and "dim 2" on the y-axis.
* X-axis (dim 1): Ranges from approximately -75 to 75, with labeled ticks at -50, 0, and 50.
* Y-axis (dim 2): Ranges from approximately -75 to 100, with labeled ticks at -75, -50, -25, 0, 25, 50, 75, and 100.
* Gridlines: Light gray dashed lines.
* **Top Marginal Plot:** A 1D density plot showing the distribution of "dim 1" for both "General Text" (blue) and "Medical Text" (red).
* **Right Marginal Plot:** A 1D density plot showing the distribution of "dim 2" for both "General Text" (blue) and "Medical Text" (red).
* **Legend:** Located in the top-center of the plot.
* "General Text": Represented by blue lines.
* "Medical Text": Represented by red lines.
### Detailed Analysis
* **General Text (Blue):**
* **Main Plot:** The contours show two distinct clusters. One is centered around dim1 = -50 and dim2 = 20, and the other is centered around dim1 = 25 and dim2 = -25.
* **Top Marginal Plot:** The distribution for dim 1 shows two peaks, one around -50 and another around 25.
* **Right Marginal Plot:** The distribution for dim 2 shows a single peak around 25.
* **Medical Text (Red):**
* **Main Plot:** The contours show a cluster centered around dim1 = 20 and dim2 = -10.
* **Top Marginal Plot:** The distribution for dim 1 shows a single peak around 20.
* **Right Marginal Plot:** The distribution for dim 2 shows a single peak around -10.
### Key Observations
* "General Text" has a bimodal distribution along "dim 1", suggesting two distinct groups or characteristics within the general text data.
* "Medical Text" has a more concentrated distribution compared to "General Text" in both dimensions.
* The clusters of "General Text" and "Medical Text" overlap slightly, indicating some similarity between the two types of text.
### Interpretation
The contour plot visualizes the differences in the distribution of "General Text" and "Medical Text" across two dimensions. The bimodal distribution of "General Text" along "dim 1" suggests that general text data may be composed of two distinct sub-groups. "Medical Text", on the other hand, appears to be more homogeneous, with a single cluster in the 2D space. The marginal distributions provide further insight into the distribution of each text type along individual dimensions. The overlap between the two distributions suggests that there are some shared characteristics between "General Text" and "Medical Text", but overall, they exhibit distinct patterns.
</details>
(a) Vanilla
<details>
<summary>figures/8B_only_grad_Layer30.jpg Details</summary>
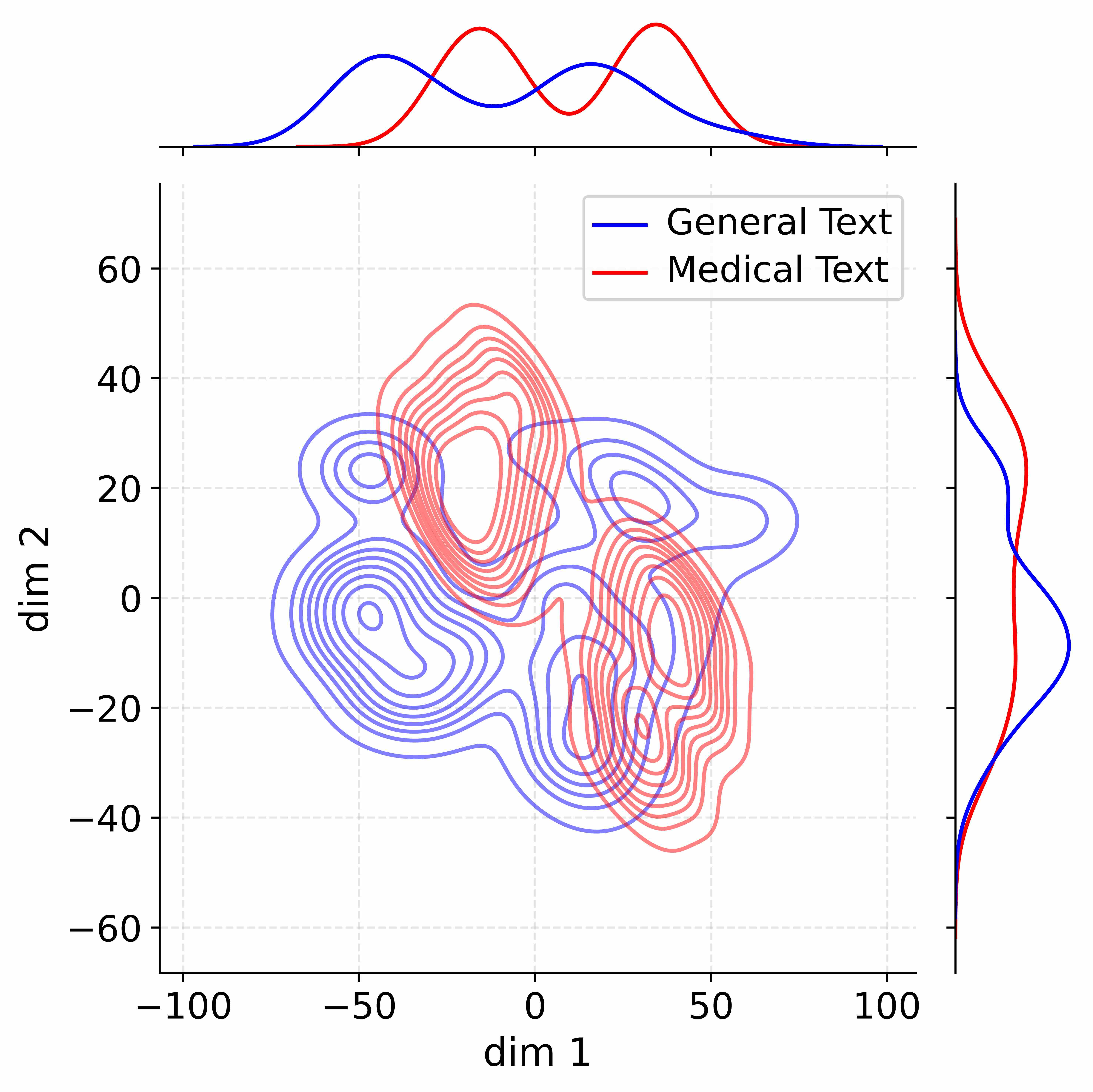
### Visual Description
## Contour Plot: General Text vs. Medical Text
### Overview
The image is a contour plot comparing the distribution of "General Text" and "Medical Text" across two dimensions (dim 1 and dim 2). The plot includes marginal distributions along the top and right edges. The "General Text" data is represented by blue lines, while the "Medical Text" data is represented by red lines.
### Components/Axes
* **Main Plot:** A 2D contour plot showing the distribution of data points.
* X-axis (dim 1): Ranges from -100 to 100, with tick marks at -100, -50, 0, 50, and 100.
* Y-axis (dim 2): Ranges from -60 to 60, with tick marks at -60, -40, -20, 0, 20, 40, and 60.
* **Top Marginal Plot:** Shows the distribution of data along the x-axis (dim 1).
* Blue line: Represents the distribution of "General Text" along dim 1. It has two peaks, one around -50 and another around 20.
* Red line: Represents the distribution of "Medical Text" along dim 1. It has two peaks, one around 20 and another around 50.
* **Right Marginal Plot:** Shows the distribution of data along the y-axis (dim 2).
* Blue line: Represents the distribution of "General Text" along dim 2. It has a single peak around 0.
* Red line: Represents the distribution of "Medical Text" along dim 2. It has a single peak around 0, but is more concentrated than the blue line.
* **Legend:** Located in the top-right corner.
* Blue line: "General Text"
* Red line: "Medical Text"
### Detailed Analysis
* **General Text (Blue):**
* In the main plot, the contours are concentrated in two main regions: one around (-50, 0) and another around (0, 0).
* The top marginal plot shows two peaks, indicating that "General Text" data is distributed across two distinct values of dim 1.
* The right marginal plot shows a single peak around 0, indicating that "General Text" data is concentrated around 0 for dim 2.
* **Medical Text (Red):**
* In the main plot, the contours are concentrated in two main regions: one around (0, 30) and another around (30, 0).
* The top marginal plot shows two peaks, indicating that "Medical Text" data is distributed across two distinct values of dim 1.
* The right marginal plot shows a single peak around 0, indicating that "Medical Text" data is concentrated around 0 for dim 2.
### Key Observations
* The distributions of "General Text" and "Medical Text" are distinct but overlapping.
* "General Text" has a higher concentration around (-50, 0), while "Medical Text" has a higher concentration around (30, 0).
* Both distributions are centered around 0 for dim 2.
### Interpretation
The contour plot visualizes the differences in the distribution of "General Text" and "Medical Text" across two dimensions. The distinct concentrations suggest that these two types of text have different characteristics in terms of dim 1 and dim 2. The overlap indicates that there are some similarities between the two types of text. The marginal distributions provide additional information about the distribution of each type of text along each dimension. The plot suggests that "General Text" and "Medical Text" can be distinguished based on their distribution in this two-dimensional space.
</details>
(b) W/o stage1
<details>
<summary>figures/8B_final_grad_Layer30.jpg Details</summary>
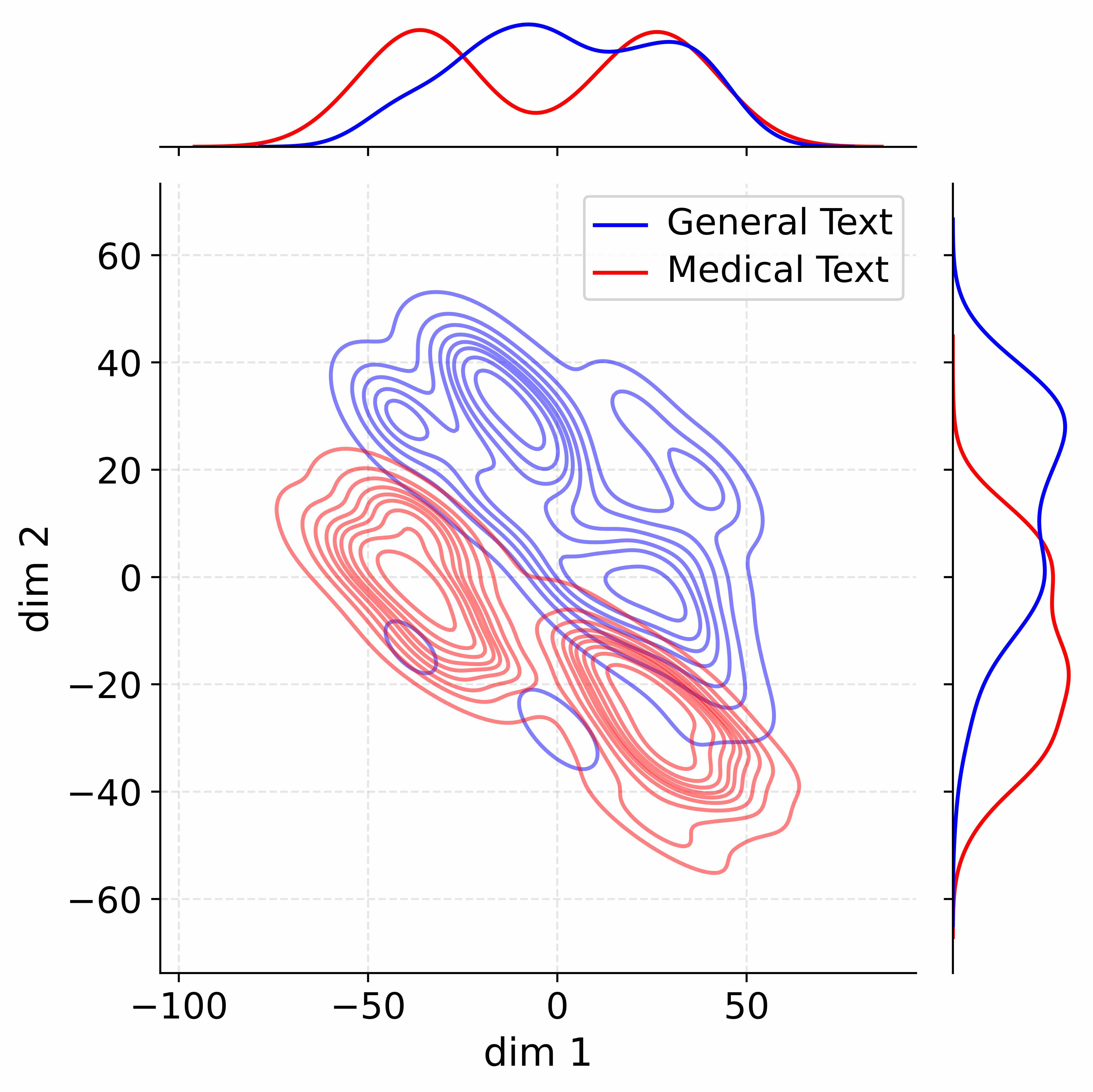
### Visual Description
## Contour Plot: Text Distribution in Two Dimensions
### Overview
The image is a contour plot showing the distribution of "General Text" and "Medical Text" along two dimensions, labeled "dim 1" and "dim 2". Marginal distributions are shown along the top and right edges of the plot.
### Components/Axes
* **X-axis (dim 1):** Ranges from -100 to 50, with tick marks at -100, -50, 0, and 50.
* **Y-axis (dim 2):** Ranges from -60 to 60, with tick marks at -60, -40, -20, 0, 20, 40, and 60.
* **Contour Lines:** Represent density levels for each type of text.
* **Marginal Distributions:** Density plots along the top (for dim 1) and right (for dim 2).
* **Legend (top-center):**
* Blue line: "General Text"
* Red line: "Medical Text"
### Detailed Analysis
**1. General Text (Blue):**
* **Contour Plot:** The blue contours are concentrated in two main regions. One is centered around dim1 = -25 and dim2 = 30, and the other is centered around dim1 = 30 and dim2 = -10.
* **Marginal Distribution (Top):** The blue line shows two peaks. One peak is around -25 and the other is around 30.
* **Marginal Distribution (Right):** The blue line shows a peak around 20.
**2. Medical Text (Red):**
* **Contour Plot:** The red contours are concentrated in two main regions. One is centered around dim1 = -40 and dim2 = -10, and the other is centered around dim1 = 0 and dim2 = -20.
* **Marginal Distribution (Top):** The red line shows two peaks. One peak is around -40 and the other is around 0.
* **Marginal Distribution (Right):** The red line shows a peak around -10.
### Key Observations
* The "General Text" distribution has two distinct clusters, one with a positive value for dim 2 and one with a negative value.
* The "Medical Text" distribution also has two clusters, both with negative values for dim 2.
* There is some overlap between the two distributions, particularly around dim1 = 0 and dim2 = -20.
### Interpretation
The contour plot visualizes the distribution of "General Text" and "Medical Text" across two dimensions. The separation of the contour clusters suggests that these two types of text occupy different regions in the two-dimensional space defined by "dim 1" and "dim 2". The marginal distributions provide further insight into the distribution of each text type along each individual dimension. The overlap between the distributions indicates some similarity or shared characteristics between the two types of text. The nature of "dim 1" and "dim 2" is not specified, so further context is needed to interpret the meaning of these dimensions and the observed distributions.
</details>
(c) ADEPT
Figure 8: Kernel Density Estimation of activations for Qwen3-8B-Base, showing that our layer extension strategy enables clear parameter decoupling. We expand layers 26, 28, 29, and 30.
<details>
<summary>figures/Llama_8B_Layer30.jpg Details</summary>
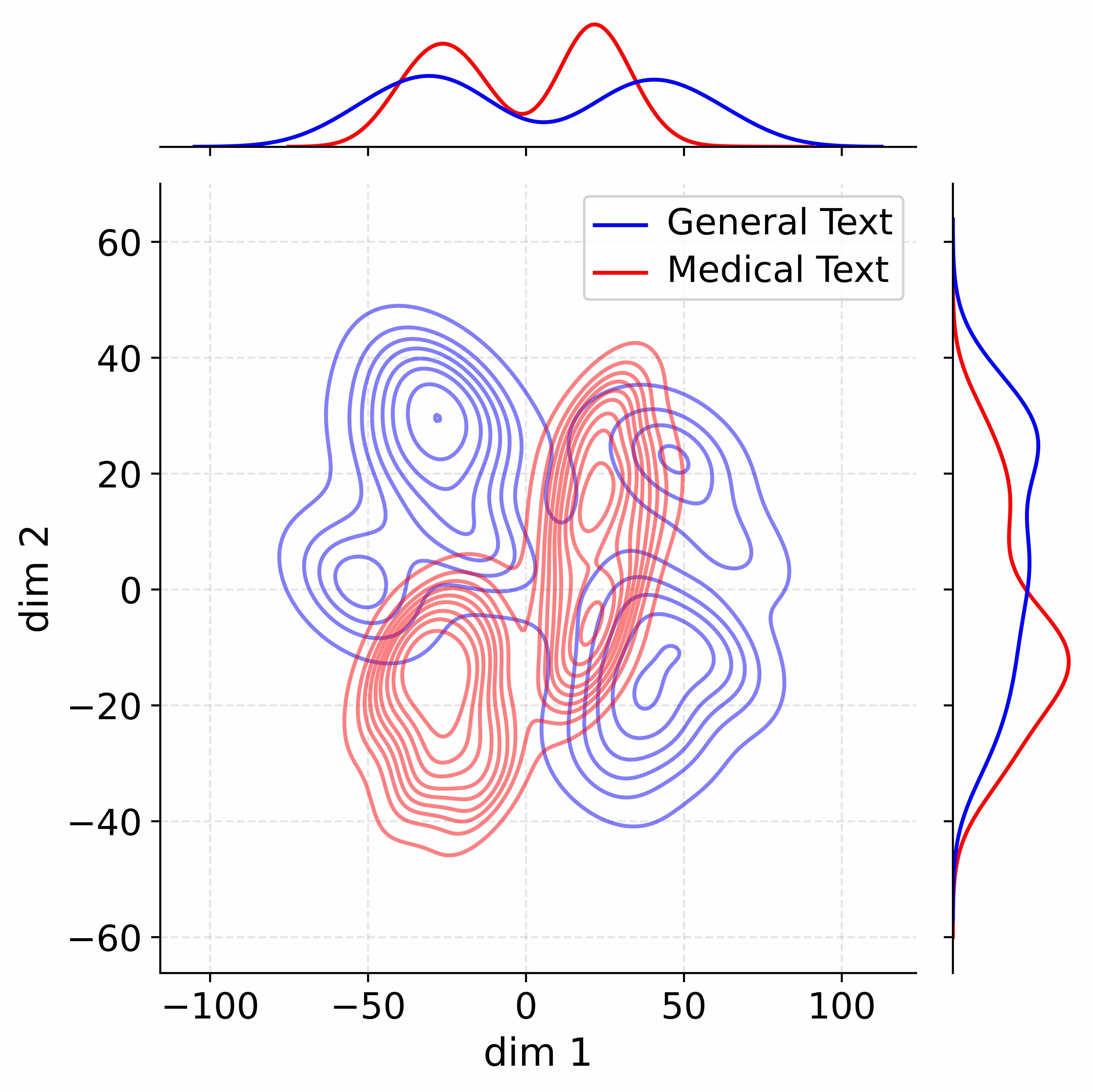
### Visual Description
## Density Plot: General Text vs. Medical Text
### Overview
The image is a density plot comparing "General Text" and "Medical Text" across two dimensions, labeled "dim 1" and "dim 2". The plot includes density contours for both categories, along with marginal density plots along the top and right axes.
### Components/Axes
* **Main Plot:**
* X-axis: "dim 1", ranging from -100 to 100, with tick marks at -100, -50, 0, 50, and 100.
* Y-axis: "dim 2", ranging from -60 to 60, with tick marks at -60, -40, -20, 0, 20, 40, and 60.
* Gridlines: Light gray dashed lines.
* Contours:
* Blue contours represent "General Text".
* Red contours represent "Medical Text".
* **Top Marginal Plot:**
* X-axis: Aligned with the main plot's "dim 1" axis.
* Y-axis: Density.
* Curves:
* Blue curve represents the marginal density of "General Text" along "dim 1".
* Red curve represents the marginal density of "Medical Text" along "dim 1".
* **Right Marginal Plot:**
* Y-axis: Aligned with the main plot's "dim 2" axis.
* X-axis: Density.
* Curves:
* Blue curve represents the marginal density of "General Text" along "dim 2".
* Red curve represents the marginal density of "Medical Text" along "dim 2".
* **Legend:** Located in the top-right corner.
* "General Text" - Blue line.
* "Medical Text" - Red line.
### Detailed Analysis or Content Details
* **General Text (Blue):**
* Main Plot: The blue contours show two main clusters. One is centered around dim1 = -50 and dim2 = 20, and the other is centered around dim1 = 50 and dim2 = 0.
* Top Marginal Plot: The blue curve shows two peaks, one around dim1 = -50 and another around dim1 = 50.
* Right Marginal Plot: The blue curve shows a peak around dim2 = 20.
* **Medical Text (Red):**
* Main Plot: The red contours show one main cluster centered around dim1 = 0 and dim2 = -20, and another smaller cluster around dim1 = 30 and dim2 = 30.
* Top Marginal Plot: The red curve shows two peaks, one around dim1 = 0 and another around dim1 = 30.
* Right Marginal Plot: The red curve shows a peak around dim2 = -20.
### Key Observations
* The "General Text" distribution has two distinct clusters, while the "Medical Text" distribution has a more concentrated cluster.
* The marginal distributions confirm the clustering observed in the main plot.
* There is some overlap between the "General Text" and "Medical Text" distributions, but they are largely distinct.
### Interpretation
The density plot visualizes the distribution of "General Text" and "Medical Text" across two dimensions. The separation of the clusters suggests that "dim 1" and "dim 2" are able to differentiate between the two types of text. The "General Text" distribution is more spread out, indicating greater variability, while the "Medical Text" distribution is more concentrated, suggesting more consistency. The marginal distributions provide further insight into the distribution of each text type along each individual dimension.
</details>
(a) Vanilla
<details>
<summary>figures/Llama_8B_only_grad_Layer30.jpg Details</summary>
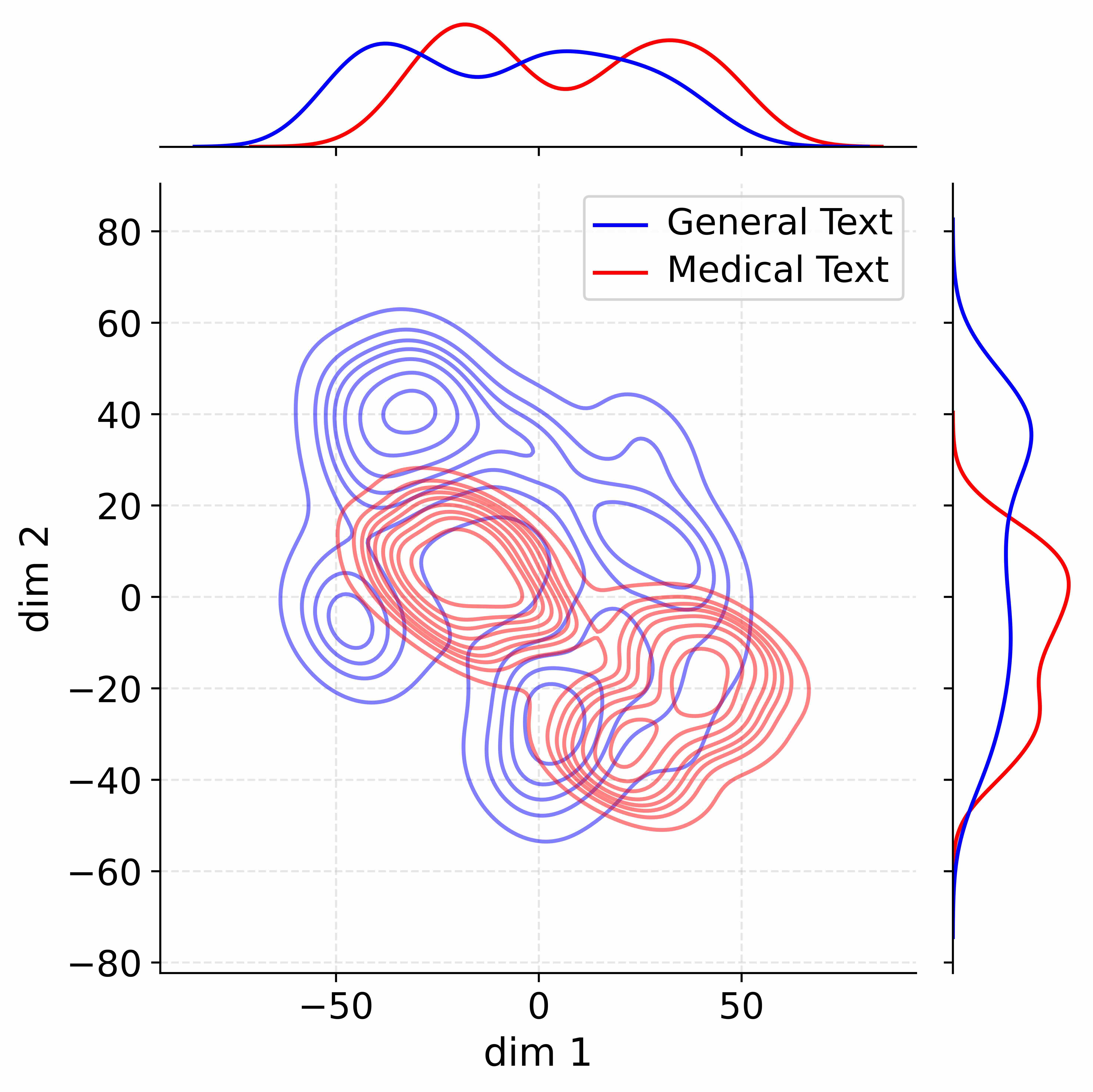
### Visual Description
## Contour Plot: General Text vs. Medical Text
### Overview
The image is a contour plot comparing the distribution of "General Text" and "Medical Text" across two dimensions (dim 1 and dim 2). The plot includes marginal density plots along the top and right edges, showing the distribution of each text type along each individual dimension.
### Components/Axes
* **Main Plot:** A 2D contour plot showing the density distribution of "General Text" (blue) and "Medical Text" (red).
* X-axis (dim 1): Ranges from -50 to 50.
* Y-axis (dim 2): Ranges from -80 to 80.
* Gridlines: Light gray dashed lines.
* **Top Marginal Plot:** Density plot showing the distribution of "General Text" (blue) and "Medical Text" (red) along dim 1.
* **Right Marginal Plot:** Density plot showing the distribution of "General Text" (blue) and "Medical Text" (red) along dim 2.
* **Legend:** Located in the top-center of the plot.
* "General Text" - Blue line
* "Medical Text" - Red line
### Detailed Analysis or ### Content Details
**Main Plot:**
* **General Text (Blue):**
* The highest density area is centered around dim 1 = -50 and dim 2 = 40.
* The contours suggest a cluster of data points in the upper-left quadrant.
* **Medical Text (Red):**
* The highest density area is centered around dim 1 = 40 and dim 2 = -20.
* The contours suggest a cluster of data points in the lower-right quadrant.
* There is some overlap between the "General Text" and "Medical Text" distributions, particularly around the center of the plot.
**Top Marginal Plot (dim 1):**
* **General Text (Blue):**
* The distribution peaks around dim 1 = -50.
* The distribution has a smaller peak around dim 1 = 20.
* **Medical Text (Red):**
* The distribution peaks around dim 1 = 40.
* The distribution has a smaller peak around dim 1 = -10.
**Right Marginal Plot (dim 2):**
* **General Text (Blue):**
* The distribution peaks around dim 2 = 60.
* **Medical Text (Red):**
* The distribution peaks around dim 2 = -20.
### Key Observations
* "General Text" is primarily concentrated in the upper-left quadrant, while "Medical Text" is concentrated in the lower-right quadrant.
* The marginal distributions confirm that "General Text" tends to have higher values along dim 2 and lower values along dim 1, while "Medical Text" tends to have lower values along dim 2 and higher values along dim 1.
* There is some overlap between the two distributions, indicating that some data points share similar values along both dimensions.
### Interpretation
The contour plot visualizes the differences in the distribution of "General Text" and "Medical Text" across two dimensions. The separation of the two clusters suggests that these dimensions are useful for distinguishing between the two types of text. The marginal distributions provide further insight into the characteristics of each text type along each individual dimension. The overlap between the distributions indicates that the separation is not perfect, and some data points may be misclassified if only these two dimensions are considered.
</details>
(b) W/o stage1
<details>
<summary>figures/Llama_8B_final_grad_Layer30.jpg Details</summary>
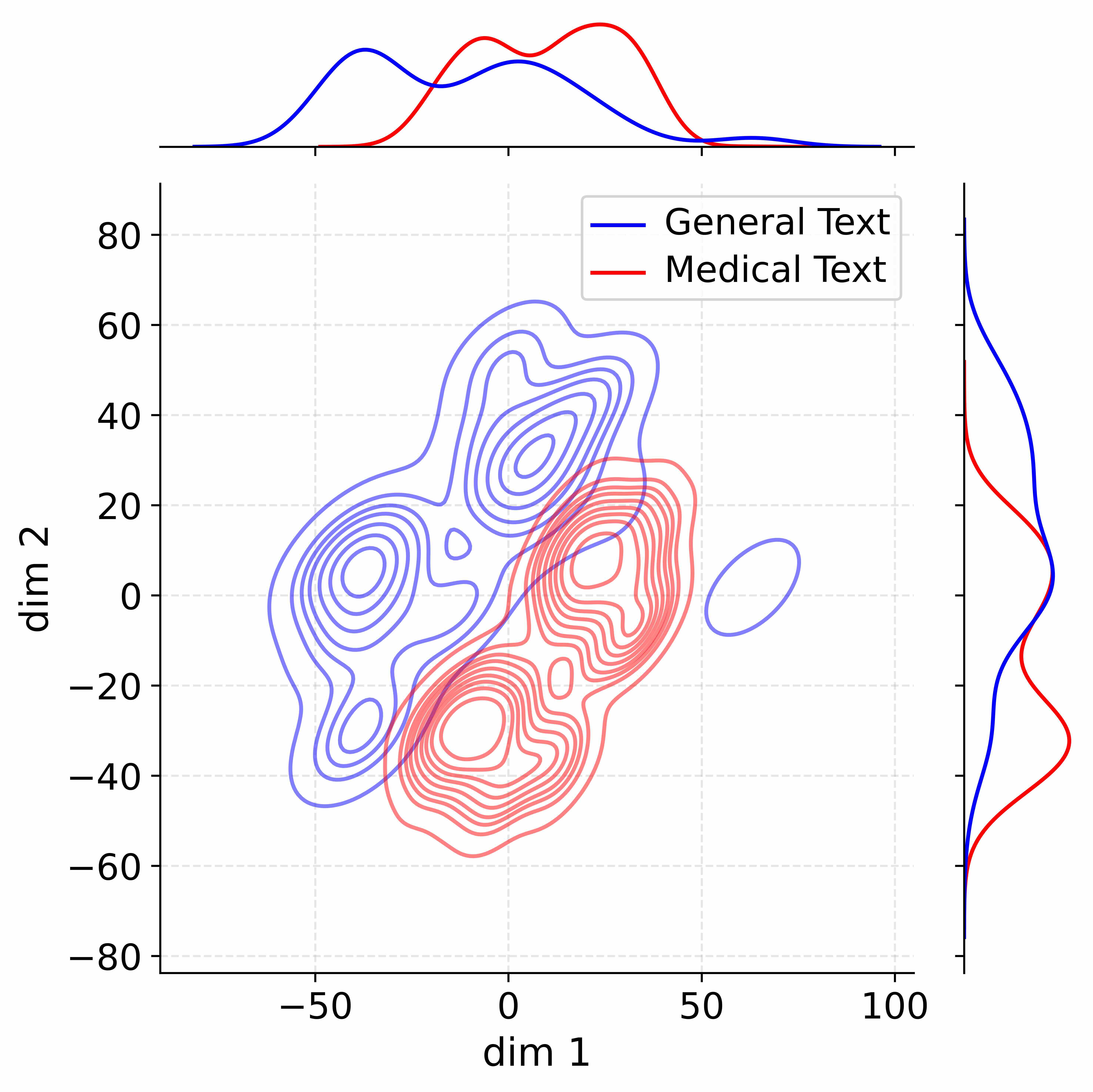
### Visual Description
## Density Plot: General Text vs. Medical Text
### Overview
The image is a density plot comparing "General Text" and "Medical Text" across two dimensions, labeled "dim 1" and "dim 2". The plot includes contour lines representing the density of each type of text, along with marginal density plots along the top and right axes.
### Components/Axes
* **Main Plot:** A 2D density plot with "dim 1" on the x-axis and "dim 2" on the y-axis.
* **X-axis (dim 1):** Ranges from approximately -50 to 100, with tick marks at -50, 0, 50, and 100.
* **Y-axis (dim 2):** Ranges from approximately -80 to 80, with tick marks at -80, -60, -40, -20, 0, 20, 40, 60, and 80.
* **Legend:** Located at the top-right of the main plot.
* **Blue Line:** Represents "General Text".
* **Red Line:** Represents "Medical Text".
* **Top Marginal Plot:** Shows the density distribution of "dim 1" for both "General Text" (blue) and "Medical Text" (red).
* **Right Marginal Plot:** Shows the density distribution of "dim 2" for both "General Text" (blue) and "Medical Text" (red).
* **Gridlines:** Light gray dashed lines provide visual reference points within the main plot.
### Detailed Analysis
* **General Text (Blue):**
* **Main Plot:** The density contours are concentrated in the upper-left quadrant, indicating higher density values for "General Text" in this region. The center of the highest density is around dim1 = -40 and dim2 = 10.
* **Top Marginal Plot:** The blue line shows a peak around dim 1 = -40, indicating a higher concentration of "General Text" along this dimension.
* **Right Marginal Plot:** The blue line shows a peak around dim 2 = 10, indicating a higher concentration of "General Text" along this dimension.
* **Medical Text (Red):**
* **Main Plot:** The density contours are concentrated in the lower-right quadrant, indicating higher density values for "Medical Text" in this region. The center of the highest density is around dim1 = 20 and dim2 = -20.
* **Top Marginal Plot:** The red line shows a peak around dim 1 = 20, indicating a higher concentration of "Medical Text" along this dimension.
* **Right Marginal Plot:** The red line shows a peak around dim 2 = -20, indicating a higher concentration of "Medical Text" along this dimension.
### Key Observations
* The density plots for "General Text" and "Medical Text" are clearly separated, suggesting distinct characteristics in the two-dimensional space.
* "General Text" is more concentrated in the region where dim 1 is negative and dim 2 is positive.
* "Medical Text" is more concentrated in the region where dim 1 is positive and dim 2 is negative.
* There is some overlap between the two density plots, indicating some similarity between "General Text" and "Medical Text" in certain regions of the two-dimensional space.
### Interpretation
The density plot visualizes the distribution of "General Text" and "Medical Text" across two dimensions, "dim 1" and "dim 2". The separation of the density contours suggests that these two types of text have distinct characteristics in this two-dimensional space. The marginal density plots provide further insight into the distribution of each type of text along each dimension. The plot could be used to understand the differences in the feature space of general and medical texts, potentially for classification or clustering purposes. The dimensions "dim 1" and "dim 2" are not defined, but they likely represent some underlying features extracted from the text data.
</details>
(c) ADEPT
Figure 9: Kernel Density Estimation of activations for Llama3-8B, showing that our layer extension strategy enables clear parameter decoupling. We expand layers 22, 23, 24, and 28.
Appendix D Detailed Importance Distribution
<details>
<summary>figures/importance_1.7B_new.jpg Details</summary>
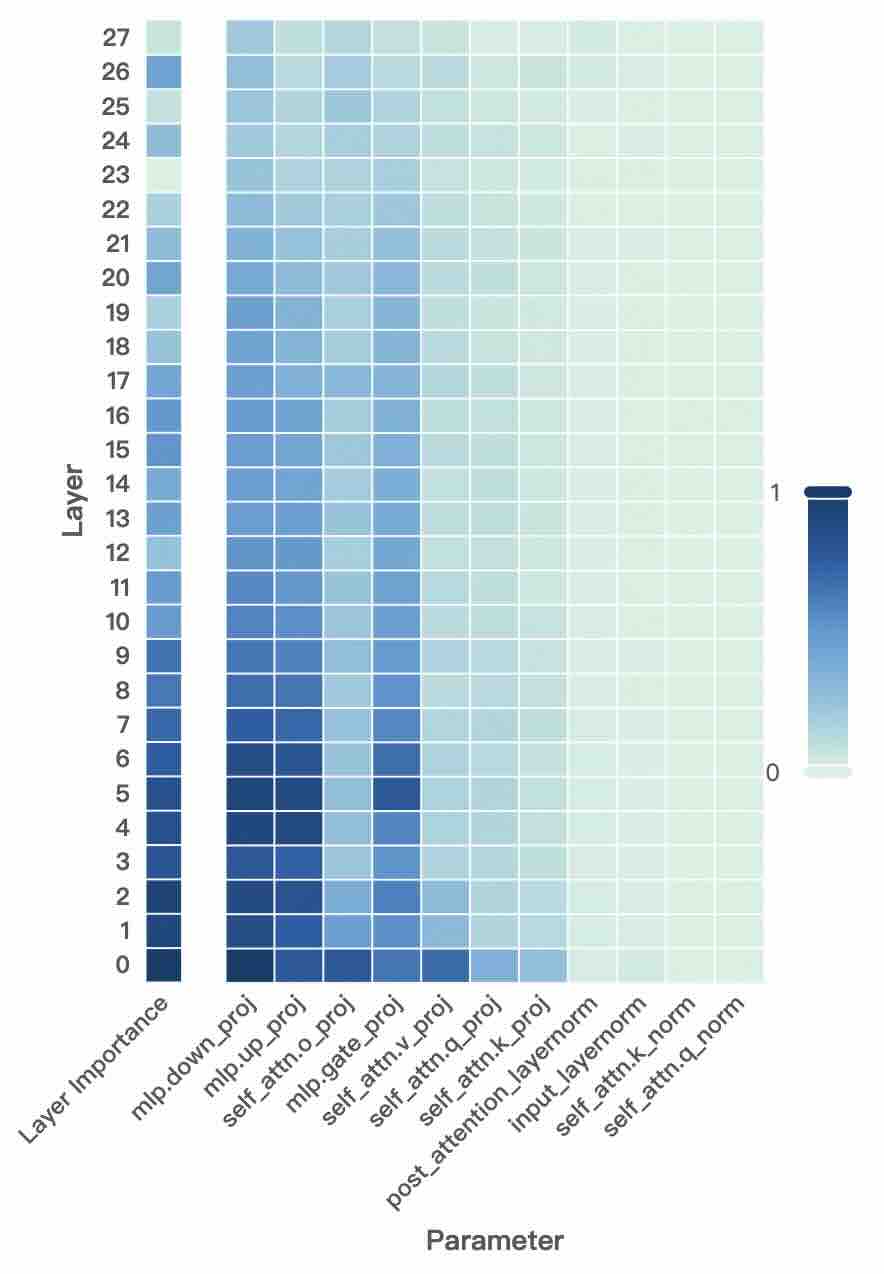
### Visual Description
## Heatmap: Layer Importance vs. Parameter
### Overview
The image is a heatmap visualizing the importance of different layers in a neural network with respect to various parameters. The heatmap uses a blue-to-green color gradient, where darker blue indicates higher importance and lighter green indicates lower importance. The y-axis represents the layer number (0-27), and the x-axis represents different parameters within the network.
### Components/Axes
* **Y-axis:** "Layer" with numerical labels from 0 to 27, incrementing by 1. Also includes the label "Layer Importance" at the bottom of the y-axis.
* **X-axis:** "Parameter" with the following labels:
* mlp.down\_proj
* mlp.up\_proj
* self\_attn.o\_proj
* mlp.gate\_proj
* self\_attn.v\_proj
* self\_attn.q\_proj
* self\_attn.k\_proj
* post\_attention\_layernorm
* input\_layernorm
* self\_attn.k\_norm
* self\_attn.q\_norm
* **Color Legend:** Located on the right side of the heatmap. Dark blue corresponds to a value of 1, and light green corresponds to a value of 0. The color gradient represents values between 0 and 1.
### Detailed Analysis or ### Content Details
The heatmap shows the relative importance of each layer (0-27) for each parameter.
* **mlp.down\_proj:** Layers 0-14 are dark blue, indicating high importance. Layers 15-27 transition to lighter shades of blue, indicating decreasing importance.
* **mlp.up\_proj:** Layers 0-14 are dark blue, indicating high importance. Layers 15-27 transition to lighter shades of blue, indicating decreasing importance.
* **self\_attn.o\_proj:** Layers 0-14 are dark blue, indicating high importance. Layers 15-27 transition to lighter shades of blue, indicating decreasing importance.
* **mlp.gate\_proj:** Layers 0-7 are dark blue, indicating high importance. Layers 8-14 transition to lighter shades of blue, indicating decreasing importance. Layers 15-27 transition to light green, indicating low importance.
* **self\_attn.v\_proj:** Layers 0-7 are dark blue, indicating high importance. Layers 8-14 transition to lighter shades of blue, indicating decreasing importance. Layers 15-27 transition to light green, indicating low importance.
* **self\_attn.q\_proj:** Layers 0-7 are dark blue, indicating high importance. Layers 8-14 transition to lighter shades of blue, indicating decreasing importance. Layers 15-27 transition to light green, indicating low importance.
* **self\_attn.k\_proj:** Layers 0-7 are dark blue, indicating high importance. Layers 8-14 transition to lighter shades of blue, indicating decreasing importance. Layers 15-27 transition to light green, indicating low importance.
* **post\_attention\_layernorm:** All layers (0-27) are light green, indicating low importance.
* **input\_layernorm:** All layers (0-27) are light green, indicating low importance.
* **self\_attn.k\_norm:** All layers (0-27) are light green, indicating low importance.
* **self\_attn.q\_norm:** All layers (0-27) are light green, indicating low importance.
### Key Observations
* The parameters `mlp.down_proj`, `mlp.up_proj`, and `self_attn.o_proj` show high importance for the lower layers (0-14) and decreasing importance for the higher layers (15-27).
* The parameters `mlp.gate_proj`, `self_attn.v_proj`, `self_attn.q_proj`, and `self_attn.k_proj` show high importance for the very lower layers (0-7), decreasing importance for the middle layers (8-14), and low importance for the higher layers (15-27).
* The parameters `post_attention_layernorm`, `input_layernorm`, `self_attn.k_norm`, and `self_attn.q_norm` consistently show low importance across all layers.
### Interpretation
The heatmap suggests that the lower layers (0-14) of the neural network are more critical for the `mlp.down_proj`, `mlp.up_proj`, and `self_attn.o_proj` parameters. The parameters `mlp.gate_proj`, `self_attn.v_proj`, `self_attn.q_proj`, and `self_attn.k_proj` are only important in the very lower layers (0-7). The parameters `post_attention_layernorm`, `input_layernorm`, `self_attn.k_norm`, and `self_attn.q_norm` may not significantly contribute to the network's performance, or their importance is distributed differently across the network architecture. This information can be used to optimize the network architecture, potentially by focusing on the lower layers for specific parameters or by simplifying the network by removing or reducing the influence of the less important parameters.
</details>
Figure 10: Layer-wise and parameter-wise importance distribution of Qwen3-1.7B-Base model
<details>
<summary>figures/importance_4B_new.jpg Details</summary>
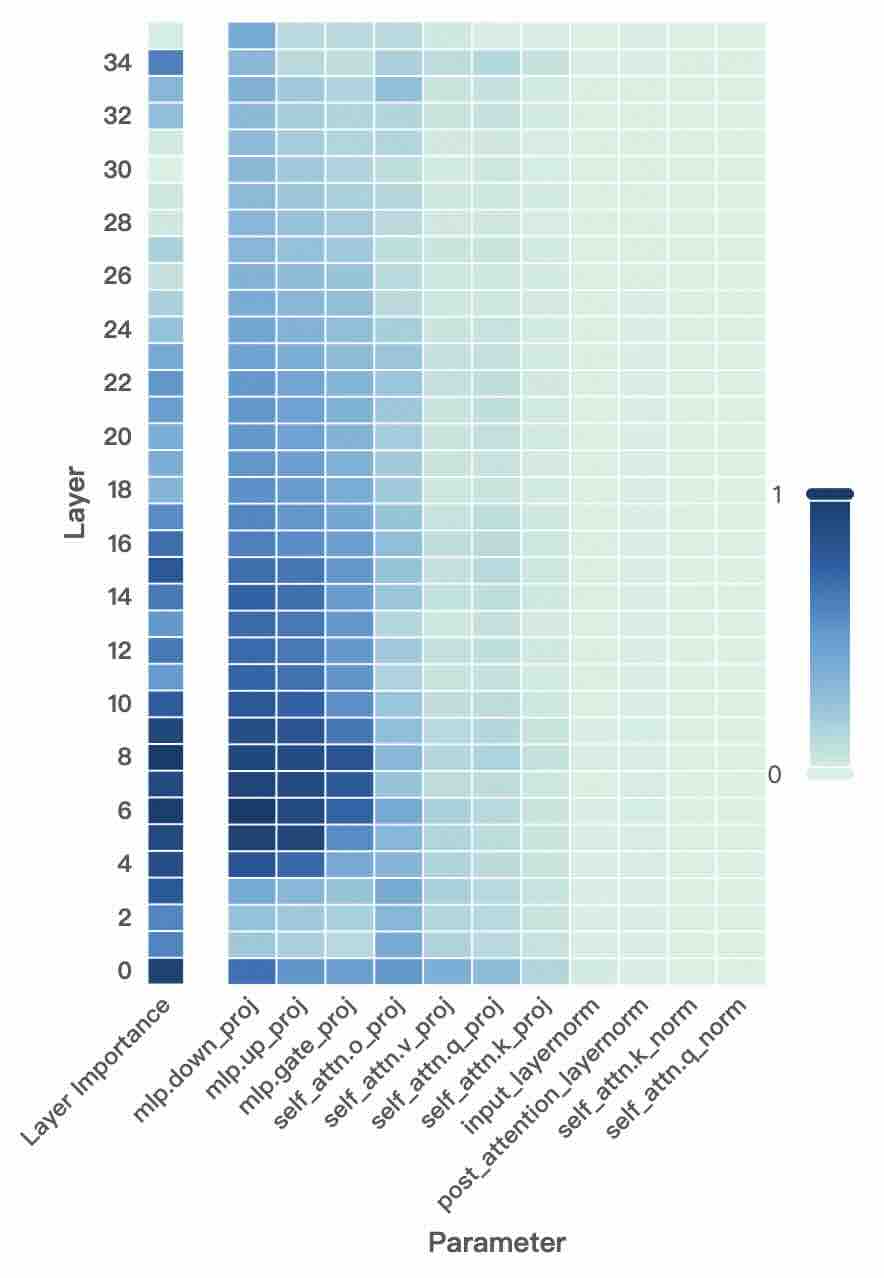
### Visual Description
## Heatmap: Layer Importance vs. Parameter
### Overview
The image is a heatmap visualizing the importance of different layers in a neural network with respect to various parameters. The color intensity represents the level of importance, ranging from dark blue (high importance, value of 1) to light green (low importance, value of 0). The y-axis represents the layer number, and the x-axis represents the different parameters.
### Components/Axes
* **X-axis (Parameter):**
* mlp.down\_proj
* mlp.up\_proj
* mlp.gate\_proj
* self\_attn.o\_proj
* self\_attn.v\_proj
* self\_attn.q\_proj
* self\_attn.k\_proj
* input\_layernorm
* post\_attention\_layernorm
* self\_attn.k\_norm
* self\_attn.q\_norm
* **Y-axis (Layer):** Numerical values from 0 to 34, incrementing by 2.
* **Legend:** A vertical color gradient from light green (0) to dark blue (1), indicating the level of importance. Located on the right side of the heatmap.
* **Title:** Layer Importance (y-axis label) vs Parameter (x-axis label)
### Detailed Analysis
The heatmap shows the importance of each layer (0-34) for each parameter. Darker blue indicates higher importance, while lighter green indicates lower importance.
* **mlp.down\_proj:** High importance (dark blue) from layer 0 to approximately layer 16, then gradually decreasing to light green by layer 34.
* **mlp.up\_proj:** High importance (dark blue) from layer 0 to approximately layer 16, then gradually decreasing to light green by layer 34.
* **mlp.gate\_proj:** High importance (dark blue) from layer 4 to approximately layer 8, then gradually decreasing to light green by layer 34.
* **self\_attn.o\_proj:** Starts with a medium blue around layer 0, decreasing to light green by layer 34.
* **self\_attn.v\_proj:** Starts with a medium blue around layer 0, decreasing to light green by layer 34.
* **self\_attn.q\_proj:** Starts with a light blue around layer 0, decreasing to light green by layer 34.
* **self\_attn.k\_proj:** Starts with a light blue around layer 0, decreasing to light green by layer 34.
* **input\_layernorm:** Light green across all layers, indicating low importance.
* **post\_attention\_layernorm:** Light green across all layers, indicating low importance.
* **self\_attn.k\_norm:** Light green across all layers, indicating low importance.
* **self\_attn.q\_norm:** Light green across all layers, indicating low importance.
### Key Observations
* The parameters `mlp.down_proj`, `mlp.up_proj`, and `mlp.gate_proj` show the highest importance in the lower layers (0-16).
* The parameters `input_layernorm`, `post_attention_layernorm`, `self_attn.k_norm`, and `self_attn.q_norm` have consistently low importance across all layers.
* The importance of most parameters decreases as the layer number increases.
### Interpretation
The heatmap suggests that the lower layers of the neural network are more critical for the `mlp.down_proj`, `mlp.up_proj`, and `mlp.gate_proj` parameters. The layernorm parameters (`input_layernorm`, `post_attention_layernorm`, `self_attn.k_norm`, and `self_attn.q_norm`) appear to have minimal impact on the network's performance across all layers, based on this importance metric. The decreasing importance of most parameters with increasing layer number could indicate that the initial layers are responsible for extracting fundamental features, while later layers refine these features with less reliance on the specified parameters.
</details>
Figure 11: Layer-wise and parameter-wise importance distribution of the Qwen3-4B-Base model
<details>
<summary>figures/importance_8B_new.jpg Details</summary>
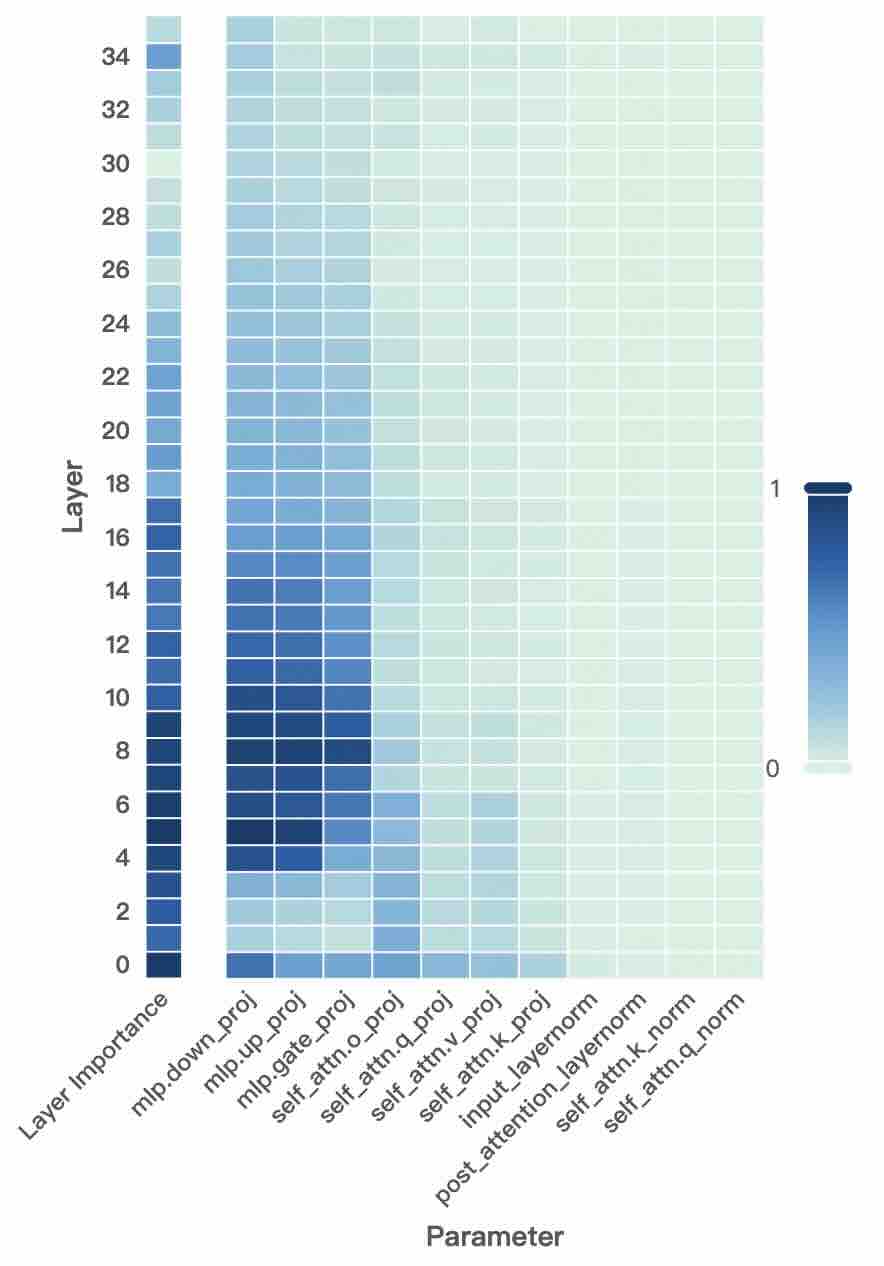
### Visual Description
## Heatmap: Layer Importance vs. Parameter
### Overview
The image is a heatmap visualizing the importance of different layers (y-axis) with respect to various parameters (x-axis). The color intensity represents the degree of importance, ranging from light green (low importance, near 0) to dark blue (high importance, near 1).
### Components/Axes
* **Y-axis:** "Layer" with numerical labels from 0 to 34 in increments of 2. Also includes "Layer Importance" label.
* **X-axis:** "Parameter" with the following labels:
* mlp.down\_proj
* mlp.up\_proj
* mlp.gate\_proj
* self\_attn.o\_proj
* self\_attn.q\_proj
* self\_attn.v\_proj
* self\_attn.k\_proj
* input\_layernorm
* post\_attention\_layernorm
* self\_attn.k\_norm
* self\_attn.q\_norm
* **Color Legend:** Located on the right side of the heatmap. Dark blue corresponds to a value of 1, and light green corresponds to a value of 0. The color gradient represents intermediate values.
### Detailed Analysis
The heatmap shows the relative importance of each parameter for each layer.
* **Layer Importance:** The leftmost column shows the importance of each layer overall. The lower layers (approximately 2 to 16) appear to have higher importance (darker blue) compared to the upper layers (lighter blue/green).
* **mlp.down\_proj, mlp.up\_proj, mlp.gate\_proj:** These parameters show high importance (dark blue) for layers approximately 4 to 12. The importance decreases as the layer number increases.
* **self\_attn.o\_proj, self\_attn.q\_proj, self\_attn.v\_proj, self\_attn.k\_proj:** These parameters show moderate importance (various shades of blue) for layers approximately 2 to 20, with some variation in intensity.
* **input\_layernorm, post\_attention\_layernorm, self\_attn.k\_norm, self\_attn.q\_norm:** These parameters generally show low importance (light green) across all layers.
### Key Observations
* Lower layers (4-12) are more important for the "mlp" parameters.
* The "layernorm" parameters have consistently low importance across all layers.
* The layer importance is concentrated in the lower layers.
### Interpretation
The heatmap suggests that the lower layers of the model are more critical for the "mlp" parameters, indicating that these layers might be responsible for initial feature extraction or processing. The "layernorm" parameters, on the other hand, seem to have a less significant role in the model's performance, as indicated by their low importance across all layers. The overall layer importance is concentrated in the lower layers, which could mean that these layers are crucial for the model's learning process.
</details>
Figure 12: Layer-wise and parameter-wise importance distribution of the Qwen3-8B-Base model
<details>
<summary>figures/importance_Llama_new.jpg Details</summary>
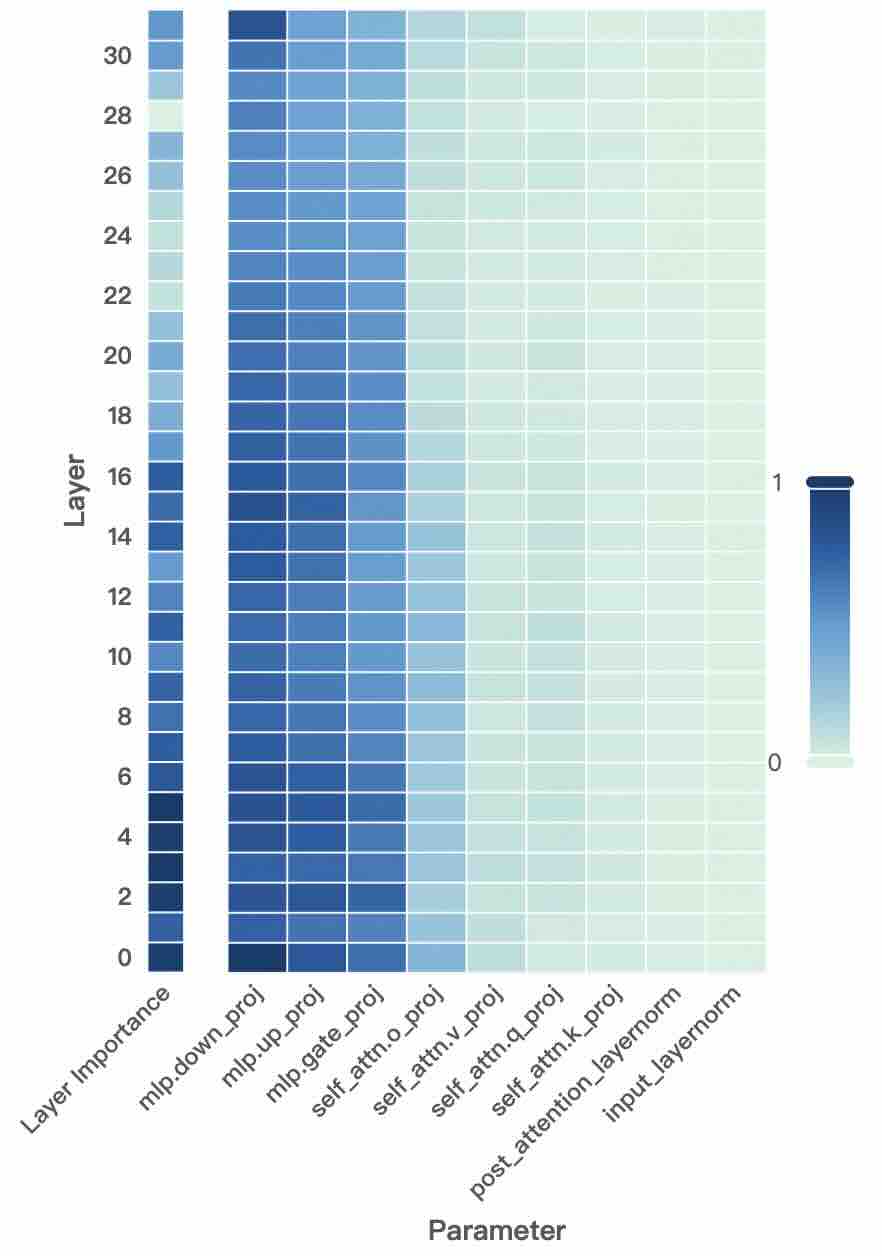
### Visual Description
## Heatmap: Layer Importance vs. Parameter
### Overview
The image is a heatmap visualizing the importance of different layers (y-axis) for various parameters (x-axis) in a neural network. The color intensity represents the degree of importance, with darker blue indicating higher importance and lighter shades indicating lower importance.
### Components/Axes
* **Y-axis:** "Layer" with numerical scale from 2 to 30 in increments of 2.
* **X-axis:** "Parameter" with the following categories:
* Layer Importance
* mlp.down\_proj
* mlp.up\_proj
* mlp.gate\_proj
* self\_attn.o\_proj
* self\_attn.v\_proj
* self\_attn.q\_proj
* self\_attn.k\_proj
* post\_attention\_layernorm
* input\_layernorm
* **Color Legend:** Located on the right side of the heatmap, ranging from dark blue (representing a value of 1) to light green/white (representing a value of 0).
### Detailed Analysis
The heatmap shows the importance of each layer for each parameter.
* **Layer Importance:** The "Layer Importance" parameter shows high importance (dark blue) across all layers, from layer 2 to layer 30.
* **mlp.down\_proj, mlp.up\_proj, mlp.gate\_proj:** These parameters also show high importance (dark blue) across all layers.
* **self\_attn.o\_proj:** This parameter shows high importance (dark blue) from layer 2 to approximately layer 16, then gradually decreases in importance (lighter shades of blue) towards layer 30.
* **self\_attn.v\_proj, self\_attn.q\_proj, self\_attn.k\_proj:** These parameters show a similar trend to "self\_attn.o\_proj," with high importance in lower layers and decreasing importance in higher layers, transitioning to light blue/green.
* **post\_attention\_layernorm, input\_layernorm:** These parameters show low importance (light green/white) across all layers.
### Key Observations
* The "Layer Importance," "mlp.down\_proj," "mlp.up\_proj," and "mlp.gate\_proj" parameters are consistently important across all layers.
* The importance of "self\_attn.o\_proj," "self\_attn.v\_proj," "self\_attn.q\_proj," and "self\_attn.k\_proj" parameters decreases as the layer number increases.
* The "post\_attention\_layernorm" and "input\_layernorm" parameters have low importance across all layers.
### Interpretation
The heatmap suggests that certain parameters (Layer Importance, mlp.down\_proj, mlp.up\_proj, mlp.gate\_proj) are crucial for all layers of the neural network. Self-attention related parameters (self\_attn.o\_proj, self\_attn.v\_proj, self\_attn.q\_proj, self\_attn.k\_proj) are more important in the lower layers, possibly indicating that these layers are responsible for capturing initial contextual information. The layernorm parameters (post\_attention\_layernorm, input\_layernorm) appear to have a less significant role in the network's performance, at least according to this importance metric. The data demonstrates a clear distinction in the importance of different parameters across the layers of the neural network.
</details>
Figure 13: Layer-wise and parameter-wise importance distribution of the Llama3-8B model
To investigate which layers should be expanded, we conduct a comprehensive importance analysis at both the layer and parameter levels. Specifically, we compute the importance scores for each layer and parameter across multiple models, and visualize their detailed distributions (see Figure 10, Figure 11, Figure 12, and Figure 13). Our analysis yields the following key observations:
1. Layer and parameter importance alignment. Overall, the distributions of layer-wise importance and parameter-wise importance are highly aligned across all four models. This alignment is expected, as both metrics are fundamentally computed under the same principle—estimating the impact of masking out (setting to zero) a given layer or parameter on model performance. Since parameter importance essentially decomposes the contribution at the layer level, this consistency reflects the intrinsic, nested relationship between the two. It also indicates that layer-level and parameter-level interventions affect the model’s predictive capability in a coherent manner.
2. High importance in lower layers and the penultimate layer exception. A notable pattern across all models is that the most important layers tend to be concentrated in the lower (early to middle) layers of the network, with importance values generally decreasing towards higher layers. This pattern suggests that the early layers play a critical role in the overall function of the model.
One plausible explanation, is that lower layers are responsible for capturing general syntactic properties and foundational compositionality (Clark et al., 2019; Hewitt & Manning, 2019), such as basic grammar and phrase structure. In contrast, deeper layers are typically responsible for integrating more task- or context-specific semantic information. This division of labor (earlier layers for generic linguistic structure, deeper layers for task semantics) naturally results in higher sensitivity to interventions at the bottom layers. This also provides a theoretical basis for layer expansion in deep layers.
An interesting exception observed in all models is that the penultimate layer does not follow this general trend: its importance appears elevated relative to immediately adjacent layers. This may stem from the model’s need to consolidate high-level semantic features just before producing the output prediction. The penultimate layer may act as a “bottleneck” for aggregating information necessary for the final decision or token generation—potentially as a final representation refinement step. Similar phenomena have been observed in works such as Intrinsic Dimensionality Explains the Effectiveness of Language Model Pruning (Aghajanyan et al., 2021), which highlight the special role of upper- and penultimate layers in output formation.
3. Intra- and inter-family patterns: Qwen vs. Llama models.
Qwen family: Across the Qwen models (Qwen3-1.7B, 4B, 8B), the overall trends are similar:
- Importance is strongly concentrated in the lower and middle layers, particularly within the first 10 layers, regardless of total model depth.
- Among parameters, mlp.down_proj and mlp.up_proj typically dominate in the most important layers, suggesting that feed-forward (MLP) components contribute substantially to the information processing in the Qwen series.
- With increasing model size (from 1.7B to 8B), the importance distribution appears to spread out slightly, showing less sharpness at the very bottom—possibly reflecting increased capacity and redundancy in larger networks.
Cross-family: Comparing Qwen models to Llama3-8B, we observe both notable similarities and differences:
- Both model families consistently exhibit high importance in MLP-related parameters (mlp.down_proj, mlp.up_proj, and mlp.gate_proj), especially in the most important layers. This underscores the universal role of the feed-forward network in transforming and integrating information beyond the capabilities of self-attention alone.
- Llama3-8B shows a broader distribution of importance across layers, with non-negligible values extending further into the middle and upper layers, suggesting a more distributed processing pipeline. In contrast, Qwen models tend to concentrate importance more in the lower layers.
- The dominance of MLP components in Llama3-8B is somewhat less pronounced than in Qwen, with parameter importance appearing more diffuse overall. These inter-family differences may be attributable to variations in architecture (such as normalization, attention mechanisms, or feed-forward design), pre-training data, or other modeling choices, leading to distinct strategies of information flow and representation across the network depth.
Appendix E Layer-wise Importance Estimation Methods Comparison
To investigate which layers contribute most to model performance, we employed four different strategies to compute layer-wise importance:
1. Cumulate importance of parameters: For each parameter $p$ in a layer, we compute the product $p\frac{∂\mathcal{L}}{∂ p}$ , and sum across all parameters in the layer:
$$
I_{\text{layer}}=\sum_{p\in\text{layer}}p\frac{\partial\mathcal{L}}{\partial p} \tag{11}
$$
1. Module-wise rank aggregation: For each module (e.g., attention, MLP, normalization), we calculate the importance score, rank layers by their score within each module, and aggregate rankings to obtain a total rank for each layer.
1. Masking out: For each layer, we mask out its parameters (i.e., set to zero) and evaluate the change in loss:
$$
I_{\text{layer}}=\mathcal{L}(\text{model with layer $l$ masked})-\mathcal{L}(\text{original model}) \tag{12}
$$
1. Fisher information: For each parameter $p$ in a layer, using the Fisher information approximation
$$
F(p)=\mathbb{E}\left[\left(\frac{\partial\log p(y|x)}{\partial p}\right)^{2}\right] \tag{13}
$$
Layer-level Fisher importance is obtained by summing over all parameters in the layer.
To further understand the significance and robustness of these metrics, we conducted a preliminary experiment on the Qwen3-1.7B-Base in the medical domain with dev subset of MMLU, CMMLU to detect the importance of layers, focusing on how different gradient computation strategies affect downstream performance.
Table 8: Performance of different expansion methods on medical-domain tasks (best result in each column is bolded). The numbers in parentheses after each method in the table indicate which layers were expanded. The Qwen3-1.7B-Base model has a total of 28 layers, indexed from 0 to 27.
| Methods Name | mmlu | cmmlu | medqa | cmb | mmcu |
| --- | --- | --- | --- | --- | --- |
| Qwen3-1.7B-Base | 62.57 | 66.86 | 48.39 | 63.67 | 69.17 |
| Uniformly Expansion (6,13,20,27) | 59.06 | 64.98 | 48.78 | 64.25 | 70.10 |
| Uniformly Expansion for first 16 layers (3,7,11,15) | 59.60 | 64.91 | 48.78 | 64.07 | 69.80 |
| Uniformly Expansion for last 16 layers (15,19,23,27) | 61.60 | 66.15 | 49.32 | 65.55 | 71.09 |
| Importance Cumulatation (23,24,25,27) | 62.63 | 66.81 | 50.19 | 63.85 | 69.48 |
| Rank Aggregation (22,24,25,27) | 62.72 | 66.86 | 50.57 | 63.97 | 69.78 |
| Masking Out (22,23,25,27) | 62.80 | 66.89 | 50.75 | 65.43 | 71.98 |
| Fisher (23,24,25,26) | 61.84 | 66.43 | 49.15 | 64.13 | 68.82 |
Table 8 compares the effect of different layer selection methods for expansion on a variety of medical-domain tasks using Qwen3-1.7B-Base. Several key observations can be made:
1. Similarity of selected layers across methods. All importance calculation methods lead to the selection of similar layers for expansion. For instance, the layers chosen by methods such as Importance Cumulatation (23,24,25,27), Rank Aggregation (22,24,25,27), Masking Out (22,23,25,27), and Fisher (23,24,25,26) significantly overlap, especially in the last 6 layers of the model (layers 22 and above). This convergence strongly validates our previous observations that general capability-critical layers tend to be concentrated in the latter half of the model in Appendix D.
In addition, the results show that uniform expansion into the last 16 layers (Uniformly Expansion for last 16 layers (15,19,23,27)) consistently outperforms expansion into the first 16 layers (Uniformly Expansion for first 16 layers (3,7,11,15)) or uniformly across all layers, further supporting the result in Appendix D.
2. Robustness of expansion results across methods. Despite minor variability in the specific layers chosen by each method, the final performance of all importance-based expansion approaches is consistently better than both the vanilla baseline and uniform expansion. For example, on the MedQA dataset, all methods using calculated importance exceed the baseline score (e.g., Masking Out achieves 50.75 vs. baseline 48.39), and on MMLU-med, Rank Aggregation achieves 67.95 versus the baseline 66.49. Crucially, the differences in scores among Masking Out, Rank Aggregation, Importance Cumulatation, and Fisher are relatively small for most tasks (typically less than 2 points), indicating that the overall framework is robust to the choice of importance calculation technique. Since our principal contribution is the training paradigm rather than the specific importance metric, for subsequent experiments, we employ the masking out approach, which demonstrated the strongest effect in preliminary experiment.
Appendix F Theoretical Analysis
Our theoretical analysis relies on several simplifying assumptions as outlined below. We discuss the rationality and limitations of each assumption:
1. Linearized Model Structure: We model the transformer as a stack of $L$ independent residual blocks, effectively ignoring cross-layer coupling effects such as those arising from pre-norm and residual connections.
Justification: In our layer expansion scheme, the newly added layers are always separated by at least one original frozen layer and never arranged in a cascading manner. This design substantially weakens direct coupling between newly expanded layers, which, in turn, reduces the degree of inter-layer interaction and nonlinearity affecting our analysis. And this abstraction is commonly used in theoretical studies (e.g., NTK analysis or pruning literature) to make layerwise analysis tractable.
1. Loss Function Smoothness: We assume the loss function $\ell(·,·)$ is $\beta$ -smooth and $L_{∞}$ -Lipschitz with respect to predictions.
Justification: Standard loss functions such as cross-entropy (with stability improvement) and mean squared error are widely established to satisfy these properties. These conditions allow us to relate small output perturbations to controlled changes in loss, facilitating theoretical bounds.
1. Training Dynamics: Our analysis assumes training is performed with a first-order SGD-like optimizer, disregarding effects from Adam or other adaptive methods.
Justification: First-order SGD provides well-understood theoretical properties and is commonly used in theoretical deep learning research. While Adam introduces adaptive scaling that can affect convergence, many results (e.g., generalization gap bounds) transfer qualitatively between SGD and Adam in practice.
1. NTK Regime and Sensitivity: Our analysis of layer sensitivity relies on the NTK (Neural Tangent Kernel) approximation (Jacot et al., 2018), which essentially assumes the model behaves locally linearly around its current parameters. Moreover, we should consider the model training process to be relatively stable, with no anomalous occurrences such as gradient explosion.
Justification: This assumption is particularly well-motivated in our setting for two reasons. First, our adaptation protocol only updates a small number of newly introduced parameters while keeping the vast majority of the pre-trained weights frozen and decouples parameters to maximize the retention of general capabilities. This ensures that the parameter changes remain minimal, keeping the network within the local linear (NTK) regime throughout adaptation. Second, unlike random initialization, our starting point is a well-trained model on a large general-domain corpus, which already provides robust and meaningful representations. Perturbations induced by finetuning are thus intrinsically local in the function space and less likely to induce sudden or nonlinear model behavior, further enhancing the validity of the NTK approximation.
Overall, these assumptions enable us to derive interpretable upper bounds and provide actionable layer selection criteria, but should be considered as idealizations. The correspondence between these theoretical insights and practical behavior is also validated in our empirical experiments.
F.1 Optimality of Least-Important Block Expansion for Preserving General Capabilities
Notation: Let $M_{0}$ denote the original base model, and $M_{S}^{(T)}$ denote the model after $T$ steps of adaptation, wherein only the set $S$ of $k$ layers are unfrozen and updated, and $\ell(·,y)$ is the loss function (e.g., cross-entropy) which is $L$ -Lipschitz and $\beta$ -smooth in its first argument. $\Delta^{(l)}$ represents the importance score of layer $l$ as defined below.
Layer Importance Score:
$$
\Delta^{(l)}:=\mathbb{E}_{x\sim D_{gen}}\big[\ell(M_{0}^{(-l)}(x),y(x))-\ell(M_{0}(x),y(x))\big]
$$
where $M_{0}^{(-l)}$ is $M_{0}$ with the $l$ -th layer masked out.
**Theorem F.1 (Upper Bound on Generalization Gap by Layer Importance)**
*Let $S⊂eq[L]$ be the set of layers selected for expansion/adaptation, and $G(S)$ denote the source-domain generalization gap after adaptation, i.e.,
$$
G(S):=\mathbb{E}_{x\sim D_{gen}}\left[\ell(M_{S}^{(T)}(x),y(x))-\ell(M_{0}(x),y(x))\right].
$$
Under function-preserving initialization, limited adaptation steps, and $L$ -Lipschitz and $\beta$ -smooth loss, the following upper bound holds:
$$
G(S)\leq C\sum_{l\in S}\Delta^{(l)}+O\left(k(\overline{\Delta W})^{2}\right)
$$
where $C$ is a constant depending on the learning rate, steps, loss smoothness, and initialization, and $\overline{\Delta W}$ is the maximal per-layer parameter change over adaptation.*
* Proof*
Step 1: Output Deviation Linearization. By function-preserving initialization, $M_{S}^{(0)}(x)=M_{0}(x)$ . After adaptation, since only layers in $S$ are modified and changes are small (Assumption A4), the output difference admits a first-order Taylor expansion:
$$
M_{S}^{(T)}(x)-M_{0}(x)\approx\sum_{l\in S}J_{l}(x)\ \Delta W_{l}
$$
where $J_{l}(x)=\left.\frac{∂ M}{∂ W_{l}}\right|_{W=W_{0}}$ and $\Delta W_{l}=W_{l}^{(T)}-W_{l}^{(0)}$ . Step 2: Lipschitz Property Application. By $L$ -Lipschitzness of $\ell(·,y)$ in its first argument,
$$
|\ell(M_{S}^{(T)}(x),y)-\ell(M_{0}(x),y)|\leq L\left\|M_{S}^{(T)}(x)-M_{0}(x)\right\|_{2}.
$$
Taking the expectation over $x\sim D_{gen}$ ,
$$
G(S)\leq L\;\mathbb{E}_{x}\left[\|M_{S}^{(T)}(x)-M_{0}(x)\|_{2}\right].
$$ Step 3: Breaking by Layer via Triangle Inequality. According to Assumption A1 and using the triangle inequality,
$$
\|M_{S}^{(T)}(x)-M_{0}(x)\|_{2}\leq\sum_{l\in S}\|J_{l}(x)\,\Delta W_{l}\|_{2},
$$
thus,
$$
G(S)\leq L\sum_{l\in S}\mathbb{E}_{x}\big[\|J_{l}(x)\,\Delta W_{l}\|_{2}\big].
$$ Step 4: Relating to Layer Importance Score. Recall the definition:
$$
\Delta^{(l)}=\mathbb{E}_{x}\left[\ell(M_{0}^{(-l)}(x),y)-\ell(M_{0}(x),y)\right].
$$
By Taylor expansion and Lipschitz continuity,
$$
|\ell(M_{0}^{(-l)}(x),y)-\ell(M_{0}(x),y)|\approx L\|J_{l}(x)W_{l}^{(0)}\|_{2}, \tag{0}
$$
so for small modifications,
$$
\mathbb{E}_{x}[\|J_{l}(x)\,\Delta W_{l}\|_{2}]\leq\frac{\|\Delta W_{l}\|_{2}}{\|W_{l}^{(0)}\|_{2}}\Delta^{(l)}+O(\|\Delta W_{l}\|_{2}^{2}). \tag{0}
$$
Assume $\|\Delta W_{l}\|_{2}≤\overline{\Delta W}$ for all $l∈ S$ and $\|W_{l}^{(0)}\|_{2}$ are similar or lower-bounded by $w_{0}>0$ , so
$$
G(S)\leq L\frac{\overline{\Delta W}}{w_{0}}\sum_{l\in S}\Delta^{(l)}+O\big(k(\overline{\Delta W})^{2}\big).
$$ Step 5: Optimization Control. In standard SGD (Assumption A3), $\overline{\Delta W}$ is controlled by learning rate $\eta$ , steps $T$ , batch size $N$ , and bounded gradients:
$$
\overline{\Delta W}\lesssim\frac{\eta T}{N}\ \max_{t,i}\|\nabla_{W_{l}}\ell(M_{0}(x_{i}),y_{i})\|_{2}.
$$
Thus, all learning and initialization constants can be absorbed into a scalar constant $C$ (Assumption A3 and A4). Step 6: Conclusion. Thus,
$$
G(S)\leq C\sum_{l\in S}\Delta^{(l)}+O\left(k(\overline{\Delta W})^{2}\right).
$$
which completes the proof. ∎
Due to the use of residual connections, the original block and the expanded block can be viewed as a single aggregated unit. Importantly, before training, the addition of the new block does not alter the model’s output, and thus the overall importance of the aggregated block remains exactly the same as that of the original block (i.e., $\Delta^{(l)}$ ). As a result, when we train the parameters of the new block, it is effectively equivalent to adapting the aggregated block as a whole, whose importance is still characterized by the original importance score $\Delta^{(l)}$ . This justifies why the potential impact of training the expanded layer is governed by the original layer’s importance.
The tightness of the derived upper bound hinges on both the local linearity of the expansion regime and the control over parameter updates during adaptation. In cases where the expansion layers are initialized to be function-preserving and the adaptation is performed with sufficiently small learning rates and moderate step sizes, the Taylor and Lipschitz approximations used in the proof become increasingly sharp. Thus, the upper bound is not only theoretically attainable, but also approaches the realistic generalization gap observed in practice under these conditions. This means that minimizing the sum $\sum_{l∈ S}\Delta^{(l)}$ when selecting layers for expansion is not merely a mathematical convenience—it is a principled, actionable strategy for controlling catastrophic forgetting and generalization degradation. As a consequence, our criterion provides practical guidance: by limiting updates to those layers with the lowest importance scores, practitioners can reliably minimize negative transfer from domain adaptation, especially when adapting large pre-trained models with limited new capacity.
F.2 Optimality of Importance-Based Learning Rate Adjustment for Modules
We provide a rigorous analysis of learning rate reallocation in Stage 2. Specifically, let the importance of each parameter $\theta_{j}$ in the general domain be defined as
$$
I_{\theta_{j}}=\left|\frac{\partial L_{\mathrm{gen}}}{\partial\theta_{j}}\right|
$$
where $L_{\mathrm{gen}}$ denotes the general-domain loss and $I_{\theta_{j}}$ quantifies the sensitivity of the overall performance with respect to $\theta_{j}$ . Under the constraint of a fixed average learning rate, our strategy assigns lower learning rates to parameters with high general-domain importance, and higher learning rates to those deemed less important. This importance-weighted reallocation is provably optimal for minimizing the upper bound of catastrophic forgetting in the general domain, subject to the constant average learning rate constraint. Furthermore, we formulate and analytically solve the underlying constrained optimization problem to ensure that our reallocation approach achieves relative optimality in practice.
Setup and Notation
Let $D_{gen}$ be the general domain distribution with loss $L_{gen}(\theta)$ . With $\theta^{*}$ as the original pre-trained parameters, we define parameter importance $I_{j}\triangleq\theta_{j}\frac{∂ L_{gen}}{∂\theta_{j}}|_{\theta^{*}}$ and unit importance:
$$
I_{U_{i}}\triangleq\frac{1}{|U_{i}|}\sum_{j\in U_{i}}I_{j}\in[0,1] \tag{14}
$$
under learning rate budget constraint:
$$
\sum_{i}\frac{|U_{i}|}{|\Theta_{\sim}|}lr_{U_{i}}=lr_{base} \tag{15}
$$
F.2.1 Upper Bound on Forgetting
Define forgetting as:
$$
F\triangleq L_{gen}(\theta(T))-L_{gen}(\theta^{*}) \tag{16}
$$
Assuming $L_{gen}$ is $\beta$ -smooth, the first-order Taylor expansion provides:
$$
F\leq\nabla_{\theta}L_{gen}(\theta^{*})^{\top}\Delta(T)+\frac{\beta}{2}\|\Delta(T)\|^{2} \tag{17}
$$
Due to parameter freezing, the gradient $∇_{\theta}L_{gen}(\theta^{*})$ is only non-zero for expanded parameters:
$$
\nabla_{\theta}L_{gen}(\theta^{*})=\sum_{i}\sum_{j\in U_{i}}I_{j}e_{j} \tag{18}
$$
where $I_{j}=\frac{∂ L_{gen}}{∂\theta_{j}}$ , $e_{j}$ are basis vectors.
Assuming gradient descent with per-group step size $\eta_{U_{i}}$ and $T$ steps, for each parameter $j∈ U_{i}$ (Assumption A4):
$$
\Delta_{j}(T)\approx-T\eta_{U_{i}}\frac{\partial L_{med}}{\partial\theta_{j}} \tag{19}
$$
Substitute into the smoothness bound:
$$
\displaystyle F \displaystyle\leq\sum_{i}\sum_{j\in U_{i}}I_{j}\Delta_{j}(T)+\frac{\beta}{2}\sum_{i}\sum_{j\in U_{i}}(\Delta_{j}(T))^{2} \displaystyle\leq\sum_{i}|U_{i}|\cdot|I_{U_{i}}|\cdot(T\eta_{U_{i}}G)+\frac{\beta}{2}T^{2}\sum_{i}|U_{i}|\eta_{U_{i}}^{2}G^{2} \tag{20}
$$
where $G:=\max_{j}|\frac{∂ L_{med}}{∂\theta_{j}}|$ upper-bounds the adaptation gradients.
The derived upper bound encompasses all possible learning rate allocations and ensures conservative control over catastrophic forgetting. Note that if group gradients $G$ or importance scores $I_{U_{i}}$ are heterogeneous, a more refined bound can be obtained by analyzing variance rather than worst-case values.
F.3 Optimal Importance-Driven Learning Rate Reallocation
Problem Statement: We aim to allocate learning rates $\eta_{U_{i}}$ for each parameter group $U_{i}$ so as to minimize the upper bound on forgetting:
$$
F\leq a\sum_{i}w_{i}I_{i}\eta_{U_{i}}+b\sum_{i}w_{i}\eta_{U_{i}}^{2}
$$
where $w_{i}=|U_{i}|$ is the number of parameters in group $U_{i}$ , $I_{i}=|I_{U_{i}}|$ indicates the average importance of parameters in $U_{i}$ , $a,b>0$ are constants determined by training steps, gradient norms, and the smoothness constant ( $\beta$ (Assumption A2)). The constraint is that the average learning rate remains fixed:
$$
\sum_{i}w_{i}\eta_{U_{i}}=W\eta_{avg}
$$
where $W=\sum_{i}w_{i}$ is the total number of trainable parameters.
Lagrangian Formulation: Introduce a Lagrange multiplier $\lambda$ and write the Lagrangian:
$$
\mathcal{L}(\{\eta_{U_{i}}\},\lambda)=a\sum_{i}w_{i}I_{i}\eta_{U_{i}}+b\sum_{i}w_{i}\eta_{U_{i}}^{2}+\lambda\left(\sum_{i}w_{i}\eta_{U_{i}}-W\eta_{avg}\right)
$$
Optimality Condition: Taking derivatives and setting to zero, we obtain for each $j$ :
$$
\frac{\partial\mathcal{L}}{\partial\eta_{U_{j}}}=aw_{j}I_{j}+2bw_{j}\eta_{U_{j}}+\lambda w_{j}=0
$$
$$
\Longrightarrow\eta_{U_{j}}^{*}=-\frac{a}{2b}I_{j}-\frac{\lambda}{2b}
$$
Including the constraint:
$$
\sum_{j}w_{j}\eta_{U_{j}}^{*}=W\eta_{avg}
$$
Plugging in the expression for $\eta_{U_{j}}^{*}$ gives:
$$
-\frac{a}{2b}\sum_{j}w_{j}I_{j}-\frac{\lambda}{2b}W=W\eta_{avg}
$$
Solving for $\lambda$ :
$$
\lambda=-2b\eta_{avg}-\frac{a}{W}\sum_{j}w_{j}I_{j}
$$
So the optimal learning rate for group $U_{j}$ is:
$$
\eta_{U_{j}}^{*}=\eta_{avg}-\frac{a}{2b}\left(I_{j}-\frac{1}{W}\sum_{j^{\prime}}w_{j^{\prime}}I_{j^{\prime}}\right) \tag{22}
$$
Interpretation and Guidance: When the theoretical upper bound is tight—which is often the case in well-controlled, locally linear training regimes—this result has direct practical utility. Notably, the optimal learning rate allocation $\eta_{U_{j}}^{*}$ is an affine (linear) function of the group importance $I_{j}$ . Our method, which assigns $\text{lr}_{U}=2·(1-I_{\text{unit}})·\text{lr}_{\text{base}}$ , can be viewed as a simplified implementation of the derived optimal form. By decreasing the learning rate for groups with high general-domain importance and increasing it for those with low importance, this strategy effectively minimizes the risk of catastrophic forgetting while respecting the global learning rate constraint. Thus, our approach provides actionable guidance for tailoring learning rates based on parameter importance in continual learning and domain adaptation.
Appendix G Experiment about the number of expanded layers
In Stage 1, determining the optimal number of expanded layers emerges as a crucial hyperparameter. To investigate this, we conducted systematic experiments across various model scales in the medical domain by expanding different numbers of layers. These comprehensive experiments aim to provide empirical insights into selecting the most effective layer expansion strategy, offering valuable guidance for future research in this direction.
Table 9: Comparative Performance of Different Layer Expansion Strategies across Model Scales and Medical Tasks. Bold indicates the best-performing setup for each task; underline shows the second-best. This highlights optimal and near-optimal choices for each scenario.
| Model | MMLU | CMMLU | MedQA | MMCU-Medical | CMB |
| --- | --- | --- | --- | --- | --- |
| Qwen3-1.7B | | | | | |
| Vanilla | 62.57 | 66.86 | 48.39 | 69.17 | 63.67 |
| 1-layer | 62.31 | 66.23 | 48.08 | 69.95 | 61.40 |
| 2-layer | 62.48 | 66.91 | 48.63 | 70.78 | 62.89 |
| 4-layer | 62.80 | 66.89 | 50.75 | 71.98 | 65.43 |
| 8-layer | 61.84 | 66.02 | 49.57 | 72.41 | 65.00 |
| 16-layer | 60.96 | 64.65 | 48.86 | 70.13 | 64.88 |
| Qwen3-4B | | | | | |
| Vanilla | 73.19 | 77.92 | 62.77 | 82.44 | 78.92 |
| 1-layer | 72.98 | 77.69 | 63.39 | 82.83 | 78.21 |
| 2-layer | 73.10 | 77.84 | 63.08 | 82.80 | 78.48 |
| 4-layer | 72.95 | 78.77 | 64.49 | 84.58 | 79.87 |
| 8-layer | 73.06 | 77.65 | 65.02 | 84.22 | 78.81 |
| 16-layer | 72.06 | 77.11 | 62.61 | 82.09 | 78.61 |
| Qwen3-8B | | | | | |
| Vanilla | 76.94 | 82.09 | 66.30 | 86.45 | 81.67 |
| 1-layer | 76.84 | 82.06 | 67.87 | 86.95 | 81.50 |
| 2-layer | 76.70 | 82.10 | 67.93 | 87.99 | 82.90 |
| 4-layer | 76.77 | 82.11 | 69.24 | 89.84 | 85.80 |
| 8-layer | 76.77 | 82.15 | 68.34 | 88.02 | 84.85 |
| 16-layer | 77.12 | 82.28 | 68.56 | 87.76 | 84.32 |
| LLaMA3-8B | | | | | |
| Vanilla | 65.33 | 50.83 | 58.91 | 46.29 | 35.61 |
| 1-layer | 65.29 | 51.12 | 58.97 | 50.83 | 40.45 |
| 2-layer | 65.61 | 50.98 | 59.56 | 55.92 | 47.83 |
| 4-layer | 65.25 | 51.73 | 60.82 | 63.17 | 54.65 |
| 8-layer | 65.17 | 51.92 | 61.17 | 67.03 | 61.78 |
| 16-layer | 65.12 | 52.45 | 61.92 | 70.86 | 65.31 |
For general language tasks such as MMLU and CMMLU, all models largely preserve their baseline performance regardless of the number of expanded layers. This indicates that layer expansion does not compromise the models’ general language capabilities and robustness.
However, for domain-specific medical tasks (MedQA, MMCU-Medical, and CMB), the impact of layer expansion is more pronounced. Across all Qwen model variants (1.7B, 4B, and 8B), expanding 4 layers consistently yields optimal performance, as shown by the bolded results in Table 9. Specifically, the Qwen3-1.7B, 4B, and 8B models improve on MMCU-Medical by up to 2.8%, 2.1%, and 3.4%, respectively, when increasing from baseline to 4-layer expansion. Notably, expanding beyond 4 layers (e.g., to 8 or 16 layers) does not systematically improve performance—and in several cases, results in diminishing or even degraded accuracy. This suggests that moderate layer expansion (4 layers) achieves a balance between performance gain and model stability, while excessive expansion may introduce optimization difficulties, overfitting, or disrupt the pre-trained knowledge representations, leading to suboptimal outcomes.
In contrast, the LLaMA3-8B model displays a unique trend: performance improvements are continuous as more layers are expanded, with the best results observed at expanding 16 layers. The gains are considerable for tasks like MMCU-Medical and CMB, where scores rise dramatically from 46.29% and 35.61% in the vanilla model to 70.86% and 65.31% with 16 expanded layers. This behavior contrasts with the Qwen models and is likely due to LLaMA’s more limited Chinese capability in its original configuration. The need for extensive architectural expansion reflects the necessity to build new, specialized representations to compensate for baseline deficiencies when addressing Chinese-centric tasks. Therefore, while moderate layer expansion is optimal for models pre-trained on Chinese data (Qwen), more substantial expansion may be required for models less adapted to the target language or domain (LLaMA).
Overall, these results indicate that expanding more layers does not guarantee better performance. For well-aligned models, excessive expansion may lead to interference with the original knowledge or cause optimization instability. In contrast, for models lacking target domain competence, increased expansion helps establish the missing representations, albeit at the cost of greater computational complexity.
Appendix H Take Pretrain data as Importance Source
Our previous experiments employed the dev sets of MMLU and CMMLU as benchmark datasets for gradient-based importance estimation. However, such high-quality and carefully curated benchmarks are often scarce, especially in practical industrial scenarios. To investigate the robustness of our ADEPT method under more realistic conditions where benchmark data may not be available, we explore the use of noisier pretraining data for importance estimation.
Table 10: General Competence Detection Pretrain Corpus. #Examples means the number of examples we used.
| Dataset | #Examples | Hugging Face Link |
| --- | --- | --- |
| FineWeb_Edu | 500 | HuggingFaceFW/fineweb-edu |
| FineWeb_Edu_Chinese V2.1 | 500 | HuggingFaceFW/fineweb-edu |
Specifically, we utilize the FineWebEdu and FineWebEdu-Chinese datasets (Data overview and links in Table 10), extracting the top 500 samples with the highest educational scores from the first 10,000 entries in each corpus to serve as our importance estimation set. Compared to curated benchmarks, these datasets are much more accessible in real-world applications. Furthermore, the computational cost for filtering out such high-quality samples is negligible relative to the overall cost of large-scale pretraining.
This experimental setting allows us to rigorously evaluate the robustness of ADEPT when real-world, easily accessible pretraining data replaces ideal benchmark datasets for importance-based layer expansion decisions.
Table 11: Performance comparison of ADEPT with benchmark-based and pretraining-data-based importance estimation across model scales. Bold indicates the best performance per column; underline marks the second-best.
| Model | MMLU | CMMLU | MedQA | MMCU-Medical | CMB |
| --- | --- | --- | --- | --- | --- |
| Qwen3-1.7B | | | | | |
| Vanilla | 62.57 | 66.86 | 48.39 | 69.17 | 63.67 |
| ADEPT (Benchmark) | 62.80 | 66.89 | 50.75 | 71.98 | 65.43 |
| ADEPT (PT Data) | 62.85 | 66.87 | 49.39 | 70.84 | 63.07 |
| Qwen3-4B | | | | | |
| Vanilla | 73.19 | 77.92 | 62.77 | 82.44 | 78.92 |
| ADEPT (Benchmark) | 72.95 | 78.77 | 64.49 | 84.58 | 79.87 |
| ADEPT (PT Data) | 73.14 | 77.96 | 63.94 | 83.34 | 79.62 |
| Qwen3-8B | | | | | |
| Vanilla | 76.94 | 82.09 | 66.30 | 86.45 | 81.67 |
| ADEPT (Benchmark) | 76.77 | 82.11 | 69.24 | 89.84 | 85.80 |
| ADEPT (PT Data) | 76.83 | 82.20 | 67.56 | 87.20 | 83.92 |
| LLaMA3-8B | | | | | |
| Vanilla | 65.33 | 50.83 | 58.91 | 46.29 | 35.61 |
| ADEPT (Benchmark) | 65.25 | 51.73 | 60.82 | 63.17 | 54.65 |
| ADEPT (PT Data) | 65.21 | 50.27 | 59.13 | 60.29 | 51.32 |
Table 11 summarizes the performance of our ADEPT method when the importance estimation is conducted with either high-quality benchmark data or more easily accessible pretraining data across different model scales. Overall, the results demonstrate that ADEPT not only consistently outperforms the vanilla baseline but also shows remarkable robustness across most scenarios when using pretraining data for importance calculation. In Qwen3 series models, the difference between benchmark-based and pretraining-data-based importance estimation is minimal. In several cases, the latter even slightly surpasses the benchmark version (e.g., Qwen3-1.7B on MMLU and Qwen3-8B on MMLU and CMMLU), validating the practical applicability and flexibility of our approach.
For LLaMA3-8B, ADEPT with pretraining data still yields clear improvements over the vanilla baseline on all tasks, particularly in domain-specific metrics such as MedQA and MMCU-Medical. However, compared to the benchmark-based ADEPT, the pretraining-data variant shows slightly lower performance, with a gap of approximately 1–5% across tasks. This modest drop can be attributed to two main factors: first, the inherent discrepancy between noisier pretraining data and expertly curated benchmarks introduces less precise gradient signals for importance estimation. Second, LLaMA3-8B’s weaker baseline in Chinese tasks means its optimization is more sensitive to the quality of importance source, and benefits more from highly targeted benchmark data. Nonetheless, even with this gap, the pretraining-data approach remains highly valid, especially in practical scenarios where access to dedicated benchmarks is limited.
In summary, ADEPT demonstrates strong effectiveness and robustness when layer expansion is guided by pretraining data, making it highly suitable for real-world deployment. The slight performance drop observed in LLaMA3-8B highlights the additional value of benchmark data for models or tasks with substantial baseline limitations, but does not diminish the overall utility of our method in resource-constrained settings.
Appendix I Token Distribution Shift
Following the methodology proposed by Lin et al. (2024), we conducted a comprehensive analysis of token distribution shifts between the base and aligned models using the MMLU (Massive Multitask Language Understanding) dataset. The analysis focuses on identifying and quantifying the changes in token prediction patterns that occur during the alignment process.
Our analysis procedure consists of the following steps:
1) For each position in the input text, we use the aligned model with greedy decoding to generate the output token $o_{t}$ .
2) We then examine how this token is ranked in the base model’s probability distribution $P_{base}$ . This ranking, denoted as $\eta$ , serves as our primary metric for categorizing token shifts.
3) Based on the base ranking $\eta$ , we classify each token position into three categories:
- Unshifted positions ( $\eta=1$ ): The token is top-ranked in both base and aligned models
- Marginal positions ( $1<\eta≤ 3$ ): The token has a relatively high probability in the base model
- Shifted positions ( $\eta>3$ ): The token is unlikely to be sampled by the base model
4) For shifted tokens, we calculate Rank Improvement Ratio: $\frac{\text{base\_rank}}{\text{aligned\_rank}}$
Our analysis of the MMLU dataset revealed significant distribution shifts between the base and continual pretrained models by ADEPT. Figure 14 visualizes the most significantly shifted tokens, where the size of each token is proportional to its rank improvement ratio.
<details>
<summary>figures/wordcloud_med.jpg Details</summary>

### Visual Description
## Word Cloud: Medical Research Terms
### Overview
The image is a word cloud, visually representing the frequency of different terms related to medical research. The size of each word corresponds to its frequency or importance within the context from which the words were extracted. The words are arranged randomly and colored in shades of green and blue.
### Components/Axes
* **Words:** The primary components are the individual words themselves, such as "Health," "Patient," "Drug," "Treatment," "Disease," etc.
* **Size:** The size of each word indicates its relative frequency or importance. Larger words appear more frequently.
* **Color:** The words are colored in shades of green and blue.
### Detailed Analysis or ### Content Details
Here's a breakdown of the words and their relative prominence based on size:
* **Largest Words:**
* "Health" (Green): One of the most prominent words, suggesting a strong focus on general health-related topics.
* "Patient" (Green): Indicates a significant emphasis on patient-related research.
* "Treatment" (Blue): Highlights the importance of treatment methodologies.
* "Drug" (Blue): Suggests a focus on pharmaceutical interventions.
* **Large Words:**
* "Care" (Blue): Indicates a focus on healthcare services.
* "Hospital" (Green): Suggests research related to hospital settings.
* "Disease" (Green): Indicates a focus on specific diseases.
* "Medication" (Green): Highlights the importance of pharmaceutical interventions.
* "Heart" (Blue): Suggests a focus on cardiovascular health.
* "Diabetes" (Blue): Indicates a focus on diabetes research.
* "Symptoms" (Blue): Highlights the importance of symptom identification.
* "Therapy" (Blue): Suggests a focus on therapeutic interventions.
* "Brain" (Blue): Indicates a focus on neurological research.
* "Tumor" (Green): Highlights the importance of cancer research.
* "Prescription" (Blue): Suggests a focus on pharmaceutical interventions.
* "Randomized" (Blue): Indicates a focus on randomized controlled trials.
* **Medium-Sized Words:**
* "Dental" (Green): Indicates a focus on dental health.
* "Medical" (Green): Suggests research related to medical settings.
* "RNA" (Green): Highlights the importance of RNA research.
* "Medicine" (Yellow): Indicates a focus on medical settings.
* "Results" (Green): Highlights the importance of research outcomes.
* "MRI" (Green): Suggests research related to medical imaging.
* **Smaller Words:**
* "FDA" (Blue): Indicates a focus on regulatory compliance.
* "GDPR" (Green): Indicates a focus on data privacy.
* "Research" (Green): Highlights the importance of research activities.
* "Arthritis" (Green): Suggests research related to arthritis.
* "Concentrations" (Green): Indicates a focus on chemical concentrations.
* "Cases" (Blue): Indicates a focus on case studies.
* "Illness" (Green): Suggests research related to illness.
* "Protein" (Green): Highlights the importance of protein research.
* "LOS" (Green): Length of Stay
* "Best" (Green): Indicates a focus on best practices.
* "MD" (Blue): Indicates a focus on medical doctors.
* "DICE" (Blue): Indicates a focus on dice research.
* "Cancer" (Green): Suggests research related to cancer.
* "Healthy" (Green): Indicates a focus on healthy lifestyles.
* "Investigator" (Green): Highlights the importance of research investigators.
* "CDC" (Green): Indicates a focus on the Centers for Disease Control and Prevention.
* "Warning" (Green): Highlights the importance of safety warnings.
* "Data" (Blue): Indicates a focus on data analysis.
* "Diagnosed" (Green): Highlights the importance of diagnosis.
* "Drugs" (Green): Suggests a focus on pharmaceutical interventions.
* "Stay" (Green): Indicates a focus on patient stay.
* "Study" (Green): Highlights the importance of research studies.
* "Significant" (Green): Indicates a focus on significant findings.
* "Effects" (Green): Highlights the importance of research effects.
* "Mental" (Blue): Indicates a focus on mental health.
* "Addiction" (Green): Suggests research related to addiction.
* "Diseases" (Green): Indicates a focus on specific diseases.
* "Time" (Green): Highlights the importance of time-related factors.
* "Vaccine" (Blue): Suggests research related to vaccines.
* "Hospitals" (Green): Indicates a focus on hospital settings.
* "Photograph" (Green): Highlights the importance of photographic documentation.
* "Effect" (Green): Highlights the importance of research effects.
* "Role" (Green): Indicates a focus on roles in healthcare.
* "Blood" (Green): Indicates a focus on blood-related research.
* "News" (Green): Highlights the importance of news dissemination.
* "Serious" (Green): Indicates a focus on serious conditions.
* "Researchers" (Green): Highlights the importance of research personnel.
* "Ways" (Green): Indicates a focus on different approaches.
* "Lot" (Green): Indicates a focus on lot sizes.
* "Of" (Green): Indicates a focus on relationships.
* "You" (Green): Indicates a focus on patient involvement.
* "Cell" (Green): Indicates a focus on cellular research.
* "Leading" (Green): Indicates a focus on leading research.
* "MS" (Blue): Indicates a focus on multiple sclerosis.
* "Kidney" (Green): Indicates a focus on kidney health.
* "Emergency" (Green): Indicates a focus on emergency care.
* "Dementia" (Green): Indicates a focus on dementia research.
* "Ildenafil" (Yellow): Indicates a focus on ildenafil research.
* "Diagnosis" (Green): Highlights the importance of diagnosis.
* "Izaion" (Green): Indicates a focus on ionization research.
* "UE" (Green): Indicates a focus on user experience.
* "Po" (Green): Indicates a focus on patient outcomes.
* "FS" (Green): Indicates a focus on financial services.
* "Most" (Green): Indicates a focus on most common conditions.
* "Doctor" (Green): Indicates a focus on doctor involvement.
* "Pain" (Green): Indicates a focus on pain management.
* "Ynec" (Yellow): Indicates a focus on gynecological research.
### Key Observations
* The word cloud emphasizes the importance of "Health," "Patient," "Treatment," and "Drug" in medical research.
* There is a significant focus on specific diseases like "Diabetes" and "Cancer."
* The inclusion of terms like "FDA" and "GDPR" suggests an awareness of regulatory and privacy considerations.
### Interpretation
The word cloud provides a snapshot of the key themes and areas of focus within medical research. The prominence of terms like "Health," "Patient," and "Treatment" underscores the patient-centric nature of the field. The presence of disease-specific terms indicates targeted research efforts, while the inclusion of regulatory terms highlights the importance of ethical and legal considerations. The word cloud suggests a field that is both broad in scope and highly specialized, with a strong emphasis on improving patient outcomes and advancing medical knowledge.
</details>
Figure 14: Word cloud visualization of shifted tokens. The size of each token represents its rank improvement ratio ( $\frac{\text{base\_rank}}{\text{aligned\_rank}}$ ), indicating the magnitude of distributional shift during alignment. Larger tokens indicate more significant shifts in the model’s prediction patterns.
Our analysis of the MMLU dataset revealed significant and efficient distribution shifts between the base and aligned models. Figure 14 visualizes the most significantly shifted tokens, where the size of each token is proportional to its rank improvement ratio.
The analysis revealed a notably efficient token distribution shift pattern. Specifically, only 2.18% of tokens underwent significant shifts (compared to 5.61% in full pretraining), with 88.78% remaining unshifted and 9.04% showing marginal changes (Totally 645496 tokens analyzed). This represents a more focused and efficient alignment compared to full pretraining scenarios, which typically show higher shift percentages (unshifted: 75.59%, marginal: 18.80%, shifted: 5.61%).
Most remarkably, the shifted tokens demonstrate a clear concentration in medical terminology and medicine-related concepts. Key examples include: ”prescription”, ”diagnosis”, ”symptoms”, ”diabetes”, ”arthritis”, ”tumor”, ”MRI”, ”therapy”, ”treatment”, ”hospital”, ”care”, ”patients”.
This specialized distribution stands in stark contrast to the more general token shifts observed in full pretraining, where top shifted tokens (such as <|im_end|>, ”CIF”, ”Registered”, ”progression”, ”median”) show no particular domain focus and more noise. This comparison suggests that ADEPT achieved a more targeted and efficient knowledge injection, specifically enhancing the model’s medical domain expertise while maintaining stability in other areas. The lower percentage of shifted tokens (2.18% vs 5.61%) combined with their high domain relevance indicates a more precise and economical alignment process that effectively injects medical knowledge without unnecessary perturbation of the model’s general language capabilities.
These findings suggest that domain-specific alignment can be achieved with minimal token distribution disruption while maintaining high effectiveness in knowledge injection. This efficiency in token shifting demonstrates the potential for targeted domain adaptation without the broader distributional changes typically seen in full pretraining scenarios.
Similarly, in mathematical domain alignment (Figure 15), we observed an even more efficient token distribution shift. The analysis shows only 1.24% of tokens underwent significant shifts, with 91.51% remaining unshifted and 7.25% showing marginal changes. This represents an even more concentrated alignment compared to full pretraining (unshifted: 85.45%, marginal: 10.18%, shifted: 4.37%).
The shifted tokens clearly reflect mathematical and scientific terminology, as evidenced by terms such as ”theorem”, ”quantum”, ”parameters”, ”physics”, and ”equation”. This highly focused shift pattern, utilizing merely one-third of the token shifts compared to full pretraining (1.24% vs 4.37%), demonstrates the effectiveness of our approach in precisely targeting mathematical knowledge injection while maintaining model stability in other domains.
<details>
<summary>figures/wordcloud_math.jpg Details</summary>

### Visual Description
## Word Cloud: Research Topics
### Overview
The image is a word cloud, visually representing the frequency of different terms. The size of each word corresponds to its frequency or importance. The words are related to research, academia, and scientific topics.
### Components/Axes
There are no axes in a word cloud. The key component is the collection of words themselves, with their relative sizes indicating frequency.
### Detailed Analysis or ### Content Details
Here's a transcription of the words, grouped loosely by location and size (largest to smallest):
* **Left Side:**
* algorithm
* students
* paper
* user
* talk
* scripts
* Oct
* problem
* sense
* udos
* de
* **Center:**
* mathematics
* space
* effects
* following
* editor
* physics
* answer
* previous
* quantum
* parameters
* model
* bit
* reaction
* identity
* theorem
* isation
* **Right Side:**
* question
* PhD
* results
* Rig
* value
* open
* profit
* error
* features
* code
* al
* ule
* to
* A
* chner
* case
* **Other words:**
* PE
* phenotype
* ob
* variance
* scatter
* et
* F
* Tony
* Sized
* ja
* ric
* ormal
* esium
* pton
* papers
* $$
* AG
* sh
* connected
* quant
* new
* ex
* CM
* the
* FT
* ATE
* he
* groups
* {
* }
* |
* b1
* 2
* qu
* $?$
### Key Observations
* The largest words include "algorithm," "mathematics," "space," "effects," "following," "editor," "physics," "answer," "previous," "quantum," "question," "PhD," and "results."
* There are some symbols and abbreviations scattered throughout, such as "$", "ex", "PE", "CM", "FT", "ATE", "AG", "sh", "b1", "2", "{", "}", "|", and "?".
* The words are arranged in a seemingly random fashion, typical of word clouds.
### Interpretation
The word cloud provides a visual summary of the topics most frequently discussed or emphasized in a given context (likely a collection of research papers, articles, or discussions). The prominence of terms like "algorithm," "mathematics," "physics," and "quantum" suggests a focus on quantitative and scientific research. The presence of words like "question," "results," and "PhD" indicates an academic or research-oriented environment. The cloud highlights the key themes and areas of interest within the source material.
</details>
Figure 15: Word cloud visualization of shifted tokens in mathematical domain alignment. The predominance of mathematical and scientific terminology demonstrates the precise targeting of domain-specific knowledge.
Appendix J Linear Merge of Domain-Specific Extensions: Results and Insights
Table 12: Performance comparison of Vanilla and Merged Models on multiple benchmarks (Qwen3-1.7B and Qwen3-4B).
| | MMLU | CMMLU | GSM8K | ARC-E | ARC-C | MedQA | MMCU | CMB |
| --- | --- | --- | --- | --- | --- | --- | --- | --- |
| Qwen3-1.7B | | | | | | | | |
| Vanilla | 62.57 | 66.86 | 57.62 | 81.44 | 51.19 | 48.39 | 69.17 | 63.67 |
| Merged Model | 62.70 | 65.83 | 60.80 | 81.06 | 51.94 | 48.39 | 68.61 | 64.83 |
| Qwen3-4B | | | | | | | | |
| Vanilla | 73.19 | 77.92 | 69.07 | 85.52 | 59.13 | 62.77 | 82.44 | 78.92 |
| Merged Model | 72.96 | 77.99 | 73.16 | 85.27 | 58.96 | 62.83 | 82.83 | 78.42 |
In Table 12, we compare the performance of the Vanilla model and the Merged Model, which was constructed by linearly merging the domain-specific extension layers (with equal weights of 0.5 for medical and mathematical domains) after independent training. Our results show that the merged model does not exhibit any significant collapse, and in some indicators even surpasses the original base model. For example, on the GSM8K benchmark for Qwen3-1.7B, the merged model achieves 60.80%, compared to 57.62% for the vanilla model. This demonstrates the generalization and extensibility of our method, enabling fusion across multiple vertical domains.
Our extension approach ensures that each newly added layer is separated by at least one original frozen layer, rather than being directly adjacent. This design leads to greater stability during model merging. On one hand, if the merged models were purely cascaded, the non-linear transformations introduced could lead to more unpredictable interactions between layers. On the other hand, because each layer operates within a consistent contextual environment provided by surrounding frozen layers during continual pre-training, we believe that this fixed hierarchical structure imposes constraints that make the semantic representations learned by the new layers more aligned in certain dimensions. As a result, the merging process becomes more reliable and beneficial to overall model performance.
It is worth noting that our merging strategy adopts the simplest possible weighted average. The specific merging algorithm is not the focus of this work; we believe that with more scientific weighting schemes, even better results can be obtained. Here, we hope to stimulate further research and provide preliminary insights based on our observations.
Appendix K Use of LLM
In the preparation of this article, we utilized large language models (LLM) solely for writing assistance purposes. Specifically, we employed the GPT-4.1-0414 model to polish language expressions, condense sentences, and improve the overall clarity and readability of the text. The model was used exclusively for editing and refining manuscript language and did not participate in any conceptual or technical aspects of this work.
All research ideas, theoretical proof methods, experimental designs, and visualizations were conceived, executed, and finalized by the authors without the involvement of any LLM tools. The development of new concepts, formulation and validation of proofs, experimental setups, analysis of results, and the creation of figures were performed independently by the research team. At no point was the LLM model used to generate, modify, or validate the scientific content, methodology, or results presented in this article.
We emphasize that the role of GPT-4.1-0414 in this research was strictly limited to linguistic enhancement at the writing stage, and that all substantive intellectual and scientific contributions originate solely from the authors.
Appendix L Algorithm
Algorithm 1 ADEPT
1: Pretrained LLM $M_{0}$ with layers $\{\Theta^{(1)},...,\Theta^{(L)}\}$ , domain probing corpus $\mathcal{D}_{\text{probe}}$ , continual pretraining corpus $\mathcal{D}_{\text{train}}$ , number of layers to expand $k$ , base learning rate $\text{lr}_{\text{base}}$ , update interval $T_{\text{update}}$
2: # Stage 1: General-Competence Guided Selective Layer Expansion
3: Compute base loss $\mathcal{L}_{\text{base}}←\frac{1}{|\mathcal{D}_{\text{probe}}|}\sum_{x}\ell(M_{0}(x),x)$
4: for $l← 1$ to $L$ do
5: Temporarily mask layer $l$ to get $M_{0}^{(-l)}$
6: Compute masked loss $\hat{\mathcal{L}}^{(l)}←\frac{1}{|\mathcal{D}_{\text{probe}}|}\sum_{x}\ell(M_{0}^{(-l)}(x),x)$
7: Compute importance score $\Delta^{(l)}←\hat{\mathcal{L}}^{(l)}-\mathcal{L}_{\text{base}}$
8: end for
9: Select $k$ least-important layers $\mathcal{S}_{k}←\text{LowestK}(\{\Delta^{(l)}\})$
10: for each $l∈\mathcal{S}_{k}$ do
11: Duplicate parameters $\tilde{\Theta}^{(l)}←\Theta^{(l)}$ $\triangleright$ Identity copy
12: Initialize $W_{\text{MHSA}}^{\text{out}}=0$ , $W_{\text{FFN}}^{\text{out}}=0$ $\triangleright$ Function Preserving Init
13: Freeze original $\Theta^{(l)}$ , mark $\tilde{\Theta}^{(l)}$ as trainable
14: end for
15: # Stage 2: Adaptive Unit-Wise Decoupled Tuning
16: for each training step $t$ do
17: if $t\mod T_{\text{update}}==0$ then
18: for each expanded layer $\tilde{\Theta}^{(l)}$ do
19: Partition into semantic units $\{U_{1},...,U_{n}\}$
20: for each unit $U_{i}$ do
21: Compute gradient-based importance $I_{U_{i}}←\frac{1}{|U_{i}|}\sum_{j∈ U_{i}}\theta_{j}·∇_{\theta_{j}}\mathcal{L}$
22: Assign adaptive learning rate $\text{lr}_{U_{i}}← 2·(1-I_{U_{i}})·\text{lr}_{\text{base}}$
23: end for
24: end for
25: end if
26: Sample training sequence $x=(x_{1},x_{2},...,x_{T})\sim\mathcal{D}_{\text{train}}$
27: Compute autoregressive loss:
28: $\mathcal{L}=-\sum_{t=1}^{T}\log P(x_{t}\mid x_{<t};\Theta)$
29: Update parameters $\{\tilde{\Theta}^{(l)}\}$ using adaptive learning rates $\{\text{lr}_{U_{i}}\}$
30: end for