# LLM-Based Scientific Equation Discovery via Physics-Informed Token-Regularized Policy Optimization
**Authors**: Boxiao Wang, Kai Li, Tianyi Liu, Chen Li, Junzhe Wang, Yifan Zhang, Jian Cheng
> 0009-0008-2970-7575 Institute of Automation, Chinese Academy of Sciences Beijing China
> Institute of Automation, Chinese Academy of Sciences Beijing China
> State Key Laboratory of Aerodynamics Mianyang, Sichuan China
> School of Mathematical Sciences, University of Chinese Academy of Sciences Beijing China
## Abstract
Symbolic regression aims to distill mathematical equations from observational data. Recent approaches have successfully leveraged Large Language Models (LLMs) to generate equation hypotheses, capitalizing on their vast pre-trained scientific priors. However, existing frameworks predominantly treat the LLM as a static generator, relying on prompt-level guidance to steer exploration. This paradigm fails to update the model’s internal representations based on search feedback, often yielding physically inconsistent or mathematically redundant expressions. In this work, we propose PiT-PO (Physics-informed Token-regularized Policy Optimization), a unified framework that evolves the LLM into an adaptive generator via reinforcement learning. Central to PiT-PO is a dual-constraint mechanism that rigorously enforces hierarchical physical validity while simultaneously applying fine-grained, token-level penalties to suppress redundant structures. Consequently, PiT-PO aligns LLM to produce equations that are both scientifically consistent and structurally parsimonious. Empirically, PiT-PO achieves state-of-the-art performance on standard benchmarks and successfully discovers novel turbulence models for challenging fluid dynamics problems. We also demonstrate that PiT-PO empowers small-scale models to outperform closed-source giants, democratizing access to high-performance scientific discovery.
conference: ; ;
## 1. Introduction
Symbolic Regression (SR) (Makke and Chawla, 2024b) stands as a cornerstone of data-driven scientific discovery, uniquely capable of distilling interpretable mathematical equations from observational data. Unlike black-box models that prioritize mere prediction, SR elucidates the fundamental mechanisms governing system behavior, proving instrumental in uncovering physical laws (Makke and Chawla, 2024a; Reuter et al., 2023), modeling chemical kinetics (Chen et al., 2025; Deng et al., 2023), and analyzing complex biological dynamics (Wahlquist et al., 2024; Shi et al., 2024).
However, the search for exact governing equations represents a formidable challenge, formally classified as an NP-hard problem (Virgolin and Pissis, 2022). To navigate this vast search space, algorithmic strategies have evolved from Genetic Programming (GP) (Schmidt and Lipson, 2009; Cranmer, 2023) and Reinforcement Learning (RL) (Petersen et al., 2021) to Transformer-based architectures that map numerical data directly to symbolic equations (Biggio et al., 2021; Kamienny et al., 2022; Zhang et al., 2025). Most recently, the advent of Large Language Models (LLMs) has introduced a new paradigm. Methods such as LLM-SR (Shojaee et al., 2025a) and LaSR (Grayeli et al., 2024) leverage the pre-trained scientific priors and in-context learning capabilities of LLMs to generate equation hypotheses. These approaches typically employ an evolutionary search paradigm, where candidates are evaluated, and high-performing solutions are fed back via prompt-level conditioning to steer subsequent generation.
Despite encouraging progress, existing LLM-based SR methods remain constrained by severe limitations. First, most approaches treat the LLM as a static generator, relying primarily on prompt-level, verbal guidance to steer the evolutionary search (Shojaee et al., 2025a; Grayeli et al., 2024). This “frozen” paradigm inherently neglects the opportunity to adapt and enhance the generative capability of the LLM itself based on evaluation signals, preventing the model from internalizing feedback and adjusting its generation strategies to the specific problem. Second, they typically operate in a physics-agnostic manner, prioritizing syntactic correctness over physical validity (Shojaee et al., 2025a; Grayeli et al., 2024). Without rigorous constraints, LLMs often generate equations that fit the data numerically but violate fundamental physical principles, rendering them prone to overfitting and practically unusable.
In this work, we propose to fundamentally shift the role of the LLM in SR from a static proposer to an adaptive generator. We establish a dynamic feedback loop in which evolutionary exploration and parametric learning reinforce each other: evolutionary search uncovers diverse candidate equations and generates informative evaluation signals, while parametric adaptation enables the LLM to consolidate effective symbolic patterns and guide subsequent exploration more efficiently. By employing in-search fine-tuning, i.e., updating the LLM parameters during the evolutionary search process, we move beyond purely verbal, prompt-level guidance and introduce numerical guidance that allows feedback to be directly internalized into the model parameters, progressively aligning LLM with the intrinsic properties of the target system.
To realize this vision, we introduce PiT-PO (Physics-informed Token-regularized Policy Optimization), a unified framework that bridges LLM-driven evolutionary exploration with rigorous verification. PiT-PO is built upon two technical components. 1) In-Search LLM Evolution. We implement the numerical guidance via reinforcement learning, which efficiently updates the LLM’s parameters during the search process. Instead of relying on static pre-trained knowledge, this in-search policy optimization enables LLM to dynamically align its generative distribution with the structural characteristics of the specific task, effectively transforming general scientific priors into domain-specific expertise on the fly. 2) Dual Constraints as Search Guidance. We enforce hierarchical physical constraints to ensure scientific validity, and uniquely, we incorporate a fine-grained regularization based on our proposed Support Exclusion Theorem. This theorem allows us to identify mathematically redundant terms and translate them into token-level penalties, effectively pruning the search space and guiding LLM toward physically meaningful and structurally parsimonious equations.
Comprehensive experimental results demonstrate that PiT-PO achieves state-of-the-art performance across standard SR benchmarks, including the LLM-SR Suite (Shojaee et al., 2025a) and LLM-SRBench (Shojaee et al., 2025b), and recovers the largest number of ground-truth equations among all evaluated methods. In addition to synthetic benchmarks, the effectiveness of PiT-PO is validated on an application-driven turbulence modeling task involving flow over periodic hills. PiT-PO improves upon traditional Reynolds-Averaged Navier-Stokes (RANS) approaches by producing anisotropic Reynolds stresses closer to Direct Numerical Simulation (DNS) references. The learned method shows enhanced physical consistency, with reduced non-physical extremes and better flow field predictions. Finally, PiT-PO maintains robust performance even when using resource-constrained small models, such as a quantized version of Llama-8B, and remains efficient under strict wall-clock time budgets, establishing a practical methodology for automated scientific discovery.
<details>
<summary>x1.png Details</summary>
### Visual Description
\n
## Diagram: PiT-PO System Architecture
### Overview
The image depicts a diagram illustrating the architecture of a "PiT-PO" (Physics-informed Token-Regularized Policy Optimization) system. It shows a cyclical process involving an LLM Policy, Island-Based Exploration, Physical and Theoretical Constraints, and a Policy Update mechanism. The diagram emphasizes the flow of information and the interplay between different components.
### Components/Axes
The diagram consists of the following key components:
* **LLM Policy (πθ):** A light blue, rounded rectangle positioned on the left side of the diagram.
* **Island-Based Exploration:** A rectangular box containing a series of labeled functions (f₀, f₁, ..., fₙ) connected by grey arrows.
* **Physical Constraints:** An orange rectangular box labeled with "General-Level pdims, pdiff" and "Domain-Specific pdomain".
* **Theoretical Constraints:** A red rectangular box labeled with "Support Exclusion Theorem ptok".
* **PiT-PO:** A purple circular shape representing the core optimization process.
* **GRPO:** A dark purple rectangular box within PiT-PO.
* **Policy Update:** A curved arrow at the top of the diagram indicating the direction of policy improvement.
* **Prompt Update:** A curved arrow at the bottom of the diagram indicating the direction of prompt refinement.
* **Global Reward:** A connection from Physical Constraints to PiT-PO.
* **Token-Aware Advantage Estimation:** A connection from Physical Constraints to PiT-PO.
* **Token Penalty:** A connection from Theoretical Constraints to PiT-PO.
* **Physics-informed Token-Regularized Policy Optimization:** Text labeling the process within PiT-PO.
### Detailed Analysis or Content Details
The diagram illustrates a feedback loop. The LLM Policy (πθ) generates outputs that are then used for Island-Based Exploration. This exploration generates a set of functions (f₀ to fₙ). These functions are then fed into both Physical Constraints and Theoretical Constraints. The Physical Constraints are defined by "General-Level pdims, pdiff" and "Domain-Specific pdomain". The Theoretical Constraints are defined by the "Support Exclusion Theorem ptok".
The outputs of both Physical and Theoretical Constraints are then fed into the PiT-PO module. PiT-PO contains GRPO and utilizes "Global Reward", "Token-Aware Advantage Estimation", and "Token Penalty" to perform "Physics-informed Token-Regularized Policy Optimization".
Finally, PiT-PO updates both the Policy (via "Policy Update") and the Prompt (via "Prompt Update"), completing the cycle.
The functions f₀ through fₙ are connected to both the Physical and Theoretical Constraints with grey arrows. The number of functions 'n' is not explicitly defined, but is represented by the ellipsis (...).
### Key Observations
The diagram highlights the integration of physical and theoretical constraints into the policy optimization process. The cyclical nature of the diagram suggests an iterative refinement process. The use of "Token-Aware" and "Token Penalty" indicates a focus on token-level regularization, likely within the context of a large language model. The diagram is highly conceptual and does not contain numerical data.
### Interpretation
The diagram represents a novel approach to policy optimization that leverages the strengths of LLMs while incorporating physical and theoretical constraints to improve performance and robustness. The "Island-Based Exploration" suggests a method for generating diverse candidate policies. The constraints act as a filter, ensuring that the policies are physically plausible and theoretically sound. The PiT-PO module then refines these policies using token-level regularization, potentially addressing issues such as hallucination or instability. The feedback loop ensures that the policy and prompt are continuously improved based on the constraints and optimization process. The diagram suggests a system designed to create more reliable and physically grounded LLM-based policies. The use of the term "theorem" suggests a mathematically rigorous foundation for the theoretical constraints. The diagram is a high-level architectural overview and does not provide details on the specific algorithms or implementation techniques used.
</details>
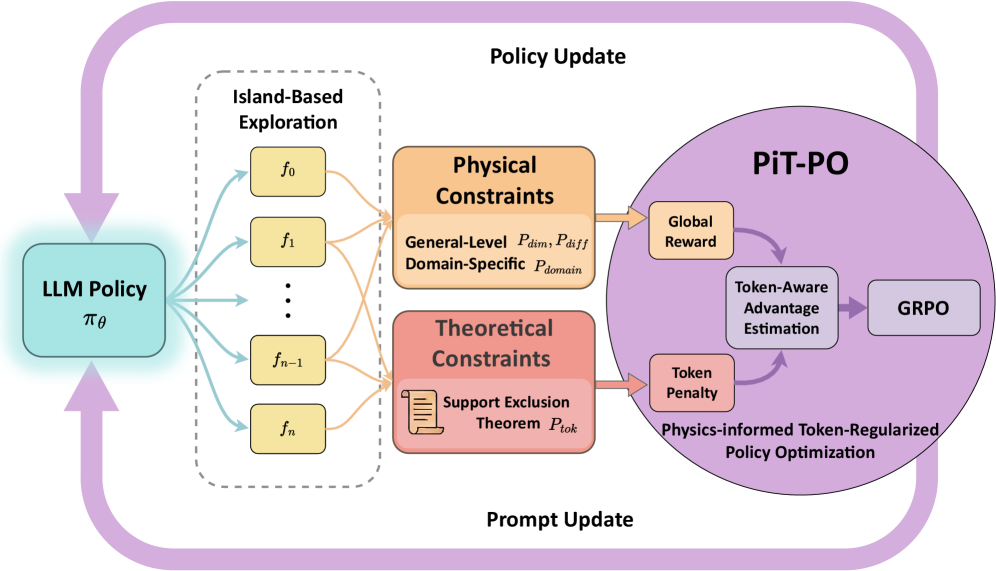
Figure 1. The overall framework of PiT-PO. PiT-PO transforms the LLM from a static proposer into an adaptive generator via a closed-loop evolutionary process. The framework integrates dual-constraint evaluation—comprising physical constraints and theoretical constraints—to generate fine-grained token-level learning signals. These signals guide the LLM policy update via reinforcement learning, ensuring the discovery of parsimonious, physically consistent equations.
## 2. Preliminaries
### 2.1. Problem Setup
In SR, a dataset of input–output observations is given:
$$
D=\{(x_{i},y_{i})\}_{i=1}^{n},\;x_{i}\in\mathbb{R}^{d},\;y_{i}\in\mathbb{R}. \tag{1}
$$
The objective is to identify a compact and interpretable function $f\in\mathcal{F}$ such that $f(x_{i})\approx y_{i}$ for the observed samples, while retaining the ability to generalize to unseen inputs.
### 2.2. LLM-based SR Methods
Contemporary LLM-based approaches reformulate SR as a iterative program synthesis task. In this paradigm, typified by frameworks such as LLM-SR (Shojaee et al., 2025a), the discovery process is decoupled into two phases: structure proposal and parameter estimation. Specifically, the LLM functions as a symbolic generator, emitting functional skeletons with placeholders for learnable coefficients. A numerical optimizer (e.g., BFGS (Fletcher, 1987)) subsequently fits these constants to the observed data. To navigate the combinatorial search space, these methods employ an evolutionary-style feedback loop: high-fitness equations are maintained in a pool to serve as in-context examples, prompting the LLM to refine subsequent generations. Our work leverages this architecture as a backbone, but fundamentally redefines the LLM’s role from a static proposer to an adaptive generator.
### 2.3. Group Relative Policy Optimization
Group Relative Policy Optimization (GRPO) (Shao et al., 2024) is a RL algorithm tailored for optimizing LLMs on reasoning tasks, characterized by its efficient baseline estimation without the need for a separate value network. In the context of LLM-based SR, the generation process is modeled as a Markov Decision Process (MDP), where LLM functions as a policy $\pi_{\theta}$ that generates a sequence of tokens $o=(t_{1},\dots,t_{L})$ given a prompt $q$ . For each $q$ , GRPO samples a group of $G$ outputs $\{o_{1},\dots,o_{G}\}$ from the sampling policy $\pi_{\theta_{old}}$ . GRPO maximizes the following surrogate loss function:
$$
\mathcal{J}_{GRPO}(\theta)=\mathbb{E}_{q\sim P(Q),\{o_{i}\}\sim\pi_{\theta_{old}}}\left[\frac{1}{G}\sum_{i=1}^{G}\left(\frac{1}{L_{i}}\sum_{k=1}^{L_{i}}\mathcal{L}^{clip}_{i,k}(\theta)-\beta\mathbb{D}_{KL}(\pi_{\theta}||\pi_{ref})\right)\right], \tag{2}
$$
where $\pi_{ref}$ is the reference policy to prevent excessive deviation, and $\beta$ controls the KL-divergence penalty. The clipping term $\mathcal{L}^{clip}_{i,k}(\theta)$ ensures trust region updates:
$$
\mathcal{L}^{clip}_{i,k}(\theta)=\min\left(\frac{\pi_{\theta}(t_{i,k}|q,o_{i,<k})}{\pi_{\theta_{old}}(t_{i,k}|q,o_{i,<k})}\hat{A}_{i},\;\text{clip}\left(\frac{\pi_{\theta}(t_{i,k}|q,o_{i,<k})}{\pi_{\theta_{old}}(t_{i,k}|q,o_{i,<k})},1-\epsilon,1+\epsilon\right)\hat{A}_{i}\right). \tag{3}
$$
Here, $\epsilon$ is the clipping coefficient. A distinctive feature of GRPO lies in its advantage estimation, it computes the advantage $\hat{A}_{i}$ by standardizing the reward $R(o_{i})$ relative to the group:
$$
\hat{A}_{i}=\frac{R(o_{i})-\text{mean}(\{R(o_{j})\})}{\text{std}(\{R(o_{j})\})}. \tag{4}
$$
Consequently, every token within the sequence $o_{i}$ is assigned the exact same feedback signal. This coarse granularity treats valid and redundant terms indistinguishably, a limitation that our work addresses by introducing fine-grained, token-level regularization.
## 3. Method
We propose PiT-PO (Physics-informed Token-regularized Policy Optimization), a framework that evolves LLM into an adaptive, physics-aware generator. PiT-PO establishes a closed-loop evolutionary process driven by two synergistic mechanisms: (1) a dual-constraint evaluation system that rigorously assesses candidates through hierarchical physical verification and theorem-guided redundancy pruning; and (2) a novel policy optimization strategy that updates LLM using fine-grained, token-level feedback derived from these constraints. This combination effectively aligns the LLM with the intrinsic structure of the problem, guiding the LLM toward solutions that are not only numerically accurate but also structurally parsimonious and scientifically consistent.
### 3.1. Dual-Constraint Learning Signals
Navigating the combinatorial space of symbolic equations requires rigorous guidance. We employ a reward system driven by dual constraints: physical constraints delineate the scientifically valid region, while theoretical constraints drive the search toward simpler equations by identifying and pruning redundant terms.
#### 3.1.1. Hierarchical Physical Constraints.
To ensure scientific validity, we construct a hierarchical filter that categorizes constraints into two levels: general properties and domain-specific priors.
General-Level Constraints. We enforce fundamental physical properties applicable across scientific disciplines. To prune physically impossible structures (e.g., adding terms with mismatched units), we assign penalty-based rewards for Dimensional Homogeneity ( $P_{dim}$ ) and Differentiability ( $P_{diff}$ ). The former strictly penalizes equations with unit inconsistencies, while the latter enforces smoothness on the data-defined domain.
Domain-Specific Constraints. To tackle specialized tasks, we inject expert knowledge as inductive biases. We define the domain-specific penalty $P^{(j)}_{domain}$ to penalize candidate equations that violate the $j$ -th domain-specific constraint. Taking the turbulence modeling task (detailed in Appendix E.4) as a representative instantiation, we enforce four rigorous constraints: (1) Realizability (Pope, 2000), ensuring the Reynolds stress tensor has positive eigenvalues; (2) Boundary Condition Consistency (Monkewitz, 2021), requiring stresses to decay to zero at the wall; (3) Asymptotic Scaling (Tennekes and Lumley, 1972; WANG et al., 2019), enforcing the cubic relationship between stress and wall distance in the viscous sublayer; and (4) Energy Consistency (Pope, 2000; MOCHIZUKI and OSAKA, 2000), aligning predicted stress with turbulent kinetic energy production.
This hierarchical design effectively embeds physical consistency as a hard constraint in the reward function, prioritizing scientific validity over mere empirical fitting.
#### 3.1.2. Theorem-Guided Mathematical Constraints
While physical constraints ensure validity, they do not prevent mathematical redundancy. To rigorously distinguish between essential terms and redundant artifacts, we introduce the Support Exclusion Theorem.
Let $\mathcal{S}$ denote the full support set containing all candidate basis functions $\{\phi_{j}\}$ . The ground truth equation is $f^{*}=\sum_{j\in\mathcal{S}^{\prime}}a_{j}\phi_{j}$ , where $\mathcal{S}^{\prime}\subseteq\mathcal{S}$ is the true support set (i.e., the indices of basis functions that truly appear in the governing equation), and $\{a_{j}\}_{j\in\mathcal{S}^{\prime}}$ are the corresponding true coefficients. Consider a candidate equation $f=\sum_{j\in\mathcal{K}}b_{j}\phi_{j}$ , where $\mathcal{K}\subseteq\mathcal{S}$ represents the current support set (i.e., the selected terms in the skeleton), $\mathbf{b}=\{b_{j}\}_{j\in\mathcal{K}}$ are the optimized coefficients derived from the data. We define the empirical Gram matrix of these basis functions as $G\in\mathbb{R}^{|\mathcal{S}|\times|\mathcal{S}|}$ and the corresponding Projection Matrix as $T$ , where $T_{ij}:=G_{ji}/G_{ii}$ .
** Theorem 3.1 (Support Exclusion Theorem)**
*Assume the ground-truth support is finite and satisfies $|\mathcal{S}^{\prime}|\leq M$ , and let the true function coefficients be bounded by $A\leq|a_{j}|\leq B$ for all $j\in\mathcal{S}^{\prime}$ . A term $\phi_{i}$ ( $i\in\mathcal{K}$ ) is theoretically guaranteed to be a false discovery (not in the true support $\mathcal{S}^{\prime}$ ) if its fitted coefficient magnitude satisfies:
$$
|b_{i}|<A-\left(\underbrace{\sum_{j\in\mathcal{K},j\neq i}(B+|b_{j}|)|T_{ij}|}_{\text{Internal Interference}}+\underbrace{B\sum_{k=1}^{m}s_{(k)}}_{\text{External Interference}}\right). \tag{5}
$$
$s(k)$ denotes the $k$ -th largest value in $\{|T_{i\ell}|:\ell\in\mathcal{S}\setminus\mathcal{K}\}$ , and $m:=\min\!\big(M-1,\;|\mathcal{S}\setminus\mathcal{K}|\big)$ .*
Detailed definitions of all notations and the rigorous proof of Theorem 3.1 are provided in Appendix B. This theorem formalizes the intuition that coefficients of redundant terms (absent from the true support $\mathcal{S}^{\prime}$ ) have significantly smaller magnitudes than those of valid components.
Specifically, after fitting $\mathbf{b}$ , we compute the normalized coefficient ratio $\tau_{i}=|b_{i}|/(\sum_{j}|b_{j}|+\epsilon)$ . We introduce a threshold $\rho\in(0,1)$ to identify potentially redundant terms. Terms satisfying $\tau_{i}>\rho$ incur no penalty, while components with $\tau_{i}\leq\rho$ are considered redundant. To suppress these redundancies, we define a token penalty for each token in redundant term $i$ :
$$
P_{tok}=p\cdot\max\left(0,-\log\left(|b_{i}|+\epsilon\right)\right), \tag{6}
$$
where $p>0$ is a scaling coefficient. We use a logarithmic scale to impose stronger penalties on terms with smaller coefficients.
By integrating this penalty into the policy optimization, we guide the LLM to reduce the probability of generating redundant terms, thereby steering the optimization toward parsimonious equations.
### 3.2. Token-Aware Policy Update
Our proposed PiT-PO effectively operationalize the hierarchical constraints and theoretical insights derived in Section 3.1. Unlike standard GRPO that assign a uniform scalar reward to the entire generated sequence, our method transitions the learning process from coarse-grained sequence scoring to fine-grained token-level credit assignment. This ensures that the policy not only learns to generate physically valid equations but also explicitly suppresses theoretically redundant terms.
#### 3.2.1. Global Reward with Gated Constraints
The optimization is driven by a composite global reward, $R_{global}$ , which balances fitting accuracy, structural parsimony, and physical consistency. Formally, for a sampled equation $o_{i}$ , the rewards are defined as follows:
Fitting Accuracy ( $R_{fit}$ ). We use the normalized log-MSE to encourage precise data fitting:
$$
R_{fit}=-\alpha\log(\text{MSE}+\epsilon), \tag{7}
$$
where $MSE=\frac{1}{n}\sum_{i=1}^{n}(y_{i}-\hat{y}_{i})^{2}$ .
Complexity Penalty ( $P_{cplx}$ ). Adhering to Occam’s Razor, we penalize structural complexity based on the Abstract Syntax Tree (AST) (Neamtiu et al., 2005) node count:
$$
P_{cplx}=\lambda_{len}\cdot\text{Length}(\text{AST}). \tag{8}
$$
In PiT-PO, each equation generated by the LLM is represented as a Python function and parsed into an AST, where each node corresponds to a variable or operator. The total node count provides a meaningful estimate of structural complexity.
Gated Physical Penalty ( $P_{phy}$ ). Imposing strict physical constraints too early can hinder exploration, causing the model to discard potentially promising functional forms. We therefore activate physical penalties only after the candidate equation reaches a baseline fitting accuracy threshold ( $\delta_{gate}$ ). Specifically, we define
where $\mathbb{1}(\cdot)$ is the indicator function. This mechanism effectively creates a soft curriculum: it allows “free” exploration in the early stages and enforcing strict physical compliance only after the solution enters a plausible region.
The total reward $R_{global}$ is then formulated as:
$$
R_{global}(o_{i})=R_{fit}(o_{i})-P_{cplx}(o_{i})-P_{phy}(o_{i}). \tag{10}
$$
#### 3.2.2. Fine-Grained Advantage Estimation
Standard GRPO applies a uniform advantage across all tokens in a sequence. We refine this by synthesizing the global reward with the token-level penalty $P_{tok}$ (Equation 6). Specifically, we define the token-aware advantage $\hat{A}_{i,k}$ for the $k$ -th token in the $i$ -th sampled equation as:
$$
\hat{A}_{i,k}=\underbrace{\frac{R_{global}(o_{i})-\mu_{group}}{\sigma_{group}}}_{\text{Global Standardization}}-\underbrace{P_{i,k}}_{\text{Local Pruning}}. \tag{11}
$$
Here, the first term standardizes the global reward against the group statistics ( $\mu_{group},\sigma_{group}$ ), reinforcing equations that satisfy multi-objective criteria relative to their peers. The second term, $P_{i,k}$ , applies a targeted penalty to suppress redundancy. Specifically, we set $P_{i,k}=0$ if token $k$ belongs to a non-redundant term, and $P_{i,k}=P_{tok}$ otherwise. This ensures that penalties are applied exclusively to tokens contributing to mathematically redundant structures, while valid terms remain unaffected.
Substituting this token-aware advantage into the GRPO objective, the policy gradient update of our PiT-PO becomes:
$$
\nabla\mathcal{J}_{PiT-PO}\propto\sum_{i,k}\hat{A}_{i,k}\nabla\log\pi_{\theta}(t_{i,k}|o_{i,<k}). \tag{12}
$$
This creates a dual-pressure optimization landscape: global rewards guide the policy toward physically consistent and accurate equations, while local penalties surgically excise redundant terms. This ensures the final output aligns with the sparse, underlying physical laws rather than merely overfitting numerical data.
Input: Dataset $D=\{(x_{i},y_{i})\}_{i=1}^{n}$ ; LLM $\pi_{\theta}$ ; number of islands $N$ ; group size $G$ ; iterations $T$ .
Output: Best equation $o^{*}$ .
1
2 1ex
3 Initialize $o^{*}$ , $s^{*}$ , and buffers $\mathcal{B}_{j}\leftarrow\emptyset$ for $j=1,\dots,N$
4
5 for $t\leftarrow 1$ to $T$ do
// Stage 1: Island-Based Exploration
6 for $j\leftarrow 1$ to $N$ do
$q_{j}\leftarrow\textsc{BuildPrompt}(D,\mathcal{B}_{j})$
// in-context rule
$\{o_{i}\}_{i=1}^{G}\sim\pi_{\theta}(\cdot\mid q_{j})$
// sample a group
7 for $i\leftarrow 1$ to $G$ do
$(R_{i},\{P_{i,k}\})\leftarrow\textsc{DualConstraintEval}(o_{i},D)$
// $R_{i}=R_{\mathrm{global}}(o_{i})$
8 $\mathcal{B}_{j}\leftarrow\mathcal{B}_{j}\cup\{(q_{j},o_{i},R_{i},\{P_{i,k}\})\}$
9 if $R_{i}>s^{*}$ then
10 $o^{*}\leftarrow o_{i}$ ; $s^{*}\leftarrow R_{i}$
11
12
13
// Stage 2: In-Search LLM Evolution
14 $\theta\leftarrow\textsc{PiT-PO\_Update}(\theta,\{\mathcal{B}_{j}\}_{j=1}^{N},\pi_{{\theta}})$
// Stage 3: Hierarchical Selection
15 $\{\mathcal{B}_{j}\}_{j=1}^{N}\leftarrow\textsc{SelectAndReset}(\{\mathcal{B}_{j}\}_{j=1}^{N})$
16 return $o^{*}$
Algorithm 1 PiT-PO Overall Training Pipeline
### 3.3. Overall Training Pipeline
We orchestrate the PiT-PO framework through a closed-loop evolutionary RL cycle. As illustrated in Algorithm 1, the training process iterates through three synergistic phases:
Phase 1: Island-Based Exploration (Data Generation). To prevent premature convergence to local optima, a common pitfall in SR, we employ a standard multi-island topology ( $N$ islands) to structurally enforce search diversity (Cranmer, 2023; Romera-Paredes et al., 2024). Each island $j$ maintains an isolated experience buffer $\mathcal{B}_{j}$ , evolving its own lineage of equations. This information isolation allows distinct islands to cultivate diverse functional forms independently.
Phase 2: In-Search LLM Evolution (Policy Update). This phase transforms the collected data into parametric knowledge. We aggregate the trajectories from all $N$ islands into a global batch to perform policy optimization using PiT-PO. By minimizing the loss, the model explicitly lowers the probability of generating mathematically redundant tokens and physically inconsistent structures. To ensure computational efficiency during this iterative search, we implement the update using Low-Rank Adaptation (LoRA) (Hu et al., 2021).
Phase 3: Hierarchical Selection (Population Management). We apply standard survival-of-the-fittest mechanisms to maintain population quality. Local buffers $\mathcal{B}_{j}$ are updated by retaining only top-performing candidates, while underperforming islands are periodically reset with high-fitness seeds to escape local optima.
This cycle establishes a reciprocal reinforcement mechanism: the island-based exploration maintains search diversity, while policy update consolidates these findings into the model weights, progressively transforming the LLM into a domain-specialized scientific discoverer.
## 4. Experiments
### 4.1. Setup
Benchmarks. To provide a comprehensive evaluation of PiT-PO, we adopt two widely used benchmarks to compare against state-of-the-art baselines:
LLM-SR Suite (Shojaee et al., 2025a). This suite comprises four tasks spanning multiple scientific domains: Oscillation 1 & 2 (Nonlinear Oscillatory Systems) feature dissipative couplings and non-polynomial nonlinearities with explicit forcing, making recovery of the correct interaction terms from trajectory data non-trivial; E. coli Growth (Monod, 1949; Rosso et al., 1995) models multivariate population dynamics with strongly coupled, multiplicative effects from nutrients, temperature, and acidity; and Stress-Strain (Aakash et al., 2019) uses experimental measurements of Aluminum 6061-T651 and exhibits temperature-dependent, piecewise non-linear deformation behavior. Detailed information about these tasks is provided in Appendix D.1.
LLM-SRBench: (Shojaee et al., 2025b) To evaluate generalization beyond canonical forms, we adopt the comprehensive LLM-SRBench benchmark, which contains 239 tasks organized into two complementary subsets, LSR-Transform and LSR-Synth. LSR-Transform changes the prediction target to rewrite well-known physics equations into less common yet analytically equivalent forms, producing 111 transformed tasks. This design aims to reduce reliance on direct memorization of canonical templates and tests whether a method can recover the same physical law under non-trivial variable reparameterizations. Complementarily, LSR-Synth composes equations from both known scientific terms and synthetic but plausible terms to further assess discovery beyond memorized templates: candidate terms are proposed by an LLM under domain context, assembled into full equations, and then filtered through multiple checks, including numerical solvability, contextual novelty, and expert plausibility, yielding 128 synthetic tasks. Further details are given in Appendix D.2.
Baselines.
We compare PiT-PO against representative baselines spanning both classical and LLM-based SR methods. For the four tasks in the LLM-SR Suite, we include GPlearn, a genetic programming-based SR approach; PySR (Grayeli et al., 2024), which couples evolutionary search with symbolic simplification; uDSR (Landajuela et al., 2022), which replaces the RNN policy in DSR with a pretrained Transformer and employs neural-guided decoding; RAG-SR (Zhang et al., 2025), which incorporates structure retrieval to assist equation generation; and LLM-SR (Shojaee et al., 2025a). For the broader LLM-SRBench benchmark, we further compare against leading LLM-based SR methods, including SGA (Ma et al., 2024), which integrates LLM-driven hypothesis proposal with physics-informed parameter optimization in a bilevel search framework, and LaSR (Grayeli et al., 2024), which leverages abstract symbolic concepts distilled from prior equations to guide hybrid LLM–evolutionary generation.
Evaluation metrics. We evaluate methods using Accuracy to Tolerance and Normalized Mean Squared Error (NMSE). For a tolerance $\tau$ , we report $\mathrm{Acc}_{\mathrm{all}}(\tau)$ (Biggio et al., 2021) and $\mathrm{Acc}_{\mathrm{avg}}(\tau)$ based on relative error: $\mathrm{Acc}_{\mathrm{all}}(\tau)=\mathbbm{1}\!\bigl(\max_{1\leq i\leq N_{\mathrm{test}}}\bigl|\tfrac{\hat{y}_{i}-y_{i}}{y_{i}}\bigr|\leq\tau\bigr)$ and $\mathrm{Acc}_{\mathrm{avg}}(\tau)=\tfrac{1}{N_{\mathrm{test}}}\sum_{i=1}^{N_{\mathrm{test}}}\mathbbm{1}\!\bigl(\bigl|\tfrac{\hat{y}_{i}-y_{i}}{y_{i}}\bigr|\leq\tau\bigr)$ , where $\hat{y}_{i}$ and $y_{i}$ denote the predicted and ground-truth values at the $i$ -th test point, respectively. We additionally report $\mathrm{NMSE}=\frac{1}{N_{\mathrm{test}}}\sum_{i=1}^{N_{\mathrm{test}}}\frac{(\hat{y}_{i}-y_{i})^{2}}{\mathrm{Var}(y)}$ to assess overall numerical accuracy. We additionally adopt the Symbolic Accuracy (SA) metric (Shojaee et al., 2025b), which directly measures whether the discovered equation recovers the correct symbolic form (i.e., whether it is mathematically equivalent to the ground-truth equation up to fitted constants).
Hyperparameter Configurations. All experiments were run for 2,500 search iterations. To ensure a fair comparison, all hyperparameters related to LLM generation and search were kept consistent with the default configuration of LLM-SR. For the in-search policy optimization specific to PiT-PO, we use a learning rate of $1\times 10^{-6}$ , a group size of $G=4$ , and a multi-island setting of $N=4$ , resulting in an effective per-device batch size of $G\times N=16$ . The coefficient of the KL regularization term was set to 0.01, and the LoRA rank was set to $r=16$ . In addition, experiments were conducted on a single NVIDIA RTX 3090 using 4-bit quantized Llama-3.2-1B-Instruct, Llama-3.2-3B-Instruct, and Llama-3.1-8B-Instruct (Kassianik et al., 2025) to evaluate the training stability and performance transferability of PiT-PO across different parameter scales under constrained compute and memory budgets. More details are in Appendix A.
| GPlern uDSR PySR | 0.11 1.78 3.80 | 0.0972 0.0002 0.0003 | 0.05 0.36 7.02 | 0.2000 0.0856 0.0002 | 0.76 1.12 2.80 | 1.0023 0.5059 0.4068 | 28.43 59.15 70.60 | 0.3496 0.0639 0.0347 |
| --- | --- | --- | --- | --- | --- | --- | --- | --- |
| RAG-SR | 39.47 | 1.49e-6 | 0.43 | 0.0282 | 2.04 | 0.2754 | 76.28 | 0.0282 |
| LLM-SR (Mixtral) | 100.00 | 1.32e-11 | 99.98 | 1.18e-11 | 2.88 | 0.0596 | 71.44 | 0.0276 |
| LLM-SR (4o-mini) | 99.92 | 8.84e-12 | 99.97 | 8.70e-10 | 5.52 | 0.0453 | 85.33 | 0.0245 |
| LLM-SR (Llama-3.1-8B) | 59.58 | 1.17e-6 | 99.96 | 9.66e-10 | 4.86 | 0.0555 | 77.74 | 0.0246 |
| LLM-SR (Llama-3.2-3B) | 39.45 | 1.76e-6 | 66.34 | 6.99e-7 | 1.08 | 0.3671 | 74.78 | 0.0324 |
| LLM-SR (Llama-3.2-1B) | 3.28 | 4.47e-4 | 7.81 | 0.0002 | 1.70 | 0.5801 | 30.35 | 0.3801 |
| PiT-PO (Llama-3.1-8B) | 100.00 | 6.41e-31 | 99.99 | 2.11e-13 | 10.42 | 0.0090 | 84.45 | 0.0136 |
| PiT-PO (Llama-3.2-3B) | 100.00 | 7.58e-31 | 99.97 | 9.77e-10 | 7.01 | 0.0248 | 84.54 | 0.0156 |
| PiT-PO (Llama-3.2-1B) | 99.95 | 1.34e-11 | 99.97 | 1.70e-8 | 4.76 | 0.0240 | 76.91 | 0.1767 |
Table 1. Overall performance on LLM-SR Suite.
### 4.2. PiT-PO Demonstrates Superior Equation Discovery Capability
As evidenced in Table 1, PiT-PO establishes a new state-of-the-art on LLM-SR Suite. It consistently dominates baseline methods across all metrics, achieving the highest accuracy while maintaining the lowest NMSE in nearly all test cases. Crucially, when controlling for the LLM backbone, PiT-PO yields a substantial performance margin over LLM-SR, validating the effectiveness of our in-search policy optimization framework. Notably, PiT-PO is the only approach to successfully identify the exact ground-truth equation for the Oscillator 1. This structural precision extends to the larger-scale LLM-SRBench (Table 2), where PiT-PO achieves the highest symbolic accuracy across all categories. These results collectively demonstrate that PiT-PO not only fits data numerically but excels in uncovering the true underlying equations.
These quantitative gains are not accidental but stem from PiT-PO’s structural awareness. Analysis of the iterative trajectories of LLM-SR and PiT-PO in Appendix C.4 corroborates this conclusion: the iterations of LLM-SR remain persistently influenced by clearly incorrect terms, which violate physical meaning despite providing strong numerical fits, as well as by additional nuisance terms. Consequently, the search of LLM-SR often stagnates in a low-MSE regime without reaching the correct structure. In contrast, once PiT-PO enters the same regime, the dual constraints rapidly eliminate terms that improve fitting performance but are structurally incorrect. This behavior highlights the central advantage of the proposed dual-constraint learning signals, in which the physical constraints and token-level penalties provide data-driven signals for precise structural correction, guiding the LLM toward the true underlying equation rather than mere numerical overfitting.
<details>
<summary>x2.png Details</summary>
### Visual Description
\n
## Chart: NMSE vs. Iteration for Different Methods
### Overview
The image presents four separate line charts, each depicting the Normalized Mean Squared Error (NMSE) on a logarithmic scale against the number of iterations. Two methods, "LLM-SR" (blue) and "PIT-PO" (red), are compared across four different scenarios: Oscillation 1, Oscillation 2, E. coli Growth, and Stress-Strain. Each chart includes a shaded region representing the standard deviation around the mean NMSE for each method.
### Components/Axes
* **X-axis:** Iteration, ranging from 0 to 2500.
* **Y-axis:** NMSE (log scale), ranging from 10^-25 to 10^0 (1).
* **Legend:**
* LLM-SR (Blue line)
* PIT-PO (Red line)
* **Chart Titles:**
* Oscillation 1 (Top-Left)
* Oscillation 2 (Top-Right)
* E. coli Growth (Bottom-Left)
* Stress-Strain (Bottom-Right)
### Detailed Analysis
**Oscillation 1 (Top-Left):**
* The blue line (LLM-SR) starts at approximately 0.1 and generally slopes downward, with some fluctuations, reaching around 10^-8 by iteration 2500.
* The red line (PIT-PO) begins at approximately 10^-6 and decreases more rapidly initially, leveling off around 10^-15 by iteration 2500.
* The shaded regions indicate a relatively large standard deviation for both methods, especially at lower iteration counts.
**Oscillation 2 (Top-Right):**
* The blue line (LLM-SR) starts at approximately 0.01 and decreases steadily, reaching around 10^-6 by iteration 2500.
* The red line (PIT-PO) starts at approximately 10^-4 and decreases rapidly, reaching around 10^-11 by iteration 2500.
* The shaded regions are smaller than in Oscillation 1, suggesting less variability.
**E. coli Growth (Bottom-Left):**
* The blue line (LLM-SR) starts at approximately 1 and decreases with fluctuations, reaching around 0.2 by iteration 2500.
* The red line (PIT-PO) starts at approximately 0.5 and decreases with more pronounced fluctuations, reaching around 0.05 by iteration 2500.
* The shaded regions are substantial, indicating high variability in both methods.
**Stress-Strain (Bottom-Right):**
* The blue line (LLM-SR) starts at approximately 0.2 and decreases gradually, reaching around 0.1 by iteration 2500.
* The red line (PIT-PO) starts at approximately 0.1 and decreases more rapidly, reaching around 0.02 by iteration 2500.
* The shaded regions are moderate in size.
### Key Observations
* In all four scenarios, the PIT-PO method (red line) generally achieves lower NMSE values than the LLM-SR method (blue line), especially at higher iteration counts.
* The standard deviation (shaded regions) is significant in all scenarios, indicating that the performance of both methods can vary considerably.
* The E. coli Growth scenario exhibits the highest variability, as evidenced by the largest shaded regions.
* The rate of NMSE decrease slows down as the number of iterations increases in all scenarios, suggesting diminishing returns.
### Interpretation
The data suggests that the PIT-PO method consistently outperforms the LLM-SR method in terms of minimizing the Normalized Mean Squared Error across the four tested scenarios. This indicates that PIT-PO is a more accurate or efficient method for these particular problems. The large standard deviations highlight the sensitivity of both methods to initial conditions or other factors. The diminishing returns observed at higher iteration counts suggest that there is a point beyond which further iterations provide only marginal improvements in accuracy. The varying levels of variability across scenarios may be related to the inherent complexity or noise in the data for each scenario. The E. coli Growth scenario, with its high variability, may represent a more challenging problem to model accurately. The logarithmic scale of the Y-axis emphasizes the substantial reduction in error achieved by both methods, particularly the PIT-PO method.
</details>
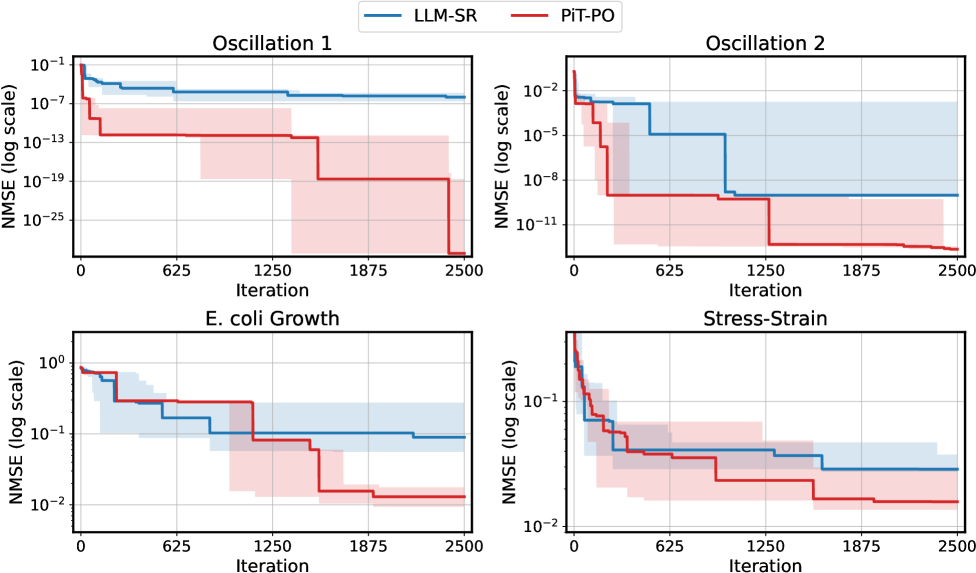
Figure 2. NMSE trajectories (log scale) over search iterations for LLM-SR and PiT-PO (Llama-3.1-8B) on LLM-SR Suite. Lines denote the median over seeds, and shaded regions indicate the min–max range.The remaining iteration curves for smaller backbones (3B and 1B) are deferred to Appendix C.1.
| Direct Prompting SGA LaSR | 3.61 2.70 5.41 | 1.801 0.909 45.94 | 0.3697 0.3519 0.0021 | 0.00 0.00 0.00 | 0.00 8.33 27.77 | 0.0644 0.0458 2.77e-4 | 0.00 0.00 4.16 | 0.00 0.00 16.66 | 0.5481 0.2416 2.73e-4 | 0.00 0.00 4.54 | 0.00 2.27 25.02 | 0.0459 0.1549 0.0018 | 0.00 0.00 8.21 | 0.00 12.12 64.22 | 0.0826 0.0435 7.44e-5 |
| --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- |
| LLM-SR | 30.63 | 38.55 | 0.0101 | 8.33 | 66.66 | 8.01e-6 | 25.30 | 58.33 | 1.04e-6 | 6.97 | 34.09 | 1.23e-4 | 4.10 | 88.12 | 1.15e-7 |
| PiT-PO | 34.23 | 46.84 | 0.0056 | 13.89 | 77.78 | 4.13e-7 | 29.17 | 70.83 | 9.37e-8 | 11.36 | 40.91 | 6.57e-5 | 12.00 | 92.00 | 1.18e-8 |
Table 2. Overall performance on LLM-SRBench (Llama-3.1-8B-Instruct).
### 4.3. PiT-PO Empowers Lightweight Backbones to Rival Large Models
As shown in Table 1, the performance of PiT-PO with the Llama-3.1-8B, Llama-3.2-3B, and Llama-3.2-1B backbones is competitive with, and often exceeds, the performance of LLM-SR that relies on substantially larger or proprietary models, including Mixtral 8 $\times$ 7B and 4o-mini.
These results indicate that PiT-PO effectively bridges the capability gap between lightweight open-source models and large-scale commercial systems. From a practical standpoint, this reduces the barrier to entry for scientific discovery: by delivering state-of-the-art performance on consumer-grade hardware (even maintaining competitiveness with a 1B backbone), PiT-PO eliminates the dependence on massive compute and closed-source APIs, thereby democratizing access to powerful SR tools.
<details>
<summary>x3.png Details</summary>
### Visual Description
\n
## Bar Chart: NMSE Comparison of Different Models
### Overview
The image presents a bar chart comparing the Normalized Mean Squared Error (NMSE) on a logarithmic scale for three different model configurations: "w/o Phy", "w/o TokenReg", and "PIT-PO". The chart distinguishes between "In-Distribution" (ID) and "Out-of-Distribution" (OOD) data.
### Components/Axes
* **X-axis:** Model Configuration - "w/o Phy", "w/o TokenReg", "PIT-PO"
* **Y-axis:** NMSE (log scale) - ranging from 10^-29 to 10^-10. The scale is logarithmic.
* **Legend:**
* ID (White bars with light gray diagonal pattern)
* OOD (Orange bars with darker orange diagonal pattern)
### Detailed Analysis
The chart displays NMSE values for both In-Distribution (ID) and Out-of-Distribution (OOD) data for each model.
* **w/o Phy:**
* ID: The blue bar representing ID data has a value of approximately 7.60e-21.
* OOD: The blue bar representing OOD data has a value of approximately 2.06e-10.
* **w/o TokenReg:**
* ID: The blue bar representing ID data has a value of approximately 2.77e-19.
* OOD: The blue bar representing OOD data has a value of approximately 9.97e-11.
* **PIT-PO:**
* ID: The orange bar representing ID data has a value of approximately 6.40e-31.
* OOD: The orange bar representing OOD data has a value of approximately 1.63e-30.
The bars for "w/o Phy" and "w/o TokenReg" are blue, indicating the ID and OOD data. The bars for "PIT-PO" are orange, indicating the ID and OOD data.
### Key Observations
* The PIT-PO model consistently exhibits the lowest NMSE values for both ID and OOD data, by several orders of magnitude.
* The NMSE values are significantly higher for OOD data compared to ID data for the "w/o Phy" and "w/o TokenReg" models.
* The difference in NMSE between ID and OOD data is less pronounced for the PIT-PO model.
### Interpretation
The data suggests that the PIT-PO model performs significantly better than the other two configurations ("w/o Phy" and "w/o TokenReg") in terms of minimizing NMSE for both in-distribution and out-of-distribution data. This indicates that the PIT-PO model is more robust and generalizes better to unseen data. The large difference in NMSE between ID and OOD data for the "w/o Phy" and "w/o TokenReg" models suggests that these models are more prone to overfitting or are less capable of handling data that deviates from the training distribution. The smaller difference for PIT-PO suggests better generalization capabilities. The logarithmic scale emphasizes the substantial differences in error rates, particularly the very low errors achieved by PIT-PO. The chart demonstrates the effectiveness of the PIT-PO approach in reducing prediction error, especially when dealing with out-of-distribution data.
</details>
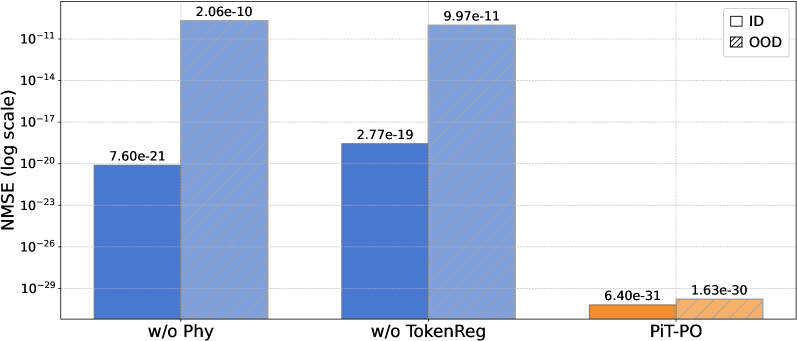
Figure 3. Ablation results of PiT-PO and its variants.
### 4.4. PiT-PO Enhances Search Efficiency and Breaks Stagnation
Figure 2 shows that PiT-PO achieves superior search efficiency in discovering accurate equations. In the early search stage, the red and blue curves are close across all four tasks: both methods primarily rely on the fitting signal (MSE) and therefore exhibit comparable per-iteration progress. As NMSE enters a lower regime, the trajectories consistently separate: PiT-PO exhibits abrupt step-wise drops while LLM-SR tends to plateau, yielding a clear red–blue gap in every subplot. Concretely, once the search reaches these lower-error regions, PiT-PO repeatedly exits stagnation and transitions to the next accuracy phase with orders-of-magnitude NMSE reductions (most prominently in Oscillation 1 and Oscillation 2, and also evident in E. coli Growth and Stress-Strain), whereas LLM-SR often remains trapped near its current error floor. This behavior confirms that the proposed dual-constraint mechanism effectively activates exactly when naive MSE feedback becomes insufficient. By penalizing physical inconsistencies and structural redundancy, PiT-PO forces the LLM to exit stagnation and transition toward the correct functional form.
While the in-search fine-tuning introduces a computational overhead, this cost is decisively outweighed by the substantial gains in performance. As detailed in Appendix C.2, PiT-PO maintains a significant performance edge even when evaluated under equivalent wall-clock time, demonstrating that the accelerated convergence speed effectively compensates for the additional training time.
### 4.5. Ablation Study
To rigorously validate the contribution of each algorithmic component, we conduct an ablation study across three settings: w/o Phy, which excludes the physics-consistency penalty $P_{\text{phy}}$ ; w/o TokenReg, which removes the redundancy-aware token-level regularization; and the full PiT-PO framework. As shown in Figure 3, removing any single component leads to a substantial deterioration of NMSE and a larger generalization gap between In-Distribution (ID) and Out-Of-Distribution (OOD) data. These empirical results underscore the necessity of the complete framework, demonstrating that the proposed dual constraints are indispensable for ensuring both search stability and robust generalization.
<details>
<summary>x4.png Details</summary>
### Visual Description
\n
## Diagram: Cyclic Boundary Condition Illustration
### Overview
The image is a diagram illustrating cyclic boundary conditions in a flow field, likely related to computational fluid dynamics or a similar simulation. It depicts the y/H versus x/H relationship for different boundary conditions (solid wall, cyclic, and varying alpha values). The diagram shows how the flow profile changes as it encounters different boundary types.
### Components/Axes
* **x-axis:** Labeled "x/H". The scale ranges from approximately 0 to 9, with markings at integer values.
* **y-axis:** Labeled "y/H". The scale ranges from approximately 0 to 3, with markings at integer values.
* **Curves:** Three distinct curves are present, each representing a different boundary condition.
* Dark Green: Represents a "solid wall" boundary condition.
* Light Brown: Represents a "solid wall" boundary condition.
* Teal/Cyan: Represents cyclic boundary conditions with varying alpha (α) values: 0.5, 0.8, and 1.0.
* **Annotations:**
* "flow direction" - A yellow rectangular box pointing to the right, indicating the direction of the flow.
* "solid wall" - Labels above and below the dark green and light brown curves.
* "cyclic" - Labels above the teal/cyan curves.
* "α = 0.5", "α = 0.8", "α = 1.0" - Labels indicating the alpha values for the cyclic boundary conditions.
* "H" - Label indicating a reference height.
* "cycliccycliccyclic" - Repeated label above the rightmost teal/cyan curves.
### Detailed Analysis or Content Details
The diagram shows the profiles of y/H as a function of x/H for different boundary conditions.
* **Solid Wall (Dark Green):** Starts at y/H ≈ 3, decreases rapidly to y/H ≈ 0 around x/H ≈ 1, and remains at y/H ≈ 0 until x/H ≈ 3.
* **Solid Wall (Light Brown):** Starts at y/H ≈ 1, decreases rapidly to y/H ≈ 0 around x/H ≈ 7, and remains at y/H ≈ 0 until x/H ≈ 9.
* **Cyclic (α = 0.5):** Starts at y/H ≈ 2.5, decreases to y/H ≈ 0 around x/H ≈ 6, and then increases again.
* **Cyclic (α = 0.8):** Starts at y/H ≈ 2.5, decreases to y/H ≈ 0 around x/H ≈ 7, and then increases again.
* **Cyclic (α = 1.0):** Starts at y/H ≈ 2.5, decreases to y/H ≈ 0 around x/H ≈ 8, and then increases again.
The cyclic boundary conditions show a phase shift as alpha increases. The curves are vertically shifted, indicating a change in the overall flow profile. The "H" label appears to indicate a reference height, possibly the height of the domain.
### Key Observations
* The solid wall boundary conditions result in a sharp drop to y/H = 0, representing a no-flow condition at the wall.
* The cyclic boundary conditions maintain continuity of the flow across the boundary, resulting in a periodic profile.
* Increasing the alpha value (α) for the cyclic boundary condition shifts the curve to the right, indicating a change in the phase of the flow.
* The repeated "cycliccycliccyclic" label suggests the cyclic nature of the boundary condition is being emphasized.
### Interpretation
This diagram illustrates the implementation of cyclic boundary conditions in a flow simulation. Cyclic boundary conditions are used when the flow is expected to repeat itself across a certain domain boundary. The alpha (α) parameter likely controls the phase shift or offset between the repeating sections of the flow. The diagram demonstrates how different alpha values affect the flow profile near the cyclic boundary. The solid wall boundary conditions serve as a contrast, showing a completely different flow behavior where the flow is stopped at the wall. The diagram is a visual aid for understanding how to apply and interpret cyclic boundary conditions in computational fluid dynamics or related fields. The diagram does not provide numerical data, but rather a qualitative illustration of the boundary conditions.
</details>
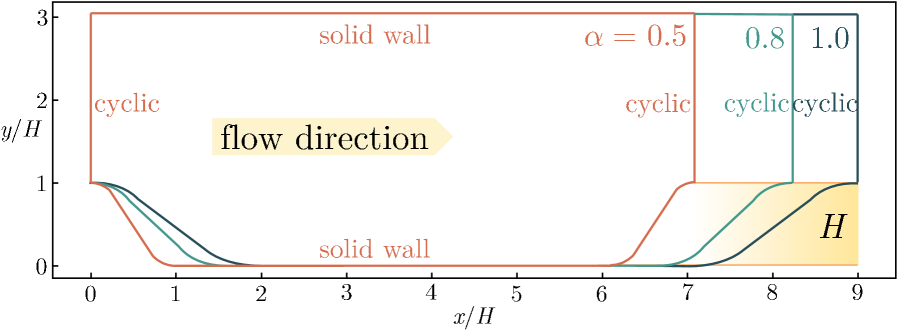
Figure 4. Schematic of the geometries for periodic hills.
### 4.6. Case Study: Turbulence Modeling
To validate the practical utility of PiT-PO in high-fidelity scientific discovery, we select the Flow over Periodic Hills (Xiao et al., 2020) (Figure 4) as a testbed. This problem is widely recognized in Computational Fluid Dynamics (CFD) (Pope, 2000) as a benchmark for Separated Turbulent Flows, presenting complex features such as strong adverse pressure gradients, massive flow detachment, and reattachment.
Problem Definition and Physics: The geometry consists of a sequence of polynomially shaped hills arranged periodically in the streamwise direction. The flow is driven by a constant body force at a bulk Reynolds number of $Re_{b}=5600$ (based on hill height $H$ and bulk velocity $U_{b}$ ). The domain height is fixed at $L_{y}/H=3.036$ , while the streamwise length $L_{x}$ varies with the slope factor $\alpha$ according to $L_{x}/H=3.858\alpha+5.142$ . Periodic boundary conditions are applied in the streamwise direction, with no-slip conditions on the walls.
The Scientific Challenge: The challenge lies in the Separation Bubble (Pope, 2000), a region where turbulence exhibits strong anisotropy due to streamline curvature. Traditional Linear Eddy Viscosity Models (LEVM) (Pope, 2000), such as the $k$ - $\omega$ SST model (Menter, 1994; Menter et al., 2003), rely on the Boussinesq hypothesis which assumes isotropic turbulence. Consequently, they systematically fail to predict key flow features, such as the separation bubble size and reattachment location.
Discovery Objective: Instead of fitting a simple curve, our goal is to discover a Non-linear Constitutive Relation for the Reynolds stress anisotropy tensor $a_{ij}$ and the dimensionless Reynolds stress anisotropy tensor $b_{ij}$ . By learning the Reynolds stress tensor $\tau_{ij}$ from high-fidelity Direct Numerical Simulation (DNS) (Pope, 2000) data, PiT-PO aims to formulate a symbolic correction term that captures the anisotropic physics missed by linear models.
Baselines: We follow standard turbulence modeling protocols and compare primarily against the standard $k$ - $\omega$ SST model of RANS. We also include LLM-SR and DSRRANS (Tang et al., 2023a), a strong SR-based turbulence modeling method specifically designed for turbulence tasks.
<details>
<summary>x5.png Details</summary>
### Visual Description
## Heatmaps: Reynolds Stress Components in Wall-Bounded Turbulence
### Overview
The image presents a 5x4 grid of heatmaps, visualizing Reynolds stress components normalized by the square of the friction velocity (uτ²). Each row represents a different turbulence model or dataset (DNS, DSRRANS, LLM-SR, PIT-PO), and each column represents a different Reynolds stress component (a₁₁, a₂₂, a₁₂, a₂₁). The heatmaps display the normalized Reynolds stress components as a function of wall-normal distance (y⁺/h) and streamwise distance (x/H).
### Components/Axes
* **Rows:** DNS (Direct Numerical Simulation), DSRRANS (Dynamic Subgrid Reynolds-Averaged Navier-Stokes), LLM-SR (Large Eddy Model - Scale-Resolved), PIT-PO (Piterbarg-Type One-Equation Model)
* **Columns:** a₁₁/uτ², a₂₂/uτ², a₁₂/uτ², a₂₁/uτ² (Reynolds stress components)
* **X-axis:** x/H (Streamwise distance, ranging from approximately 0 to 7.5)
* **Y-axis:** y⁺/h (Wall-normal distance, ranging from approximately 0 to 2.5)
* **Color Scale:** Ranges from -0.04 to 0.03, with blue representing negative values and red representing positive values. The colorbar increments are 0.004.
### Detailed Analysis or Content Details
**Row 1: DNS**
* **a₁₁/uτ²:** The heatmap shows a generally positive trend, with higher values near the wall (y⁺/h ≈ 0) and decreasing values as y⁺/h increases. Values range from approximately -0.01 to 0.02. A slight undulation is visible along the x/H axis.
* **a₂₂/uτ²:** Similar to a₁₁, this heatmap shows positive values near the wall, decreasing with increasing y⁺/h. Values range from approximately -0.02 to 0.01.
* **a₁₂/uτ²:** Displays a strong negative region near the wall, transitioning to a positive region further away from the wall. Values range from approximately -0.03 to 0.01.
* **a₂₁/uτ²:** Mirrors the pattern of a₁₂ with a strong positive region near the wall and a negative region further away. Values range from approximately -0.02 to 0.02.
**Row 2: DSRRANS**
* **a₁₁/uτ²:** Shows a broad positive region, but with less pronounced variation than DNS. Values range from approximately -0.01 to 0.02.
* **a₂₂/uτ²:** Similar to a₁₁, with a broad positive region. Values range from approximately -0.01 to 0.02.
* **a₁₂/uτ²:** Displays a negative region near the wall, but less defined than in DNS. Values range from approximately -0.02 to 0.01.
* **a₂₁/uτ²:** Mirrors a₁₂, but with less definition. Values range from approximately -0.02 to 0.01.
**Row 3: LLM-SR**
* **a₁₁/uτ²:** Shows a strong positive region near the wall, with a more pronounced peak than DNS. Values range from approximately -0.01 to 0.03.
* **a₂₂/uτ²:** Similar to a₁₁, with a strong positive region. Values range from approximately -0.01 to 0.03.
* **a₁₂/uτ²:** Displays a clear negative region near the wall, transitioning to positive values. Values range from approximately -0.03 to 0.02.
* **a₂₁/uτ²:** Mirrors a₁₂, with a clear negative region near the wall. Values range from approximately -0.03 to 0.02.
**Row 4: PIT-PO**
* **a₁₁/uτ²:** Shows a positive region near the wall, but with less variation than DNS. Values range from approximately -0.01 to 0.02.
* **a₂₂/uτ²:** Similar to a₁₁, with a positive region. Values range from approximately -0.01 to 0.02.
* **a₁₂/uτ²:** Displays a negative region near the wall, but less defined than in DNS. Values range from approximately -0.02 to 0.01.
* **a₂₁/uτ²:** Mirrors a₁₂, but with less definition. Values range from approximately -0.02 to 0.01.
**Row 5: DNS**
* **a₁₁/uτ²:** The heatmap shows a generally positive trend, with higher values near the wall (y⁺/h ≈ 0) and decreasing values as y⁺/h increases. Values range from approximately -0.01 to 0.02. A slight undulation is visible along the x/H axis.
* **a₂₂/uτ²:** Similar to a₁₁, this heatmap shows positive values near the wall, decreasing with increasing y⁺/h. Values range from approximately -0.02 to 0.01.
* **a₁₂/uτ²:** Displays a strong negative region near the wall, transitioning to a positive region further away from the wall. Values range from approximately -0.03 to 0.01.
* **a₂₁/uτ²:** Mirrors the pattern of a₁₂ with a strong positive region near the wall and a negative region further away. Values range from approximately -0.02 to 0.02.
### Key Observations
* DNS consistently shows the most detailed and complex patterns in the Reynolds stress components.
* DSRRANS and PIT-PO exhibit smoother distributions, suggesting a loss of fine-scale information.
* LLM-SR captures some of the complexity of DNS, particularly in the near-wall region.
* The a₁₂ and a₂₁ components consistently show a sign change with increasing wall-normal distance.
* The magnitude of the Reynolds stresses generally decreases with increasing wall-normal distance.
### Interpretation
The heatmaps illustrate the differences in how various turbulence models represent Reynolds stresses in wall-bounded turbulent flows. DNS, being the most computationally expensive but accurate method, serves as a benchmark. The other models (DSRRANS, LLM-SR, and PIT-PO) attempt to approximate the DNS results with varying degrees of fidelity and computational cost.
The smoother distributions observed in DSRRANS and PIT-PO suggest that these models may not fully resolve the small-scale turbulent structures that contribute significantly to Reynolds stresses. LLM-SR, as a scale-resolved model, appears to capture more of the DNS complexity, particularly in the near-wall region where the Reynolds stresses are highest.
The consistent sign change in a₁₂ and a₂₁ indicates the transfer of momentum between the different flow directions. The magnitude of these stresses reflects the intensity of this momentum transfer. The differences in the magnitude and distribution of these stresses between the models highlight the challenges in accurately modeling turbulent flows and the importance of choosing an appropriate model based on the specific application and available computational resources. The repeated DNS row suggests a possible error in the image or a deliberate comparison of two DNS runs.
</details>
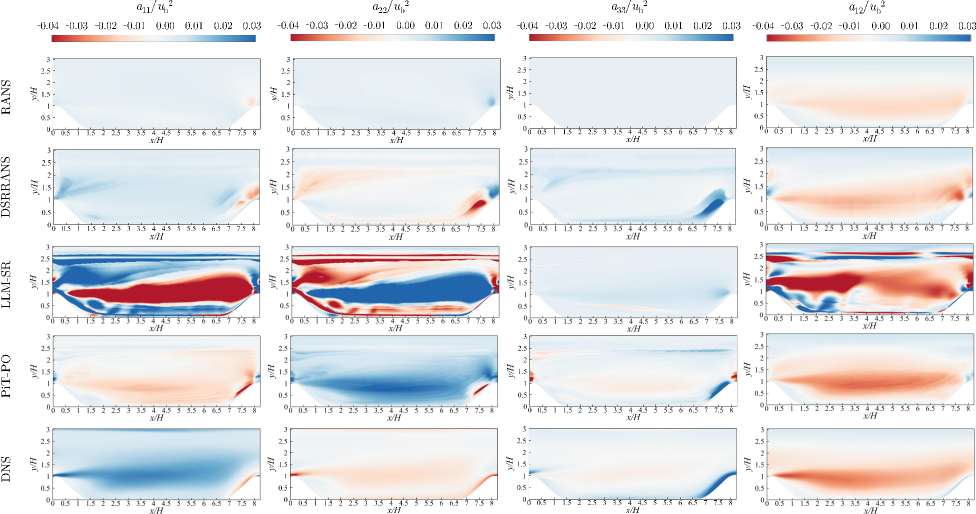
Figure 5. Comparison of the four anisotropic Reynolds stress components for periodic hill training flow using RANS, DSRRANS, LLM-SR, PiT-PO and DNS, respectively.
We cast turbulence closure modeling as a SR problem (Tang et al., 2023b) (see Appendix E for details). After obtaining the final symbolic equation, we embed it into a RANS solver of OpenFOAM (Weller et al., 1998) and run CFD simulations on the periodic-hill configuration. We compare the resulting Reynolds-stress components, mean-velocity fields, and skin-friction profiles against DNS references. Figures 5 – 7 visualize these quantities, enabling a direct assessment of physical fidelity and flow-field prediction quality.
Based on the comparative analysis of the anisotropic Reynolds stress contours (Figure 5), DSRRANS and PiT-PO show enhancement over the traditional RANS approach. Among them, PiT-PO performs the best: its contour matches the DNS reference most closely, with reduced error compared to DSRRANS and LLM-SR, demonstrating less severe non-physical extremes.
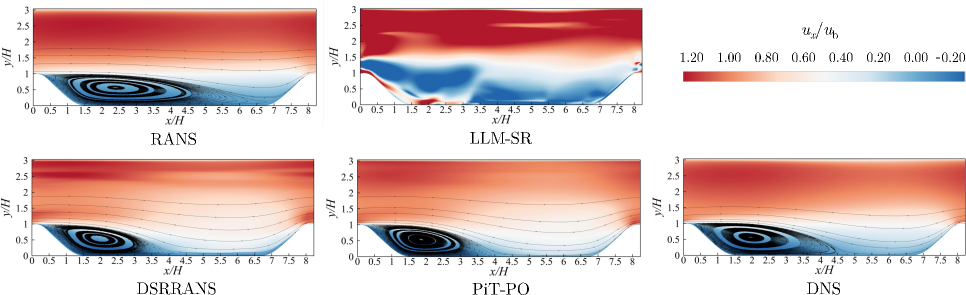
The stream-wise velocity contours illustrate the correction of the bubble size, a region of reversed flow that forms when fluid detaches from a surface. In Figure 6, PiT-PO most accurately represents the extent and shape of the recirculation zone, where fluid circulates within the separated region, closely consistent with the DNS data throughout the domain, particularly within the separation region and the recovery layer, where flow re-attaches to the surface.
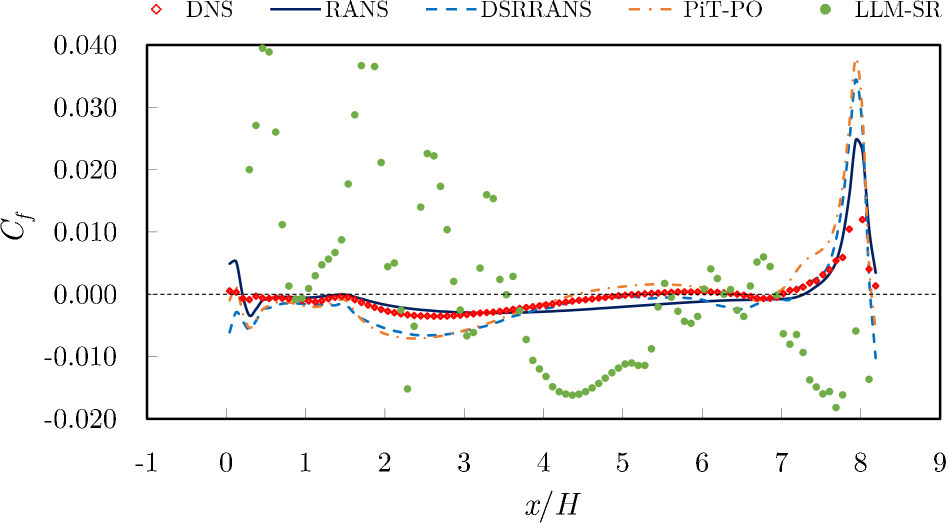
The skin friction coefficient (Figure 7), defined as the ratio of the wall stress to the dynamic pressure of the flow along the bottom wall, is a sensitive metric for predicting flow separation. The $k$ - $\omega$ SST model of RANS underestimates the magnitude of the skin friction and predicts a delayed reattachment location compared to the DNS. The learned model (PiT-PO) improves the prediction, aligning more closely with the DNS profile.
These results demonstrate that PiT-PO can generate symbolic equations tailored to turbulence modeling and that, under a posteriori CFD evaluation, the resulting predictions more closely match DNS references, which increases the practical value of LLM-based SR in real scientific and engineering workflows. With the proposed dual constraints,
PiT-PO provides targeted search and learning signals that enable the internalization of turbulence priors during equation discovery, thereby steering the model toward physically consistent and domain-relevant structures.
<details>
<summary>x6.png Details</summary>
### Visual Description
## Heatmaps: Flow Field Comparisons
### Overview
The image presents a 2x3 grid of heatmaps, each representing a different computational model of a flow field. Each heatmap visualizes the normalized streamwise velocity (u<sub>x</sub>/u<sub>b</sub>) as a function of normalized streamwise (x/H) and vertical (y/H) coordinates. The models compared are RANS, LLM-SR, DSBRANS, PIT-PO, and DNS. A color scale is provided in the top-right corner to interpret the velocity values.
### Components/Axes
* **x/H Axis:** Represents the normalized streamwise coordinate, ranging from 0 to 8, with tick marks at intervals of 0.5.
* **y/H Axis:** Represents the normalized vertical coordinate, ranging from 0 to 3, with tick marks at intervals of 0.5.
* **Color Scale:** Represents the normalized streamwise velocity (u<sub>x</sub>/u<sub>b</sub>). The scale ranges from -0.20 (dark blue) to 1.20 (dark red), with intermediate values of -0.02, 0.20, 0.40, 0.60, 0.80, and 1.00.
* **Labels:** Each heatmap is labeled with the corresponding model name: RANS, LLM-SR, DSBRANS, PIT-PO, and DNS.
* **Legend:** Located in the top-right corner, the legend maps colors to u<sub>x</sub>/u<sub>b</sub> values.
### Detailed Analysis or Content Details
Each heatmap displays a similar flow pattern, characterized by a recirculation zone near x/H = 1.0 and a region of accelerated flow further downstream. The color intensity indicates the magnitude of the normalized streamwise velocity.
**RANS:**
* The recirculation zone is visible as a dark blue region.
* The maximum positive velocity (dark red) is observed around x/H = 6.5, y/H = 1.5. The value is approximately 1.10.
* The velocity gradient is relatively smooth.
**LLM-SR:**
* The recirculation zone is more pronounced and extends further downstream compared to RANS.
* The maximum positive velocity is around x/H = 7.0, y/H = 1.5, with a value of approximately 1.15.
* There is a sharper velocity gradient near the top boundary (y/H = 3).
**DSBRANS:**
* The recirculation zone is similar in size to RANS.
* The maximum positive velocity is around x/H = 6.5, y/H = 1.5, with a value of approximately 1.10.
* The velocity gradient is smoother than LLM-SR.
**PIT-PO:**
* The recirculation zone is well-defined and similar to LLM-SR.
* The maximum positive velocity is around x/H = 7.0, y/H = 1.5, with a value of approximately 1.15.
* The velocity gradient is relatively sharp.
**DNS:**
* The recirculation zone is similar to LLM-SR and PIT-PO.
* The maximum positive velocity is around x/H = 7.0, y/H = 1.5, with a value of approximately 1.15.
* The velocity gradient is sharp, with fine-scale structures visible.
### Key Observations
* All models capture the basic flow features, but differ in the size and intensity of the recirculation zone and the sharpness of the velocity gradients.
* RANS appears to underestimate the size of the recirculation zone.
* LLM-SR, PIT-PO, and DNS show similar flow structures, with a more pronounced recirculation zone and sharper gradients.
* The maximum normalized velocity is approximately consistent across all models, around 1.10-1.15.
### Interpretation
The heatmaps compare the performance of different turbulence models in simulating a flow field. The DNS result is often considered the "ground truth" for such simulations. The comparison suggests that RANS is the least accurate model, underpredicting the recirculation zone. LLM-SR, PIT-PO, and DNS provide more realistic representations of the flow, capturing the complex flow structures with greater fidelity. The differences in velocity gradients highlight the ability of higher-fidelity models to resolve smaller-scale flow features. The consistent maximum velocity across models suggests that the overall momentum balance is reasonably well-captured by all models, but the distribution of velocity is significantly different. This data is valuable for assessing the suitability of different turbulence models for specific flow applications.
</details>
Figure 6. Non-dimensional stream-wise velocity contours obtained by the learned model and the standard $k$ - $\omega$ SST model of RANS, compared with DNS data.
## 5. Related Work
Traditional SR has been studied through several lines, including genetic programming , reinforcement learning (Petersen et al., 2021), and transformer-based generation (Biggio* et al., 2021). Genetic programming (Koza, 1990) casts equation discovery as an evolutionary search over tree-structured programs, where candidate expressions are iteratively refined via mutation and crossover. Reinforcement learning-based SR, introduced by Petersen et al. (Petersen et al., 2021), has developed into a family of policy-optimization frameworks (Mundhenk et al., 2021; Landajuela et al., 2021; Crochepierre et al., 2022; Du et al., 2023) that formulate SR as a sequential decision-making process. More recently, transformer-based models (Valipour et al., 2021; Vastl et al., 2024; Kamienny et al., 2022; Li et al., 2023; Zhang et al., 2025) have been adopted for SR, using large-scale pretraining to map numerical data directly to equations. However, these methods typically fail to incorporate scientific prior knowledge.
Recent progress in natural language processing has further enabled LLM-based SR methods, including LLM-SR (Shojaee et al., 2025a), LaSR (Grayeli et al., 2024), ICSR (Merler et al., 2024), CoEvo (Guo et al., 2025), and SR-Scientist (Xia et al., 2025). LLM-SR exploits scientific priors that are implicitly captured by LLMs to propose plausible functional forms, followed by data-driven parameter estimation. LaSR augments SR with abstract concept generation to guide hypothesis formation, while ICSR reformulates training examples as in-context prompts to elicit function generation. However, a unifying limitation across these methods is their reliance on the LLM as a frozen generator, which precludes incorporating search feedback to update the generation strategy and consequently restricts their ability to adapt to complex problems.
While some recent works, such as SOAR (Pourcel et al., 2025) and CALM (Huang et al., 2025), have begun to explore adaptive in-search tuning, they primarily focus on algorithm discovery or combinatorial optimization problems, whereas our method is specifically tailored for SR. By integrating hierarchical physical constraints and theorem-guided token regularization, PiT-PO establishes an adaptive framework capable of discovering accurate and physically consistent equations.
<details>
<summary>x7.png Details</summary>
### Visual Description
\n
## Chart: Skin Friction Coefficient vs. Normalized Distance
### Overview
The image presents a line chart comparing the skin friction coefficient (Cf) against the normalized distance (x/H) for several turbulence models. The chart displays data from Direct Numerical Simulation (DNS), Reynolds-Averaged Navier-Stokes (RANS), Delayed Detached Eddy Simulation (DSRRANS), Pit-PO, and Large Eddy Simulation (LLM-SR). The DNS data is represented as scattered points, while the other models are shown as lines.
### Components/Axes
* **X-axis:** x/H (Normalized Distance). Scale ranges from approximately -1 to 9.
* **Y-axis:** Cf (Skin Friction Coefficient). Scale ranges from approximately -0.020 to 0.040.
* **Legend:** Located at the top of the chart, identifying each data series with a specific color and line style.
* DNS (Red Diamonds)
* RANS (Solid Black Line)
* DSRRANS (Blue Dashed Line)
* PiT-PO (Orange Dotted Line)
* LLM-SR (Green Circles)
### Detailed Analysis
* **DNS (Red Diamonds):** The DNS data shows significant scatter, with Cf values ranging from approximately -0.015 to 0.035. There's a general trend of increasing Cf as x/H approaches 9, with a peak around x/H = 8.5.
* **RANS (Solid Black Line):** The RANS line starts at approximately 0.002 at x/H = -1, dips to around -0.005 at x/H = 0, remains relatively flat around -0.002 to 0.001 from x/H = 1 to 7, and then rises sharply to approximately 0.015 at x/H = 9.
* **DSRRANS (Blue Dashed Line):** The DSRRANS line begins at approximately 0.005 at x/H = -1, dips to around -0.010 at x/H = 0, fluctuates between -0.010 and 0.005 from x/H = 1 to 7, and then increases rapidly to approximately 0.035 at x/H = 9.
* **PiT-PO (Orange Dotted Line):** The PiT-PO line starts at approximately 0.001 at x/H = -1, remains relatively flat around 0.000 to 0.002 from x/H = 0 to 7, and then increases to approximately 0.010 at x/H = 9.
* **LLM-SR (Green Circles):** The LLM-SR data shows significant scatter, with Cf values ranging from approximately -0.020 to 0.030. There's a general trend of increasing Cf as x/H approaches 9, with a peak around x/H = 8.
### Key Observations
* The DNS data exhibits the highest degree of variability, indicating the complex nature of the turbulent flow.
* RANS provides the smoothest and most conservative estimate of Cf, consistently underpredicting the values observed in DNS.
* DSRRANS shows a more dynamic behavior than RANS, capturing some of the fluctuations seen in DNS, but still smoothing out much of the detail.
* PiT-PO provides a relatively stable prediction of Cf, similar to RANS but with slightly higher values.
* LLM-SR shows a similar trend to DNS, but with less scatter.
* All models show a significant increase in Cf as x/H approaches 9, suggesting a region of increased shear stress.
### Interpretation
The chart compares the performance of different turbulence models in predicting the skin friction coefficient along a surface. The DNS data serves as a benchmark, representing the most accurate (but computationally expensive) solution. The other models offer varying levels of accuracy and computational cost.
The significant scatter in the DNS data highlights the inherent complexity of turbulent flows. The RANS model, being the simplest, provides a smoothed-out representation that may be suitable for engineering applications where accuracy is less critical. DSRRANS and PiT-PO offer improvements over RANS by capturing some of the turbulent fluctuations, while LLM-SR appears to provide a good balance between accuracy and computational cost.
The sharp increase in Cf near x/H = 9 suggests a region of flow separation or adverse pressure gradient, where the shear stress at the wall is significantly increased. The differences in how each model captures this phenomenon indicate their varying abilities to resolve complex flow features. The divergence between the models at higher x/H values suggests that the choice of turbulence model can significantly impact the prediction of skin friction drag in this region.
</details>
Figure 7. Skin friction distribution along the bottom obtained by the learned model and the standard $k$ - $\omega$ SST model of RANS, compared with DNS data.
## 6. Conclusion
In this work, we introduced PiT-PO, a unified framework that fundamentally transforms LLMs from static equation proposers into adaptive, physics-aware generators for SR. By integrating in-search policy optimization with a novel dual-constraint evaluation mechanism, PiT-PO rigorously enforces hierarchical physical validity while leveraging theorem-guided, token-level penalties to eliminate structural redundancy. This synergistic design aligns generation with numerical fitness, scientific consistency, and parsimony, establishing new state-of-the-art performance on SR benchmarks. Beyond synthetic tasks, PiT-PO demonstrates significant practical utility in turbulence modeling, where the discovered symbolic corrections improve Reynolds stress and flow-field predictions. Notably, PiT-PO achieves these results using small open-source backbones, making it a practical and accessible tool for scientific communities with limited computational resources. Looking forward, we plan to extend PiT-PO to broader scientific and engineering domains by enriching the library of domain-specific constraints and validating it across more complex, real-world systems. Moreover, we anticipate that integrating PiT-PO with larger-scale multi-modal foundation models could further unlock its potential in processing heterogeneous scientific data.
## References
- (1)
- Aakash et al. (2019) B. S. Aakash, JohnPatrick Connors, and Michael D Shields. 2019. Stress-strain data for aluminum 6061-T651 from 9 lots at 6 temperatures under uniaxial and plane strain tension. Data in Brief 25 (Aug 2019), 104085. doi: 10.1016/j.dib.2019.104085
- Biggio et al. (2021) Luca Biggio, Tommaso Bendinelli, Alexander Neitz, Aurelien Lucchi, and Giambattista Parascandolo. 2021. Neural Symbolic Regression that Scales. arXiv:2106.06427 [cs.LG] https://arxiv.org/abs/2106.06427
- Biggio* et al. (2021) L. Biggio*, T. Bendinelli*, A. Neitz, A. Lucchi, and G. Parascandolo. 2021. Neural Symbolic Regression that Scales. In Proceedings of 38th International Conference on Machine Learning (ICML 2021) (Proceedings of Machine Learning Research, Vol. 139). PMLR, 936–945. https://proceedings.mlr.press/v139/biggio21a.html *equal contribution.
- Chen et al. (2025) Jindou Chen, Jidong Tian, Liang Wu, ChenXinWei, Xiaokang Yang, Yaohui Jin, and Yanyan Xu. 2025. KinFormer: Generalizable Dynamical Symbolic Regression for Catalytic Organic Reaction Kinetics. In International Conference on Representation Learning, Y. Yue, A. Garg, N. Peng, F. Sha, and R. Yu (Eds.), Vol. 2025. 67058–67080. https://proceedings.iclr.cc/paper_files/paper/2025/file/a76b693f36916a5ed84d6e5b39a0dc03-Paper-Conference.pdf
- Cranmer (2023) Miles Cranmer. 2023. Interpretable Machine Learning for Science with PySR and SymbolicRegression.jl. arXiv:2305.01582 [astro-ph.IM] https://arxiv.org/abs/2305.01582
- Crochepierre et al. (2022) Laure Crochepierre, Lydia Boudjeloud-Assala, and Vincent Barbesant. 2022. Interactive Reinforcement Learning for Symbolic Regression from Multi-Format Human-Preference Feedbacks. In IJCAI 2022- 31st International Joint Conference on Artificial Intelligence. Vienne, Austria. https://hal.science/hal-03695471
- Deng et al. (2023) Song Deng, Junjie Wang, Li Tao, Su Zhang, and Hongwei Sun. 2023. EV charging load forecasting model mining algorithm based on hybrid intelligence. Computers and Electrical Engineering 112 (2023), 109010. doi: 10.1016/j.compeleceng.2023.109010
- Du et al. (2023) Mengge Du, Yuntian Chen, and Dongxiao Zhang. 2023. DISCOVER: Deep identification of symbolically concise open-form PDEs via enhanced reinforcement-learning. arXiv:2210.02181 [cs.LG] https://arxiv.org/abs/2210.02181
- Fletcher (1987) Roger Fletcher. 1987. Practical Methods of Optimization (2nd ed.). John Wiley & Sons, Chichester, New York.
- Grayeli et al. (2024) Arya Grayeli, Atharva Sehgal, Omar Costilla-Reyes, Miles Cranmer, and Swarat Chaudhuri. 2024. Symbolic Regression with a Learned Concept Library. arXiv:2409.09359 [cs.LG] https://arxiv.org/abs/2409.09359
- Guo et al. (2025) Ping Guo, Qingfu Zhang, and Xi Lin. 2025. CoEvo: Continual Evolution of Symbolic Solutions Using Large Language Models. arXiv:2412.18890 [cs.AI] https://arxiv.org/abs/2412.18890
- Hu et al. (2021) Edward J. Hu, Yelong Shen, Phillip Wallis, Zeyuan Allen-Zhu, Yuanzhi Li, Shean Wang, Lu Wang, and Weizhu Chen. 2021. LoRA: Low-Rank Adaptation of Large Language Models. arXiv:2106.09685 [cs.CL] https://arxiv.org/abs/2106.09685
- Huang et al. (2025) Ziyao Huang, Weiwei Wu, Kui Wu, Jianping Wang, and Wei-Bin Lee. 2025. CALM: Co-evolution of Algorithms and Language Model for Automatic Heuristic Design. arXiv:2505.12285 [cs.NE] https://arxiv.org/abs/2505.12285
- Kamienny et al. (2022) Pierre-Alexandre Kamienny, Stéphane d’Ascoli, Guillaume Lample, and François Charton. 2022. End-to-end symbolic regression with transformers. arXiv:2204.10532 [cs.LG] https://arxiv.org/abs/2204.10532
- Kassianik et al. (2025) Paul Kassianik, Baturay Saglam, Alexander Chen, Blaine Nelson, Anu Vellore, Massimo Aufiero, Fraser Burch, Dhruv Kedia, Avi Zohary, Sajana Weerawardhena, Aman Priyanshu, Adam Swanda, Amy Chang, Hyrum Anderson, Kojin Oshiba, Omar Santos, Yaron Singer, and Amin Karbasi. 2025. Llama-3.1-FoundationAI-SecurityLLM-Base-8B Technical Report. arXiv:2504.21039 [cs.CR] https://arxiv.org/abs/2504.21039
- Koza (1990) J.R. Koza. 1990. Genetically breeding populations of computer programs to solve problems in artificial intelligence. In [1990] Proceedings of the 2nd International IEEE Conference on Tools for Artificial Intelligence. 819–827. doi: 10.1109/TAI.1990.130444
- Landajuela et al. (2022) Mikel Landajuela, Chak Shing Lee, Jiachen Yang, Ruben Glatt, Claudio P Santiago, Ignacio Aravena, Terrell Mundhenk, Garrett Mulcahy, and Brenden K Petersen. 2022. A Unified Framework for Deep Symbolic Regression. In Advances in Neural Information Processing Systems, S. Koyejo, S. Mohamed, A. Agarwal, D. Belgrave, K. Cho, and A. Oh (Eds.), Vol. 35. Curran Associates, Inc., 33985–33998. https://proceedings.neurips.cc/paper_files/paper/2022/file/dbca58f35bddc6e4003b2dd80e42f838-Paper-Conference.pdf
- Landajuela et al. (2021) Mikel Landajuela, Brenden K. Petersen, Soo K. Kim, Claudio P. Santiago, Ruben Glatt, T. Nathan Mundhenk, Jacob F. Pettit, and Daniel M. Faissol. 2021. Improving exploration in policy gradient search: Application to symbolic optimization. arXiv:2107.09158 [cs.LG] https://arxiv.org/abs/2107.09158
- Li et al. (2023) Wenqiang Li, Weijun Li, Linjun Sun, Min Wu, Lina Yu, Jingyi Liu, Yanjie Li, and Song Tian. 2023. Transformer-based model for symbolic regression via joint supervised learning. In International Conference on Learning Representations. https://api.semanticscholar.org/CorpusID:259298765
- Ma et al. (2024) Pingchuan Ma, Tsun-Hsuan Wang, Minghao Guo, Zhiqing Sun, Joshua B. Tenenbaum, Daniela Rus, Chuang Gan, and Wojciech Matusik. 2024. LLM and Simulation as Bilevel Optimizers: A New Paradigm to Advance Physical Scientific Discovery. In Proceedings of the 41st International Conference on Machine Learning (Proceedings of Machine Learning Research, Vol. 235), Ruslan Salakhutdinov, Zico Kolter, Katherine Heller, Adrian Weller, Nuria Oliver, Jonathan Scarlett, and Felix Berkenkamp (Eds.). PMLR, 33940–33962. https://proceedings.mlr.press/v235/ma24m.html
- Makke and Chawla (2024a) Nour Makke and Sanjay Chawla. 2024a. Data-driven discovery of Tsallis-like distribution using symbolic regression in high-energy physics. PNAS Nexus 3, 11 (10 2024), pgae467. arXiv:https://academic.oup.com/pnasnexus/article-pdf/3/11/pgae467/60816181/pgae467.pdf doi: 10.1093/pnasnexus/pgae467
- Makke and Chawla (2024b) Nour Makke and Sanjay Chawla. 2024b. Interpretable scientific discovery with symbolic regression: a review. Artificial Intelligence Review 57 (01 2024). doi: 10.1007/s10462-023-10622-0
- Menter (1994) Florian R Menter. 1994. Two-equation eddy-viscosity turbulence models for engineering applications. AIAA journal 32, 8 (1994), 1598–1605.
- Menter et al. (2003) Florian R Menter, Martin Kuntz, Robin Langtry, et al. 2003. Ten years of industrial experience with the SST turbulence model. Turbulence, heat and mass transfer 4, 1 (2003), 625–632.
- Merler et al. (2024) Matteo Merler, Katsiaryna Haitsiukevich, Nicola Dainese, and Pekka Marttinen. 2024. In-Context Symbolic Regression: Leveraging Large Language Models for Function Discovery. In Proceedings of the 62nd Annual Meeting of the Association for Computational Linguistics (Volume 4: Student Research Workshop). Association for Computational Linguistics, 589–606. doi: 10.18653/v1/2024.acl-srw.49
- MOCHIZUKI and OSAKA (2000) Shinsuke MOCHIZUKI and Hideo OSAKA. 2000. Management of a Stronger Wall Jet by a Pair of Streamwise Vortices. Reynolds Stress Tensor and Production Tenns. TRANSACTIONS OF THE JAPAN SOCIETY OF MECHANICAL ENGINEERS Series B 66, 646 (2000), 1309–1317. doi: 10.1299/kikaib.66.646_1309
- Monkewitz (2021) Peter A. Monkewitz. 2021. Asymptotics of streamwise Reynolds stress in wall turbulence. Journal of Fluid Mechanics 931 (Nov. 2021). doi: 10.1017/jfm.2021.924
- Monod (1949) Jacques Monod. 1949. THE GROWTH OF BACTERIAL CULTURES. Annual Review of Microbiology 3, Volume 3, 1949 (1949), 371–394. doi: 10.1146/annurev.mi.03.100149.002103
- Mundhenk et al. (2021) T. Nathan Mundhenk, Mikel Landajuela, Ruben Glatt, Claudio P. Santiago, Daniel M. Faissol, and Brenden K. Petersen. 2021. Symbolic Regression via Neural-Guided Genetic Programming Population Seeding. arXiv:2111.00053 [cs.NE] https://arxiv.org/abs/2111.00053
- Neamtiu et al. (2005) Iulian Neamtiu, Jeffrey S. Foster, and Michael Hicks. 2005. Understanding source code evolution using abstract syntax tree matching. In Proceedings of the 2005 International Workshop on Mining Software Repositories (St. Louis, Missouri) (MSR ’05). Association for Computing Machinery, New York, NY, USA, 1–5. doi: 10.1145/1083142.1083143
- Petersen et al. (2021) Brenden K. Petersen, Mikel Landajuela, T. Nathan Mundhenk, Claudio P. Santiago, Soo K. Kim, and Joanne T. Kim. 2021. Deep symbolic regression: Recovering mathematical expressions from data via risk-seeking policy gradients. arXiv:1912.04871 [cs.LG] https://arxiv.org/abs/1912.04871
- Pope (2000) Stephen B. Pope. 2000. Turbulent Flows. Cambridge University Press. doi: 10.1017/cbo9780511840531
- Pourcel et al. (2025) Julien Pourcel, Cédric Colas, and Pierre-Yves Oudeyer. 2025. Self-Improving Language Models for Evolutionary Program Synthesis: A Case Study on ARC-AGI. arXiv:2507.14172 [cs.LG] https://arxiv.org/abs/2507.14172
- Reuter et al. (2023) Julia Reuter, Hani Elmestikawy, Fabien Evrard, Sanaz Mostaghim, and Berend van Wachem. 2023. Graph Networks as Inductive Bias for Genetic Programming: Symbolic Models for Particle-Laden Flows. In Genetic Programming, Gisele Pappa, Mario Giacobini, and Zdenek Vasicek (Eds.). Springer Nature Switzerland, Cham, 36–51.
- Romera-Paredes et al. (2024) Bernardino Romera-Paredes, Mohammadamin Barekatain, Alexander Novikov, Matej Balog, M Pawan Kumar, Emilien Dupont, Francisco JR Ruiz, Jordan S Ellenberg, Pengming Wang, Omar Fawzi, et al. 2024. Mathematical discoveries from program search with large language models. Nature 625, 7995 (2024), 468–475.
- Rosso et al. (1995) L Rosso, J. R. Lobry, S Bajard, and J. P. Flandrois. 1995. Convenient Model To Describe the Combined Effects of Temperature and pH on Microbial Growth. Applied and Environmental Microbiology 61, 2 (Feb 1995), 610–6. doi: 10.1128/aem.61.2.610-616.1995
- Schmidt and Lipson (2009) Michael Schmidt and Hod Lipson. 2009. Distilling Free-Form Natural Laws from Experimental Data. Science 324, 5923 (2009), 81–85. arXiv:https://www.science.org/doi/pdf/10.1126/science.1165893 doi: 10.1126/science.1165893
- Shao et al. (2024) Zhihong Shao, Peiyi Wang, Qihao Zhu, Runxin Xu, Junxiao Song, Xiao Bi, Haowei Zhang, Mingchuan Zhang, Y. K. Li, Y. Wu, and Daya Guo. 2024. DeepSeekMath: Pushing the Limits of Mathematical Reasoning in Open Language Models. arXiv:2402.03300 [cs.CL] https://arxiv.org/abs/2402.03300
- Shi et al. (2024) Hao Shi, Weili Song, Xinting Zhang, Jiahe Shi, Cuicui Luo, Xiang Ao, Hamid Arian, and Luis Seco. 2024. AlphaForge: A Framework to Mine and Dynamically Combine Formulaic Alpha Factors. arXiv:2406.18394 [q-fin.CP] https://arxiv.org/abs/2406.18394
- Shojaee et al. (2025a) Parshin Shojaee, Kazem Meidani, Shashank Gupta, Amir Barati Farimani, and Chandan K Reddy. 2025a. LLM-SR: Scientific Equation Discovery via Programming with Large Language Models. arXiv:2404.18400 [cs.LG] https://arxiv.org/abs/2404.18400
- Shojaee et al. (2025b) Parshin Shojaee, Ngoc-Hieu Nguyen, Kazem Meidani, Amir Barati Farimani, Khoa D Doan, and Chandan K Reddy. 2025b. LLM-SRBench: A New Benchmark for Scientific Equation Discovery with Large Language Models. arXiv:2504.10415 [cs.CL] https://arxiv.org/abs/2504.10415
- Tang et al. (2023a) Hongwei Tang, Yan Wang, Tongguang Wang, and Linlin Tian. 2023a. Discovering explicit Reynolds-averaged turbulence closures for turbulent separated flows through deep learning-based symbolic regression with non-linear corrections. Physics of Fluids 35, 2 (2023), 025118. arXiv:https://doi.org/10.1063/5.0135638 doi: 10.1063/5.0135638
- Tang et al. (2023b) Hongwei Tang, Yan Wang, Tongguang Wang, and Linlin Tian. 2023b. Discovering explicit Reynolds-averaged turbulence closures for turbulent separated flows through deep learning-based symbolic regression with non-linear corrections. Physics of Fluids 35, 2 (Feb. 2023). doi: 10.1063/5.0135638
- Tennekes and Lumley (1972) Henk Tennekes and John L. Lumley. 1972. A First Course in Turbulence. The MIT Press. doi: 10.7551/mitpress/3014.001.0001
- Valipour et al. (2021) Mojtaba Valipour, Bowen You, Maysum Panju, and Ali Ghodsi. 2021. SymbolicGPT: A Generative Transformer Model for Symbolic Regression. arXiv:2106.14131 [cs.LG] https://arxiv.org/abs/2106.14131
- Vastl et al. (2024) Martin Vastl, Jonáš Kulhánek, Jiří Kubalík, Erik Derner, and Robert Babuška. 2024. SymFormer: End-to-End Symbolic Regression Using Transformer-Based Architecture. IEEE Access 12 (2024), 37840–37849. doi: 10.1109/ACCESS.2024.3374649
- Virgolin and Pissis (2022) Marco Virgolin and Solon P. Pissis. 2022. Symbolic Regression is NP-hard. arXiv:2207.01018 [cs.NE] https://arxiv.org/abs/2207.01018
- Wahlquist et al. (2024) Ylva Wahlquist, Jesper Sundell, and Kristian Soltesz. 2024. Learning pharmacometric covariate model structures with symbolic regression networks. Journal of Pharmacokinetics and Pharmacodynamics 51, 2 (2024), 155–167. doi: 10.1007/s10928-023-09887-3
- WANG et al. (2019) Wei WANG, Yang ZHANG, and Lili CHEN. 2019. An application of the shear stress transport low-Reynolds-number k-ε turbulence model on turbulent flows. ACTA AERODYNAMICA SINICA 37, 3 (2019), 419–425. doi: 10.7638/kqdlxxb-2016.0158
- Weller et al. (1998) H.G. Weller, Gavin Tabor, Hrvoje Jasak, and Christer Fureby. 1998. A Tensorial Approach to Computational Continuum Mechanics Using Object Orientated Techniques. Computers in Physics 12 (11 1998), 620–631. doi: 10.1063/1.168744
- Xia et al. (2025) Shijie Xia, Yuhan Sun, and Pengfei Liu. 2025. SR-Scientist: Scientific Equation Discovery With Agentic AI. arXiv:2510.11661 [cs.AI] https://arxiv.org/abs/2510.11661
- Xiao et al. (2020) Heng Xiao, Jin-Long Wu, Sylvain Laizet, and Lian Duan. 2020. Flows over periodic hills of parameterized geometries: A dataset for data-driven turbulence modeling from direct simulations. Computers & Fluids 200 (2020), 104431. doi: 10.1016/j.compfluid.2020.104431
- Zhang et al. (2025) Hengzhe Zhang, Qi Chen, Bing XUE, Wolfgang Banzhaf, and Mengjie Zhang. 2025. RAG-SR: Retrieval-Augmented Generation for Neural Symbolic Regression. In The Thirteenth International Conference on Learning Representations. https://openreview.net/forum?id=NdHka08uWn
| Fitting log-MSE scale | $\alpha$ | 1.0 | Scaling in $R_{\mathrm{fit}}=-\alpha\log(\mathrm{MSE}+\epsilon)$ . |
| --- | --- | --- | --- |
| Complexity penalty weight | $\lambda_{\mathrm{len}}$ | 5e-3 | Weight in $P_{\mathrm{cplx}}=\lambda_{\mathrm{len}}\cdot\mathrm{Length}(\mathrm{AST})$ . |
| Dimensional homogeneity penalty weight | $P_{\mathrm{dim}}$ | 1.0 | Penalty/reward weight for unit inconsistency (general-level constraint). |
| Differentiability penalty weight | $P_{\mathrm{diff}}$ | 0.5 | Penalty/reward weight for non-smoothness on the data-defined domain (general-level constraint). |
| Domain-specific constraint weight (j-th) | $P_{\mathrm{domain}}^{(j)}$ | 0.5 | Weight for the $j$ -th domain prior (e.g., realizability, wall BC, asymptotic scaling, energy consistency). |
| Gating threshold for physics constraints | $\delta_{\mathrm{gate}}$ | 1e-3 * MSE_{initial} | Activate physics penalties only after fitting reaches a sufficiently low-error regime. |
| Numerical stability constant in $\tau_{i}$ | $\epsilon$ | 1e-50 | Used in $\tau_{i}=\frac{|b_{i}|}{\sum_{j}|b_{j}|+\epsilon}$ to avoid division by zero. |
| Redundancy threshold | $\rho$ | 1e-2 | A term is considered redundant if $\tau_{i}\leq\rho$ . |
| Token penalty scale | $p$ | 0.5 | Scaling coefficient in $P_{\mathrm{tok}}=p\cdot\max(0,-\log(|b_{i}|+\epsilon))$ . |
Table 3. Hyperparameters for dual-constraint evaluation and token-level signal synthesis.
## Appendix A Additional Hyperparameters of PiT-PO
Table 3 lists the additional hyperparameters of PiT-PO, including (i) general-level physical penalties (dimensional homogeneity and differentiability), (ii) domain-specific constraint penalties, (iii) the gated activation threshold for physical constraints, and (iv) theorem-guided redundancy detection and token-level penalization.
## Appendix B Support Exclusion Theorem: Theory and Practice
### B.1. Preliminaries: Empirical Function Space and Orthogonality
** Definition B.1 (Basis functions / dictionary)**
*Let $\mathcal{X}\subseteq\mathbb{R}^{d}$ be the domain, and let $\{\phi_{j}:\mathcal{X}\to\mathbb{R}\}_{j\in\mathcal{S}}$ be a collection of candidate basis (dictionary) functions indexed by $\mathcal{S}$ . For any subset $\mathcal{K}\subseteq\mathcal{S}$ , denote the corresponding model subspace by
$$
\mathrm{span}\{\phi_{j}:j\in\mathcal{K}\}.
$$*
** Definition B.2 (Target functionf∗f^{*})**
*Assume the ground-truth target function admits a sparse expansion over the dictionary:
$$
f^{*}(x)=\sum_{j\in\mathcal{S}^{\prime}}a_{j}\,\phi_{j}(x),
$$
where $\mathcal{S}^{\prime}\subseteq\mathcal{S}$ is the true support set (indices of nonzero terms) and satisfies $|\mathcal{S}^{\prime}|\leq M$ . Moreover, the true coefficients are bounded as
$$
A\leq|a_{j}|\leq B,\qquad\forall j\in\mathcal{S}^{\prime}.
$$*
** Definition B.3 (Empirical inner product and empirical norm)**
*Given a finite dataset $D=\{x_{n}\}_{n=1}^{N}\subset\mathcal{X}$ , for any two functions $f,g:\mathcal{X}\to\mathbb{R}$ , define the empirical inner product by
$$
\langle f,g\rangle_{D}:=\frac{1}{N}\sum_{n=1}^{N}f(x_{n})\,g(x_{n}),
$$
and the induced empirical norm by
$$
\|f\|_{D}:=\sqrt{\langle f,f\rangle_{D}}.
$$
The empirical function space
$$
\mathcal{F}_{D}:=\{(f(x_{1}),\ldots,f(x_{N})):\ f:\mathcal{X}\to\mathbb{R}\}
$$
is a finite-dimensional inner-product space under $\langle\cdot,\cdot\rangle_{D}$ .*
** Definition B.4 (Empirical Orthogonality)**
*Two functions $f,g\in\mathcal{F}_{D}$ are empirically orthogonal if $\langle f,g\rangle_{D}=0$ .*
Convention. Unless stated otherwise, throughout this appendix we write $\langle\cdot,\cdot\rangle$ and $\|\cdot\|$ to mean the empirical inner product $\langle\cdot,\cdot\rangle_{D}$ and the empirical norm $\|\cdot\|_{D}$ .
### B.2. Proof of the Support Exclusion Theorem
** Theorem B.5 (Support Exclusion Theorem)**
*Let $\mathcal{K}\subseteq\mathcal{S}$ be a candidate skeleton (active set). Consider the empirical least-squares fit over $\mathcal{K}$ :
$$
b\in\arg\min_{\{b_{j}\}_{j\in\mathcal{K}}}\left\|f^{*}-\sum_{j\in\mathcal{K}}b_{j}\phi_{j}\right\|^{2}.
$$
Define the empirical Gram matrix $G\in\mathbb{R}^{|\mathcal{S}|\times|\mathcal{S}|}$ by
$$
G_{ij}:=\langle\phi_{i},\phi_{j}\rangle,\qquad i,j\in\mathcal{S},
$$
and assume $G_{ii}>0$ . For any $i\in\mathcal{S}$ , define
$$
T_{ij}:=\frac{G_{ji}}{G_{ii}},\qquad j\in\mathcal{S}.
$$
Further define the external-candidate vector
$$
u_{i}:=\big(|T_{i\ell}|\big)_{\ell\in\mathcal{S}\setminus\mathcal{K}}\in\mathbb{R}^{|\mathcal{S}\setminus\mathcal{K}|},
$$
and let $s(1)\geq s(2)\geq\cdots\geq s(|\mathcal{S}\setminus\mathcal{K}|)$ be the entries of $u_{i}$ sorted in non-increasing order. If for some $i\in\mathcal{K}$ ,
$$
|b_{i}|<A-\left(\underbrace{\sum_{j\in\mathcal{K},\,j\neq i}(B+|b_{j}|)\,|T_{ij}|}_{\textnormal{Internal Interference}}+\underbrace{B\sum_{k=1}^{|\mathcal{S}\setminus\mathcal{K}|}s(k)}_{\textnormal{External Interference}}\right),
$$
then $i\notin\mathcal{S}^{\prime}$ .*
* Proof*
Step 1: Empirical orthogonality (first-order optimality of least squares). Let
$$
g(x):=\sum_{j\in\mathcal{K}}b_{j}\phi_{j}(x),\qquad r(x):=f^{*}(x)-g(x).
$$
The objective is $\|r\|^{2}=\langle r,r\rangle$ . For any $i\in\mathcal{K}$ , the first-order optimality condition yields
$$
0=\frac{\partial}{\partial b_{i}}\|r\|^{2}=-2\langle r,\phi_{i}\rangle\quad\Longrightarrow\quad\langle r,\phi_{i}\rangle=0.
$$
Hence, the residual $r$ is empirically orthogonal to $\mathrm{span}\{\phi_{i}:i\in\mathcal{K}\}$ . Step 2: Set decomposition. Define three disjoint sets
$$
\mathcal{T}:=\mathcal{K}\cap\mathcal{S}^{\prime},\qquad\mathcal{F}:=\mathcal{K}\setminus\mathcal{S}^{\prime},\qquad\mathcal{R}:=\mathcal{S}^{\prime}\setminus\mathcal{K}.
$$
Then $\mathcal{K}=\mathcal{T}\cup\mathcal{F}$ and $\mathcal{S}^{\prime}=\mathcal{T}\cup\mathcal{R}$ . Expanding the residual gives
$$
r=\sum_{j\in\mathcal{T}}(a_{j}-b_{j})\phi_{j}+\sum_{j\in\mathcal{R}}a_{j}\phi_{j}-\sum_{j\in\mathcal{F}}b_{j}\phi_{j}.
$$ Step 3: An exact coefficient identity for $i\in\mathcal{T}$ . Fix any $i\in\mathcal{T}\subseteq\mathcal{K}$ . Using $\langle r,\phi_{i}\rangle=0$ and writing $G_{ji}=\langle\phi_{j},\phi_{i}\rangle$ , we obtain
$$
(a_{i}-b_{i})G_{ii}+\sum_{j\in\mathcal{T},\,j\neq i}(a_{j}-b_{j})G_{ji}+\sum_{j\in\mathcal{R}}a_{j}G_{ji}-\sum_{j\in\mathcal{F}}b_{j}G_{ji}=0.
$$
Dividing by $G_{ii}>0$ and using $T_{ij}=G_{ji}/G_{ii}$ yields
$$
a_{i}-b_{i}=-\sum_{j\in\mathcal{T},\,j\neq i}(a_{j}-b_{j})T_{ij}-\sum_{j\in\mathcal{R}}a_{j}T_{ij}+\sum_{j\in\mathcal{F}}b_{j}T_{ij}.
$$ Step 4: Upper bound via internal and external interference. Taking absolute values and applying the triangle inequality gives
$$
|a_{i}-b_{i}|\leq\sum_{j\in\mathcal{T},\,j\neq i}|a_{j}-b_{j}|\,|T_{ij}|+\sum_{j\in\mathcal{R}}|a_{j}|\,|T_{ij}|+\sum_{j\in\mathcal{F}}|b_{j}|\,|T_{ij}|.
$$
For $j\in\mathcal{T}\subseteq\mathcal{S}^{\prime}$ , we have $|a_{j}|\leq B$ and thus
$$
|a_{j}-b_{j}|\leq|a_{j}|+|b_{j}|\leq B+|b_{j}|.
$$
Therefore,
$$
|a_{i}-b_{i}|\leq\sum_{j\in\mathcal{T},\,j\neq i}(B+|b_{j}|)\,|T_{ij}|+\sum_{j\in\mathcal{F}}|b_{j}|\,|T_{ij}|+B\sum_{j\in\mathcal{R}}|T_{ij}|.
$$
Since $|b_{j}|\leq B+|b_{j}|$ for all $j\in\mathcal{F}$ , we further have
$$
\sum_{j\in\mathcal{F}}|b_{j}|\,|T_{ij}|\leq\sum_{j\in\mathcal{F}}(B+|b_{j}|)\,|T_{ij}|.
$$
Combining $\mathcal{T}\setminus\{i\}$ and $\mathcal{F}$ yields the internal-interference term:
$$
|a_{i}-b_{i}|\leq\underbrace{\sum_{j\in\mathcal{K},\,j\neq i}(B+|b_{j}|)\,|T_{ij}|}_{\textnormal{Internal Interference}}+B\sum_{j\in\mathcal{R}}|T_{ij}|.
$$
For the external term, note that $\mathcal{R}\subseteq\mathcal{S}\setminus\mathcal{K}$ , and the entries $\{|T_{i\ell}|\}_{\ell\in\mathcal{S}\setminus\mathcal{K}}$ sorted in non-increasing order are $s(1)\geq\cdots\geq s(|\mathcal{S}\setminus\mathcal{K}|)$ . Hence, for any subset $\mathcal{R}$ ,
$$
\sum_{j\in\mathcal{R}}|T_{ij}|\leq\sum_{k=1}^{|\mathcal{R}|}s(k).
$$
Assuming $|\mathcal{S}^{\prime}|\leq M$ and that the current support is partially correct, i.e., $\mathcal{K}\cap\mathcal{S}^{\prime}\neq\emptyset$ , we have $|\mathcal{R}|=|\mathcal{S}^{\prime}\setminus\mathcal{K}|\leq|\mathcal{S}^{\prime}|-1\leq M-1$ . Define $m:=\min\!\big(M-1,\ |\mathcal{S}\setminus\mathcal{K}|\big)$ . Since also $|\mathcal{R}|\leq|\mathcal{S}\setminus\mathcal{K}|$ , it follows that $|\mathcal{R}|\leq m$ , and thus
$$
\sum_{j\in\mathcal{R}}|T_{ij}|\leq\sum_{k=1}^{|\mathcal{R}|}s(k)\leq\sum_{k=1}^{m}s(k).
$$
Thus,
$$
|a_{i}-b_{i}|\leq\sum_{j\in\mathcal{K},\,j\neq i}(B+|b_{j}|)\,|T_{ij}|+B\sum_{k=1}^{m}s(k).
$$ Step 5: Lower bound when $i\in\mathcal{S}^{\prime}$ . If $i\in\mathcal{S}^{\prime}$ , then $|a_{i}|\geq A$ . By the reverse triangle inequality,
$$
|a_{i}-b_{i}|\geq\big||a_{i}|-|b_{i}|\big|\geq A-|b_{i}|.
$$ Step 6: Contradiction. Assume, for contradiction, that there exists $i\in\mathcal{K}\cap\mathcal{S}^{\prime}$ satisfying
$$
|b_{i}|<A-\left(\sum_{j\in\mathcal{K},\,j\neq i}(B+|b_{j}|)\,|T_{ij}|+B\sum_{k=1}^{m}s(k)\right).
$$
Rearranging yields
$$
A-|b_{i}|>\sum_{j\in\mathcal{K},\,j\neq i}(B+|b_{j}|)\,|T_{ij}|+B\sum_{k=1}^{m}s(k).
$$
However, Step 5 implies $|a_{i}-b_{i}|\geq A-|b_{i}|$ , while the bound above implies
$$
|a_{i}-b_{i}|\leq\sum_{j\in\mathcal{K},\,j\neq i}(B+|b_{j}|)\,|T_{ij}|+B\sum_{k=1}^{m}s(k),
$$
which is a contradiction. Therefore such an $i$ cannot belong to $\mathcal{S}^{\prime}$ , i.e., $i\notin\mathcal{S}^{\prime}$ . ∎
## Appendix C Supplementary Diagnostics and Analyses
### C.1. Remaining Iteration Curves on Smaller Backbones
In the main text, we report iteration curves for the 8B backbone. Here we provide the remaining curves for 3B and 1B (Figure 8), which corroborate the observations in Section 4.4.
<details>
<summary>x8.png Details</summary>
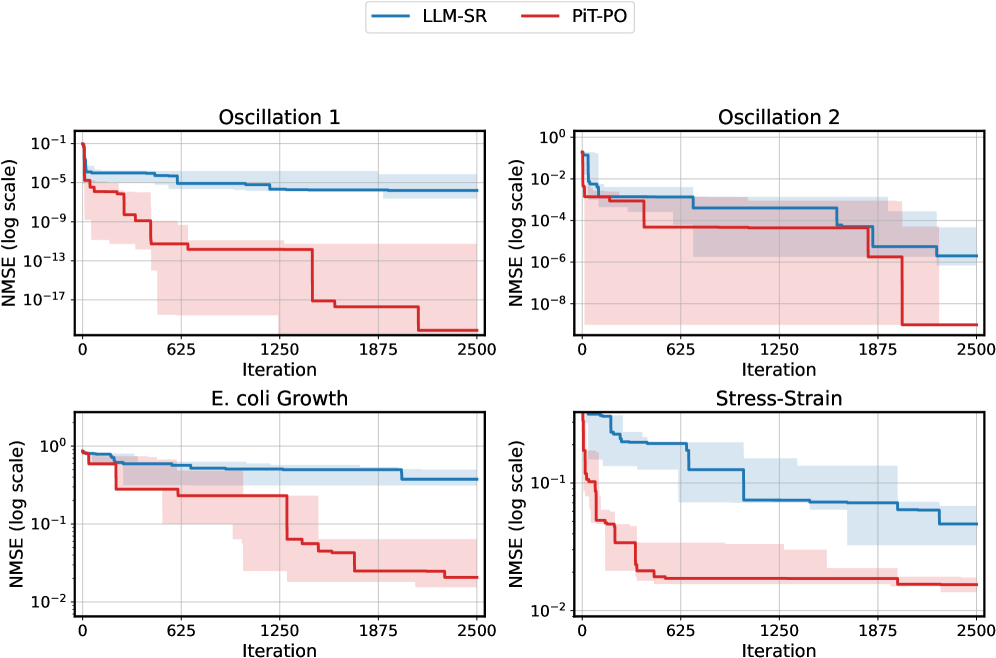
### Visual Description
## Chart: NMSE vs. Iteration for Different Models
### Overview
The image presents four separate line charts, each depicting the Normalized Mean Squared Error (NMSE) on a logarithmic scale against the number of iterations. Two models, "LLM-SR" (blue) and "PIT-PO" (red), are compared across four different datasets: "Oscillation 1", "Oscillation 2", "E. coli Growth", and "Stress-Strain". Each line represents the average NMSE for the respective model, and shaded areas indicate the standard deviation.
### Components/Axes
* **X-axis:** Iteration (ranging from 0 to 2500)
* **Y-axis:** NMSE (log scale)
* **Legend:**
* LLM-SR (Blue)
* PIT-PO (Red)
* **Chart Titles:**
* Oscillation 1 (Top-Left)
* Oscillation 2 (Top-Right)
* E. coli Growth (Bottom-Left)
* Stress-Strain (Bottom-Right)
### Detailed Analysis or Content Details
**Oscillation 1 (Top-Left):**
* **LLM-SR (Blue):** The line starts at approximately 1e-1 and slopes downward, reaching approximately 1e-13 by iteration 2500. The shaded area indicates a relatively small standard deviation throughout the iterations.
* **PIT-PO (Red):** The line begins at approximately 5e-1 and also slopes downward, but more gradually than LLM-SR. It reaches approximately 5e-10 by iteration 2500. The shaded area is wider than LLM-SR, indicating a larger standard deviation.
**Oscillation 2 (Top-Right):**
* **LLM-SR (Blue):** The line starts at approximately 1e-2 and decreases to approximately 1e-8 by iteration 2500. The standard deviation is relatively small.
* **PIT-PO (Red):** The line begins at approximately 1e-2 and decreases to approximately 1e-6 by iteration 2500. The standard deviation is larger than LLM-SR.
**E. coli Growth (Bottom-Left):**
* **LLM-SR (Blue):** The line starts at approximately 1e0 and remains relatively stable around 1e-1 to 1e-2 for most of the iterations, with a slight decrease towards the end. The standard deviation is significant.
* **PIT-PO (Red):** The line starts at approximately 1e0 and decreases more rapidly than LLM-SR, reaching approximately 1e-2 by iteration 2500. The standard deviation is also significant.
**Stress-Strain (Bottom-Right):**
* **LLM-SR (Blue):** The line starts at approximately 1e0 and decreases to approximately 1e-1 by iteration 2500. The standard deviation is relatively small.
* **PIT-PO (Red):** The line starts at approximately 1e0 and decreases more rapidly than LLM-SR, reaching approximately 5e-2 by iteration 2500. The standard deviation is larger than LLM-SR.
### Key Observations
* LLM-SR consistently outperforms PIT-PO in terms of NMSE across all four datasets, especially in "Oscillation 1" and "Oscillation 2".
* The standard deviation for LLM-SR is generally smaller than that of PIT-PO, indicating more stable performance.
* The "E. coli Growth" dataset shows the least amount of improvement in NMSE for both models, with both lines remaining relatively high.
* The "Stress-Strain" dataset shows a more significant decrease in NMSE for both models.
### Interpretation
The charts demonstrate the performance of two models, LLM-SR and PIT-PO, in predicting different datasets. The NMSE metric, plotted on a logarithmic scale, indicates the accuracy of the predictions. Lower NMSE values signify better performance.
LLM-SR consistently achieves lower NMSE values across all datasets, suggesting it is a more accurate model than PIT-PO. The smaller standard deviations associated with LLM-SR indicate that its performance is more consistent and less sensitive to variations in the data.
The "E. coli Growth" dataset presents a challenge for both models, as the NMSE values remain relatively high even after 2500 iterations. This suggests that the underlying dynamics of E. coli growth are more difficult to model accurately.
The differences in performance between the models and datasets could be attributed to the complexity of the data, the model architectures, and the optimization algorithms used during training. The logarithmic scale highlights the magnitude of the errors, making it easier to compare the performance of the models across different datasets. The shaded areas representing standard deviation provide insight into the robustness of each model's performance.
</details>
(a) Llama-3B
<details>
<summary>x9.png Details</summary>
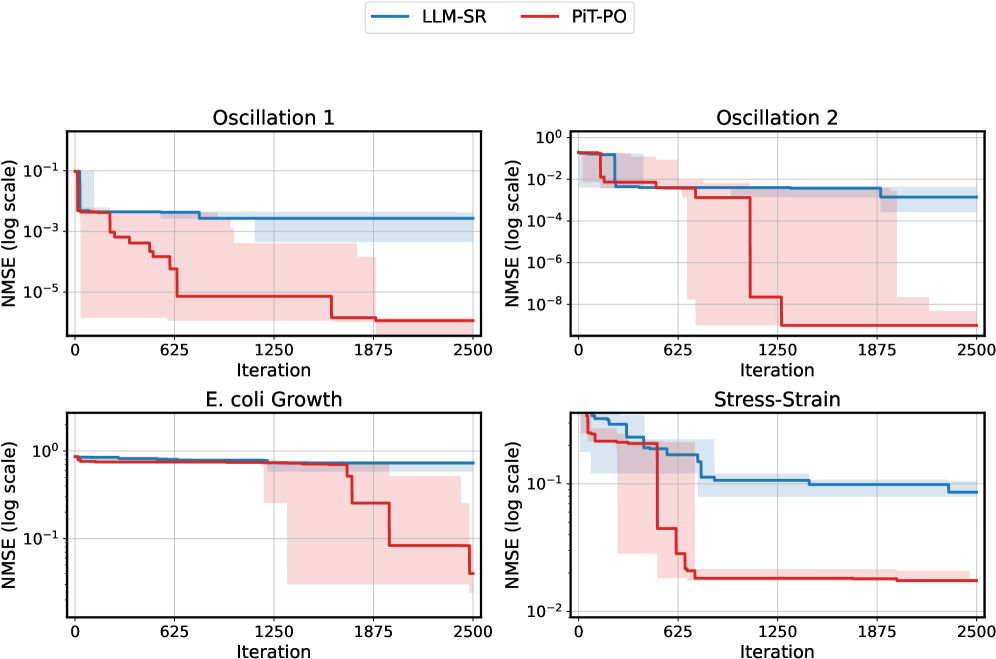
### Visual Description
## Chart: NMSE vs. Iteration for Different Models
### Overview
The image presents four separate line charts, each depicting the Normalized Mean Squared Error (NMSE) on a logarithmic scale against the number of iterations. Two models, "LLM-SR" (represented by a blue line) and "PIT-PO" (represented by a red line), are compared across four different scenarios: Oscillation 1, Oscillation 2, E. coli Growth, and Stress-Strain. Shaded regions around each line indicate the standard deviation or confidence interval.
### Components/Axes
* **X-axis:** Iteration, ranging from 0 to 2500.
* **Y-axis:** NMSE (log scale). The scale varies for each chart, but generally spans several orders of magnitude.
* **Legend:** Located at the top-right of the image, it identifies the lines:
* LLM-SR (Blue)
* PIT-PO (Red)
* **Chart Titles:** Each subplot has a title indicating the scenario:
* Oscillation 1 (Top-Left)
* Oscillation 2 (Top-Right)
* E. coli Growth (Bottom-Left)
* Stress-Strain (Bottom-Right)
* **Gridlines:** Present in all charts to aid in reading values.
### Detailed Analysis or Content Details
**Oscillation 1 (Top-Left):**
* Both lines start at approximately 10^-1.
* The blue line (LLM-SR) shows a consistent downward trend, decreasing to approximately 10^-5 by iteration 2500.
* The red line (PIT-PO) also decreases, but more erratically, ending at approximately 10^-3 by iteration 2500.
* The shaded region around the blue line is relatively narrow, indicating lower variance. The red line's shaded region is wider, suggesting higher variance.
**Oscillation 2 (Top-Right):**
* Both lines start around 10^-2.
* The blue line (LLM-SR) initially decreases, then plateaus around 10^-5.
* The red line (PIT-PO) fluctuates around 10^-2, with some dips and rises.
* The shaded regions are relatively narrow for both lines.
**E. coli Growth (Bottom-Left):**
* Both lines start around 10^0.
* The blue line (LLM-SR) decreases in steps, reaching approximately 10^-1 by iteration 2500.
* The red line (PIT-PO) decreases more rapidly and in larger steps, reaching approximately 10^-2 by iteration 2500.
* The shaded regions are wider, indicating higher variance.
**Stress-Strain (Bottom-Right):**
* Both lines start around 10^-1.
* The blue line (LLM-SR) decreases gradually to approximately 10^-2.
* The red line (PIT-PO) initially decreases, then increases sharply around iteration 1250, reaching approximately 10^-1 before decreasing again to approximately 10^-2.
* The shaded regions are relatively narrow for the blue line, but wider for the red line, especially during the increase around iteration 1250.
### Key Observations
* LLM-SR generally exhibits smoother and more consistent decreases in NMSE compared to PIT-PO.
* PIT-PO shows more variability and, in the Stress-Strain scenario, a notable increase in NMSE at iteration 1250.
* The E. coli Growth scenario shows the most significant reduction in NMSE for both models.
* The logarithmic scale emphasizes the relative changes in NMSE, making it easier to compare performance across different scenarios.
### Interpretation
The charts compare the performance of two models, LLM-SR and PIT-PO, in terms of their ability to minimize the Normalized Mean Squared Error (NMSE) across four different dynamic systems. The consistent downward trend of LLM-SR in most scenarios suggests it is more stable and reliable in reducing error. The fluctuations and occasional increases in NMSE for PIT-PO indicate it may be more sensitive to the specific dynamics of the system or require more careful tuning.
The E. coli Growth scenario shows the largest error reduction, potentially indicating that the models are well-suited for modeling biological systems. The Stress-Strain scenario, with PIT-PO's increase in NMSE, suggests a potential instability or limitation of that model under certain conditions. The shaded regions around the lines represent the uncertainty or variance in the model's performance, providing a measure of confidence in the results.
The use of a logarithmic scale is crucial for visualizing the wide range of NMSE values, allowing for a clear comparison of performance even when errors are very small. The charts collectively demonstrate the importance of model selection and parameter tuning for achieving optimal performance in different dynamic systems.
</details>
(b) Llama-1B
Figure 8. NMSE versus iteration on smaller backbones, consistent with the trends in the 8B setting.
<details>
<summary>x10.png Details</summary>
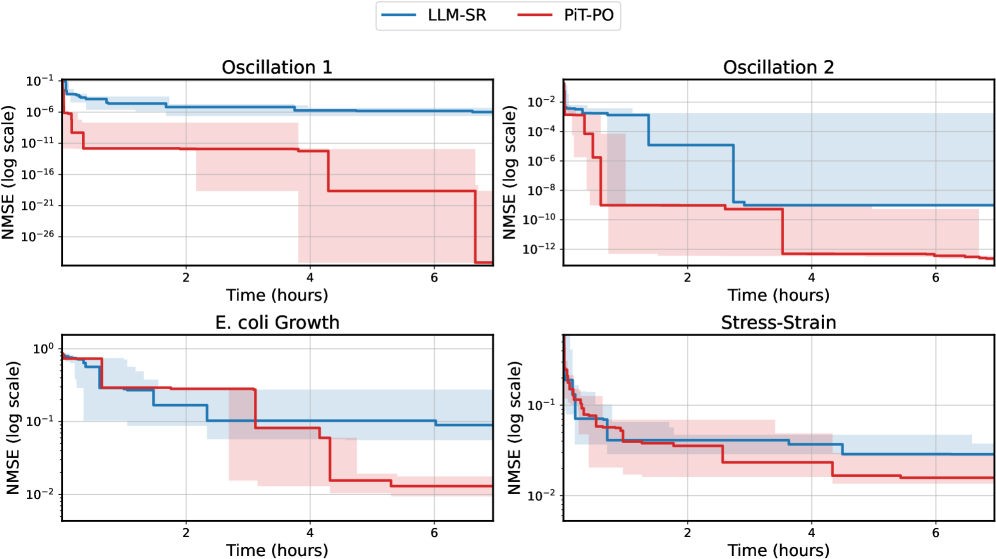
### Visual Description
\n
## Chart: NMSE over Time for Different Methods
### Overview
The image presents four line charts, each depicting the Normalized Mean Squared Error (NMSE) on a logarithmic scale against Time (in hours). Two methods, LLM-SR (blue) and PIT-PO (red), are compared across four different scenarios: Oscillation 1, Oscillation 2, E. coli Growth, and Stress-Strain. Each chart includes shaded regions representing uncertainty around the respective lines.
### Components/Axes
* **X-axis:** Time (hours), ranging from 0 to 6 hours.
* **Y-axis:** NMSE (log scale), ranging from 10<sup>-26</sup> to 10<sup>2</sup>.
* **Lines:**
* LLM-SR (blue)
* PIT-PO (red)
* **Shaded Regions:** Represent uncertainty around each line. The shading is light blue for LLM-SR and light red for PIT-PO.
* **Titles:** Each subplot has a title indicating the scenario: "Oscillation 1", "Oscillation 2", "E. coli Growth", "Stress-Strain".
* **Legend:** Located in the top-right corner, labeling the lines as "LLM-SR" and "PIT-PO".
### Detailed Analysis or Content Details
**Oscillation 1 (Top-Left):**
* LLM-SR (blue): Starts at approximately 10<sup>-1</sup> NMSE and decreases to approximately 10<sup>-26</sup> NMSE by 6 hours. The line is relatively flat between 2 and 4 hours.
* PIT-PO (red): Starts at approximately 10<sup>-1</sup> NMSE and decreases to approximately 10<sup>-16</sup> NMSE by 6 hours. The line has a step-wise decrease, remaining constant for periods before dropping.
**Oscillation 2 (Top-Right):**
* LLM-SR (blue): Starts at approximately 10<sup>-2</sup> NMSE and decreases to approximately 10<sup>-12</sup> NMSE by 6 hours. The line is relatively flat between 2 and 4 hours.
* PIT-PO (red): Starts at approximately 10<sup>-4</sup> NMSE and decreases to approximately 10<sup>-12</sup> NMSE by 6 hours. The line has a step-wise decrease, remaining constant for periods before dropping.
**E. coli Growth (Bottom-Left):**
* LLM-SR (blue): Starts at approximately 10<sup>0</sup> NMSE and decreases to approximately 10<sup>-2</sup> NMSE by 6 hours. The line is relatively flat between 2 and 4 hours.
* PIT-PO (red): Starts at approximately 10<sup>0</sup> NMSE and decreases to approximately 10<sup>-2</sup> NMSE by 6 hours. The line has a step-wise decrease, remaining constant for periods before dropping.
**Stress-Strain (Bottom-Right):**
* LLM-SR (blue): Starts at approximately 10<sup>1</sup> NMSE and decreases to approximately 10<sup>-1</sup> NMSE by 6 hours. The line is relatively flat between 2 and 4 hours.
* PIT-PO (red): Starts at approximately 10<sup>1</sup> NMSE and decreases to approximately 10<sup>-1</sup> NMSE by 6 hours. The line has a step-wise decrease, remaining constant for periods before dropping.
### Key Observations
* In all four scenarios, both LLM-SR and PIT-PO show a decreasing trend in NMSE over time, indicating improved performance as time progresses.
* LLM-SR generally achieves lower NMSE values than PIT-PO in all scenarios, especially after 4 hours.
* PIT-PO exhibits a step-wise decrease in NMSE, while LLM-SR shows a smoother, more continuous decrease.
* The uncertainty regions (shaded areas) are relatively wide, particularly in the early stages of each scenario, indicating higher variability in the NMSE estimates.
### Interpretation
The charts demonstrate the performance of two methods, LLM-SR and PIT-PO, in predicting or modeling different dynamic systems. The decreasing NMSE values suggest that both methods improve their accuracy over time. LLM-SR consistently outperforms PIT-PO across all scenarios, suggesting it is a more effective method for these types of problems. The step-wise nature of PIT-PO's NMSE reduction might indicate a discrete update or adjustment process within the method. The wide uncertainty regions early on suggest that initial predictions are less reliable, but become more stable as time progresses and more data is available. The different scenarios (Oscillation, E. coli Growth, Stress-Strain) represent diverse applications, and the consistent outperformance of LLM-SR suggests its robustness and generalizability. The logarithmic scale emphasizes the significant reduction in error achieved by both methods, even if the absolute error remains non-negligible.
</details>
(a) Llama-3.1-8B
<details>
<summary>x11.png Details</summary>
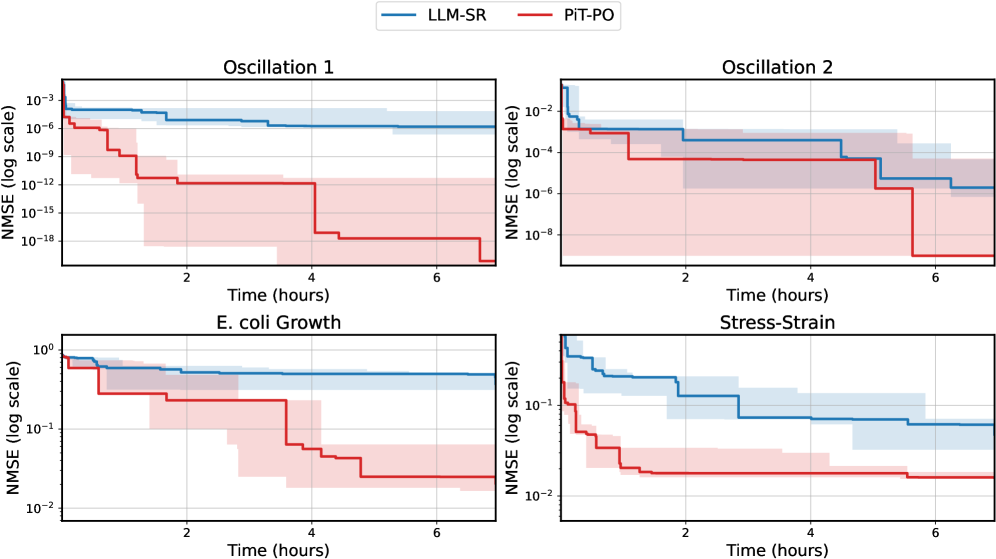
### Visual Description
\n
## Line Charts: NMSE vs. Time for Different Models
### Overview
The image presents four line charts, each depicting the Normalized Mean Squared Error (NMSE) on a logarithmic scale against Time (in hours). Two models, LLM-SR (blue) and PIT-PO (red), are compared across four different conditions: Oscillation 1, Oscillation 2, E. coli Growth, and Stress-Strain. Each line represents the mean NMSE, and the shaded area around each line represents the standard deviation.
### Components/Axes
* **X-axis:** Time (hours), ranging from 0 to 6 hours.
* **Y-axis:** NMSE (log scale), ranging from 10<sup>-18</sup> to 10<sup>1</sup>.
* **Models:**
* LLM-SR (blue)
* PIT-PO (red)
* **Conditions (Chart Titles):**
* Oscillation 1
* Oscillation 2
* E. coli Growth
* Stress-Strain
### Detailed Analysis or Content Details
**Oscillation 1 (Top-Left):**
* **LLM-SR (Blue):** The line starts at approximately 8 x 10<sup>-4</sup> NMSE at time 0, decreases steadily to approximately 2 x 10<sup>-12</sup> NMSE at time 6 hours. The shaded area indicates a relatively small standard deviation.
* **PIT-PO (Red):** The line starts at approximately 1 x 10<sup>-3</sup> NMSE at time 0, decreases rapidly to approximately 5 x 10<sup>-15</sup> NMSE at time 4 hours, and then plateaus. The shaded area is wider than for LLM-SR, indicating a larger standard deviation.
**Oscillation 2 (Top-Right):**
* **LLM-SR (Blue):** The line starts at approximately 2 x 10<sup>-2</sup> NMSE at time 0, decreases steadily to approximately 1 x 10<sup>-6</sup> NMSE at time 6 hours. The shaded area is relatively narrow.
* **PIT-PO (Red):** The line starts at approximately 3 x 10<sup>-2</sup> NMSE at time 0, decreases to approximately 2 x 10<sup>-5</sup> NMSE at time 2 hours, then decreases more slowly to approximately 5 x 10<sup>-7</sup> NMSE at time 6 hours. The shaded area is wider than for LLM-SR.
**E. coli Growth (Bottom-Left):**
* **LLM-SR (Blue):** The line starts at approximately 0.8 NMSE at time 0, decreases to approximately 0.02 NMSE at time 6 hours. The shaded area is relatively narrow.
* **PIT-PO (Red):** The line starts at approximately 1 NMSE at time 0, decreases rapidly to approximately 0.1 NMSE at time 2 hours, then decreases more slowly to approximately 0.03 NMSE at time 6 hours. The shaded area is wider than for LLM-SR.
**Stress-Strain (Bottom-Right):**
* **LLM-SR (Blue):** The line starts at approximately 0.1 NMSE at time 0, decreases to approximately 0.01 NMSE at time 6 hours. The shaded area is relatively narrow.
* **PIT-PO (Red):** The line starts at approximately 0.2 NMSE at time 0, decreases to approximately 0.02 NMSE at time 6 hours. The shaded area is wider than for LLM-SR.
### Key Observations
* In all four conditions, LLM-SR generally exhibits a lower NMSE than PIT-PO, indicating better performance.
* The standard deviation around the LLM-SR lines is consistently smaller than that around the PIT-PO lines, suggesting more stable performance.
* The rate of NMSE decrease is generally faster for both models in the initial stages (0-2 hours) and then slows down.
* The PIT-PO model shows a more pronounced plateau in the Oscillation 1 chart.
### Interpretation
The data suggests that the LLM-SR model consistently outperforms the PIT-PO model across all four tested conditions. The lower NMSE values and smaller standard deviations indicate that LLM-SR provides more accurate and reliable predictions. The initial rapid decrease in NMSE for both models likely reflects the models quickly learning the underlying dynamics of each condition. The subsequent slower decrease and eventual plateau suggest that the models are approaching their limits of accuracy or that the dynamics become more complex over time. The wider standard deviation for PIT-PO suggests that its performance is more sensitive to variations in the data or initial conditions. The different conditions (Oscillations, E. coli growth, Stress-Strain) represent different types of dynamic systems, and the consistent outperformance of LLM-SR suggests that it is a more robust and versatile model for predicting these types of systems. The logarithmic scale on the Y-axis emphasizes the magnitude of the error reduction, particularly at lower NMSE values.
</details>
(b) Llama-3.2-3B
<details>
<summary>x12.png Details</summary>
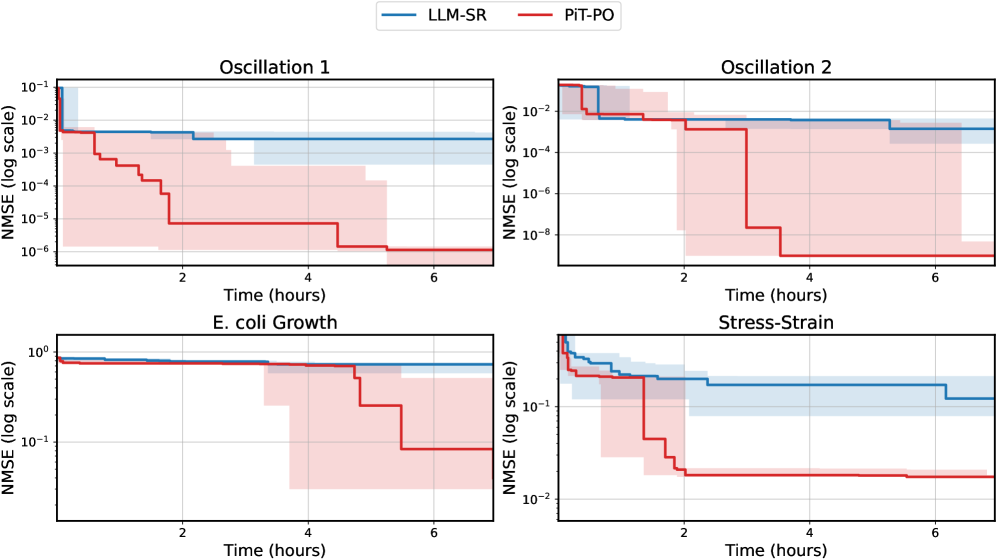
### Visual Description
\n
## Chart: NMSE vs. Time for Different Oscillations and Conditions
### Overview
The image presents four line charts, each displaying the Normalized Mean Squared Error (NMSE) on a logarithmic scale against Time (in hours). Each chart represents a different condition: Oscillation 1, Oscillation 2, E. coli Growth, and Stress-Strain. Two methods, LLM-SR (blue) and PIT-PO (red), are compared within each chart. Shaded areas around the lines represent uncertainty or variance.
### Components/Axes
* **X-axis:** Time (hours), ranging from approximately 0 to 7 hours.
* **Y-axis:** NMSE (log scale), ranging from 10<sup>-8</sup> to 10<sup>1</sup>. The scale is logarithmic.
* **Legend:**
* LLM-SR (Blue line)
* PIT-PO (Red line)
* **Chart Titles:**
* Oscillation 1 (Top-left)
* Oscillation 2 (Top-right)
* E. coli Growth (Bottom-left)
* Stress-Strain (Bottom-right)
### Detailed Analysis
**Oscillation 1 (Top-left):**
* The blue line (LLM-SR) starts at approximately 0.01 and decreases to approximately 0.0001 over the 7 hours. The line is relatively smooth.
* The red line (PIT-PO) starts at approximately 0.001 and decreases to approximately 0.00001 over the 7 hours. The line is stepped, with plateaus.
* The shaded area around the blue line is smaller than the shaded area around the red line, indicating less uncertainty for LLM-SR.
**Oscillation 2 (Top-right):**
* The blue line (LLM-SR) starts at approximately 0.02 and decreases to approximately 0.0005 over the 7 hours. The line is relatively smooth.
* The red line (PIT-PO) starts at approximately 0.01 and decreases to approximately 0.0001 over the 7 hours. The line is stepped, with plateaus.
* The shaded area around the blue line is smaller than the shaded area around the red line, indicating less uncertainty for LLM-SR.
**E. coli Growth (Bottom-left):**
* The blue line (LLM-SR) starts at approximately 0.1 and decreases to approximately 0.01 over the 7 hours. The line is relatively smooth.
* The red line (PIT-PO) starts at approximately 0.05 and decreases to approximately 0.005 over the 7 hours. The line is stepped, with plateaus.
* The shaded area around the blue line is smaller than the shaded area around the red line, indicating less uncertainty for LLM-SR.
**Stress-Strain (Bottom-right):**
* The blue line (LLM-SR) starts at approximately 0.2 and decreases to approximately 0.02 over the 7 hours. The line is relatively smooth.
* The red line (PIT-PO) starts at approximately 0.1 and decreases to approximately 0.005 over the 7 hours. The line is stepped, with plateaus.
* The shaded area around the blue line is smaller than the shaded area around the red line, indicating less uncertainty for LLM-SR.
### Key Observations
* In all four conditions, the LLM-SR method (blue line) generally exhibits lower NMSE values than the PIT-PO method (red line) at most time points.
* The PIT-PO method (red line) consistently shows a stepped pattern, suggesting discrete updates or adjustments in its error calculation.
* The uncertainty (shaded areas) around the LLM-SR lines is consistently smaller than that around the PIT-PO lines, indicating more stable and reliable performance.
* The NMSE decreases over time for both methods in all conditions, indicating improved performance or convergence.
### Interpretation
The charts demonstrate that the LLM-SR method consistently outperforms the PIT-PO method across all tested conditions (Oscillation 1, Oscillation 2, E. coli Growth, and Stress-Strain) in terms of NMSE. The lower NMSE values and smaller uncertainty intervals suggest that LLM-SR provides more accurate and reliable predictions or estimations compared to PIT-PO. The stepped nature of the PIT-PO results suggests that its calculations are performed in discrete steps, potentially leading to less smooth and potentially less accurate results. The decreasing NMSE over time for both methods indicates that both methods are learning or adapting to the data, but LLM-SR does so more effectively. The logarithmic scale emphasizes the magnitude of the error reduction, particularly at lower NMSE values. The consistent performance advantage of LLM-SR across diverse conditions suggests its robustness and generalizability.
</details>
(c) Llama-3.2-1B
Figure 9. Wall-clock efficiency under a fixed 25,000-second budget.
### C.2. Time-Cost Analysis under a Fixed Wall-Clock Budget
To complement the iteration-based efficiency analysis in Section 4.4, we evaluate search efficiency under a strict wall-clock constraint. We fix the total runtime of each method to 25,000 seconds ( $\approx$ 6.9 hours) and track the best-so-far NMSE as a function of elapsed time (Figure 9). This protocol directly answers a practical question: given the same compute time, which method returns a more accurate recovered equation?
Across all three backbones (Llama-3.1-8B, Llama-3.2-3B, and Llama-3.2-1B), PiT-PO consistently yields lower NMSE trajectories than LLM-SR under the same 25,000-second budget. In line with the behavior discussed in Section 4.4, the curves exhibit a characteristic divergence in the low-error regime: LLM-SR often plateaus after an initial reduction, while PiT-PO continues to realize step-wise decreases over time, indicating more effective late-stage credit assignment and continued progress rather than stagnation.
The advantage is most pronounced on the two oscillator systems. For 8B and 3B, PiT-PO achieves rapid early improvements and maintains additional phase transitions later in the run, reaching substantially lower NMSE within the same wall-clock budget. For the 1B model, both methods converge more slowly overall, yet PiT-PO still more reliably escapes stagnation and attains markedly lower final NMSE, suggesting that the proposed in-search tuning remains effective even in the small-model regime.
Similar trends hold for the E. coli Growth and Stress–Strain tasks, where progress is typically more gradual. Under equal wall-clock time, PiT-PO achieves a lower final NMSE and exhibits clearer late-stage improvements, whereas LLM-SR tends to flatten earlier. Importantly, these gains are obtained without increasing runtime: despite the additional overhead introduced by in-search policy optimization, PiT-PO delivers consistently better return-on-compute within a fixed time budget, validating the practical efficiency of the proposed approach.
| LLM-SR (Llama-3.1-8B) LLM-SR (Llama-3.2-3B) LLM-SR (Llama-3.2-1B) | 0.0114 0.0191 0.1182 | 8.05e-10 5.56e-5 7.57e-3 | 0.1676 809.94 1888.5 | 0.1026 0.0264 0.2689 |
| --- | --- | --- | --- | --- |
| PiT-PO (Llama-3.1-8B) | 1.63e-30 | 1.36e-11 | 0.1163 | 0.0163 |
| PiT-PO (Llama-3.2-3B) | 2.63e-30 | 8.52e-10 | 0.0237 | 0.0144 |
| PiT-PO (Llama-3.2-1B) | 5.99e-5 | 8.58e-10 | 0.3462 | 0.0124 |
Table 4. OOD evaluation: NMSE on the OOD split.
### C.3. OOD Evaluation (NMSE ${}_{\text{OOD}}$ )
We report out-of-distribution performance using NMSE on the OOD split (NMSE ${}_{\text{OOD}}$ ), summarized in Table 4. Across all three backbones (8B/3B/1B) and all four tasks, PiT-PO achieves lower NMSE ${}_{\text{OOD}}$ than LLM-SR under the same setting. In particular, PiT-PO maintains extremely small OOD errors on the two oscillator systems, and yields substantially reduced OOD NMSE on E. coli growth and Stress–Strain, where LLM-SR exhibits noticeably larger errors. These results provide consistent evidence that the improvements observed in the main text persist under OOD evaluation.
### C.4. Equation Iterative Trajectories
<details>
<summary>x13.png Details</summary>
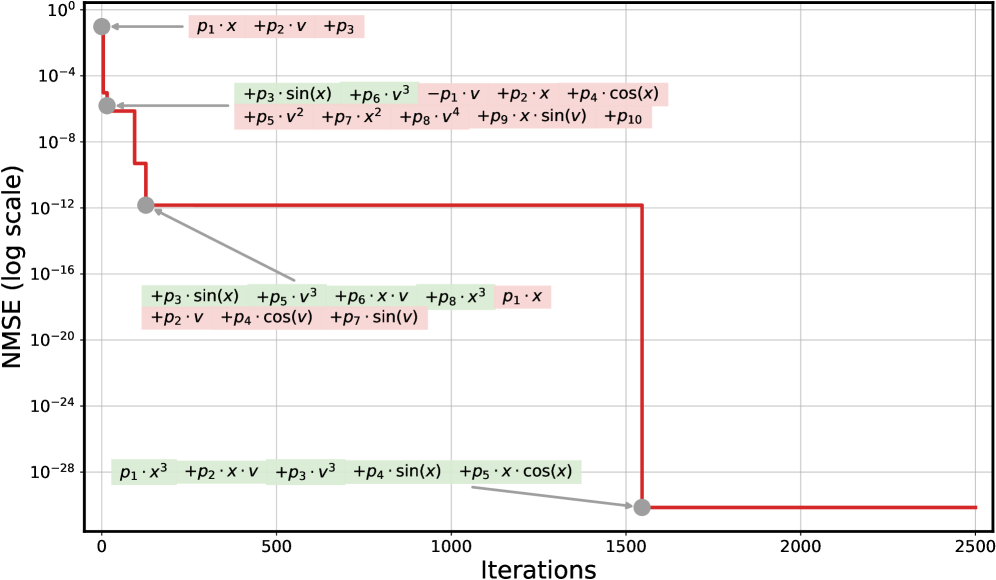
### Visual Description
## Chart: NMSE vs. Iterations for Model Complexity
### Overview
This chart displays the Negative Mean Squared Error (NMSE) on a logarithmic scale as a function of the number of iterations. The chart illustrates how the NMSE decreases with increasing model complexity (number of terms in the polynomial) and iteration count. Multiple lines represent different polynomial models, showing their convergence behavior.
### Components/Axes
* **X-axis:** Iterations (ranging from 0 to approximately 2500).
* **Y-axis:** NMSE (log scale, ranging from 10<sup>-30</sup> to 10<sup>0</sup>).
* **Lines:** Four distinct lines representing different polynomial models.
* **Markers:** Circular markers are placed at specific points along each line, likely indicating key stages or evaluations.
* **Legend:** Located in the top-left corner, labeling each line with its corresponding polynomial expression.
### Detailed Analysis
The chart shows four lines, each representing a polynomial model with increasing complexity.
* **Line 1 (Gray):** p<sub>1</sub>⋅x + p<sub>2</sub>⋅v + p<sub>3</sub>. This line starts at approximately NMSE = 6.0 x 10<sup>-7</sup> at iteration 0. It decreases steadily until approximately iteration 1500, where it drops sharply to approximately 2.0 x 10<sup>-25</sup>. The trend is initially a gentle slope downwards, followed by a near-vertical drop.
* **Line 2 (Orange):** +p<sub>3</sub>⋅sin(x) + p<sub>6</sub>⋅v<sup>3</sup> - p<sub>1</sub>⋅v + p<sub>2</sub>⋅x + p<sub>4</sub>⋅cos(x) + p<sub>5</sub>⋅v<sup>2</sup> + p<sub>7</sub>⋅x<sup>2</sup> + p<sub>8</sub>⋅v<sup>4</sup> + p<sub>9</sub>⋅x⋅sin(v) + p<sub>10</sub>. This line begins at approximately NMSE = 2.0 x 10<sup>-8</sup> at iteration 0. It decreases slowly until approximately iteration 1500, where it drops sharply to approximately 2.0 x 10<sup>-25</sup>. The trend is a very gradual decline, followed by a steep drop.
* **Line 3 (Green):** +p<sub>3</sub>⋅sin(x) + p<sub>6</sub>⋅v<sup>3</sup> + p<sub>2</sub>⋅x + p<sub>4</sub>⋅cos(v) + p<sub>7</sub>⋅sin(v). This line starts at approximately NMSE = 2.0 x 10<sup>-16</sup> at iteration 0. It remains relatively flat until approximately iteration 1500, where it drops to approximately 2.0 x 10<sup>-25</sup>. The trend is almost horizontal, then a sharp decline.
* **Line 4 (Red):** p<sub>1</sub>⋅x<sup>3</sup> + p<sub>2</sub>⋅x⋅v + p<sub>3</sub>⋅v<sup>3</sup> + p<sub>4</sub>⋅sin(x) + p<sub>5</sub>⋅x⋅cos(x). This line begins at approximately NMSE = 2.0 x 10<sup>-25</sup> at iteration 0. It remains flat until approximately iteration 1500, where it drops to approximately 2.0 x 10<sup>-25</sup>. The trend is flat.
The markers on the gray line are located at approximately:
* (0, 6.0 x 10<sup>-7</sup>)
* (500, 5.0 x 10<sup>-10</sup>)
* (1000, 2.0 x 10<sup>-12</sup>)
* (1500, 2.0 x 10<sup>-25</sup>)
The markers on the orange line are located at approximately:
* (0, 2.0 x 10<sup>-8</sup>)
* (500, 1.0 x 10<sup>-10</sup>)
* (1000, 2.0 x 10<sup>-12</sup>)
* (1500, 2.0 x 10<sup>-25</sup>)
The markers on the green line are located at approximately:
* (0, 2.0 x 10<sup>-16</sup>)
* (500, 2.0 x 10<sup>-16</sup>)
* (1000, 2.0 x 10<sup>-16</sup>)
* (1500, 2.0 x 10<sup>-25</sup>)
The markers on the red line are located at approximately:
* (0, 2.0 x 10<sup>-25</sup>)
* (500, 2.0 x 10<sup>-25</sup>)
* (1000, 2.0 x 10<sup>-25</sup>)
* (1500, 2.0 x 10<sup>-25</sup>)
### Key Observations
* All models converge to a similar NMSE value (approximately 2.0 x 10<sup>-25</sup>) after 1500 iterations.
* The simplest model (gray line) shows the most significant initial improvement in NMSE.
* The more complex models (orange, green, and red lines) start with lower NMSE values and exhibit slower initial improvement.
* The sharp drop in NMSE around iteration 1500 suggests a significant change in the optimization process or a convergence point.
* The red line remains constant throughout the iterations, indicating it has already reached its optimal performance.
### Interpretation
The chart demonstrates the relationship between model complexity, iteration count, and prediction error (NMSE). Initially, increasing model complexity leads to lower error, but beyond a certain point, adding more terms does not significantly improve performance. The convergence around iteration 1500 suggests that the optimization algorithm has reached a stable state for all models. The fact that all models converge to the same NMSE value indicates that the data may not require a highly complex model to achieve good performance. The red line's constant NMSE suggests it may have already reached its maximum representational capacity or is overfitting. This chart is likely used to evaluate the effectiveness of different polynomial models in fitting a given dataset and to determine the optimal level of model complexity. The logarithmic scale on the Y-axis emphasizes the large range of NMSE values and highlights the substantial error reduction achieved by the models.
</details>
(a) PiT-PO.
<details>
<summary>x14.png Details</summary>
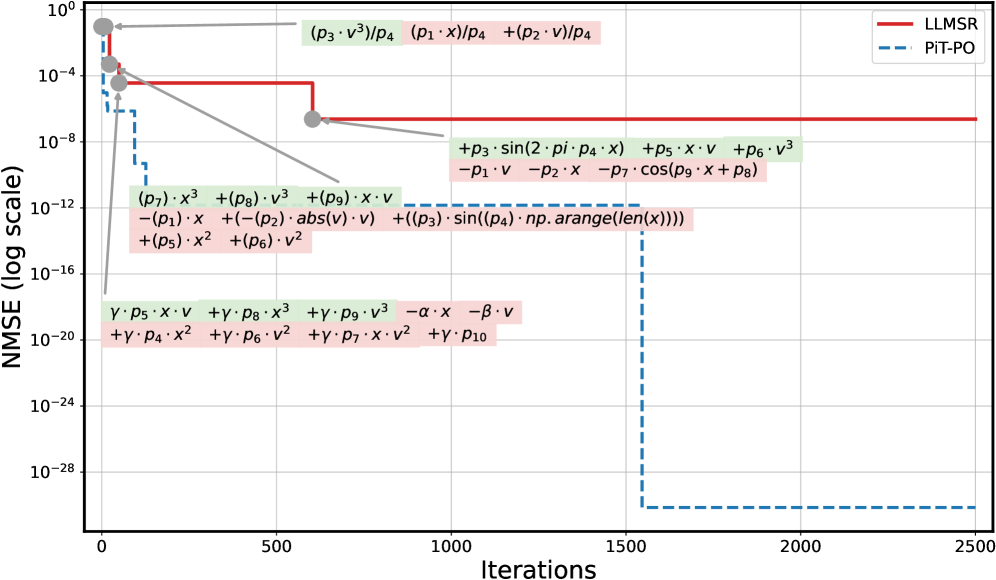
### Visual Description
## Chart: NMSE vs. Iterations for LLMSR and PIT-PO
### Overview
This chart displays the Normalized Mean Squared Error (NMSE) on a logarithmic scale against the number of iterations for two algorithms: LLMSR (Least-squares Locally Linear Model Shrinkage Regression) and PIT-PO (likely an abbreviation for a specific optimization method). The chart aims to demonstrate the convergence of these algorithms, showing how the error decreases with increasing iterations.
### Components/Axes
* **X-axis:** Iterations (linear scale, ranging from 0 to 2500)
* **Y-axis:** NMSE (log scale, ranging from 10<sup>-24</sup> to 10<sup>0</sup>)
* **Legend:**
* LLMSR (solid black line)
* PIT-PO (dashed red line)
* **Annotations:** Equations are displayed above the lines, presumably representing the error function or update rule for each algorithm.
### Detailed Analysis
**LLMSR (Black Solid Line):**
The LLMSR line starts at approximately 8.0 x 10<sup>-8</sup> NMSE at 0 iterations. It exhibits a steep downward slope initially, decreasing rapidly to approximately 1.5 x 10<sup>-12</sup> NMSE at around 500 iterations. The slope becomes less steep between 500 and 1500 iterations, reaching a plateau around 2.0 x 10<sup>-20</sup> NMSE at 1500 iterations. After 1500 iterations, the line shows a sudden drop to approximately 1.0 x 10<sup>-24</sup> NMSE at 1525 iterations, and remains relatively constant thereafter.
**PIT-PO (Red Dashed Line):**
The PIT-PO line begins at approximately 8.0 x 10<sup>-8</sup> NMSE at 0 iterations. It initially decreases at a moderate rate, reaching approximately 1.0 x 10<sup>-10</sup> NMSE at around 500 iterations. Between 500 and 1500 iterations, the line remains relatively flat, fluctuating around 1.0 x 10<sup>-10</sup> NMSE. At approximately 1500 iterations, the line experiences a sharp drop to approximately 1.0 x 10<sup>-24</sup> NMSE, and remains constant thereafter.
**Annotations (Equations):**
* **LLMSR Equation:** (p<sub>3</sub>⋅v<sup>3</sup>)/p<sub>4</sub> - (p<sub>1</sub>⋅x)/p<sub>4</sub> + (p<sub>2</sub>⋅v)/p<sub>4</sub>
* **PIT-PO Equation:** +p<sub>3</sub>⋅sin(2⋅pi⋅x) + p<sub>5</sub>⋅v + p<sub>6</sub>⋅v<sup>3</sup> - p<sub>1</sub>⋅v - p<sub>2</sub>⋅x - p<sub>7</sub>⋅cos(p<sub>9</sub>⋅x + p<sub>8</sub>)
* **LLMSR Equation:** (p<sub>7</sub>⋅x<sup>3</sup> + (p<sub>8</sub>⋅v<sup>3</sup> + (p<sub>9</sub>⋅x⋅v - (p<sub>1</sub>)⋅x +(-(p<sub>2</sub>)⋅abs(v)⋅v) + ((p<sub>3</sub>)⋅sin(p<sub>4</sub>⋅np.arange(len(x)))) + (p<sub>5</sub>)⋅x<sup>2</sup> + (p<sub>6</sub>)⋅v<sup>2</sup>
* **PIT-PO Equation:** γ⋅p<sub>5</sub>⋅x⋅v + γ⋅p<sub>8</sub>⋅x<sup>3</sup> + γ⋅p<sub>9</sub>⋅v<sup>3</sup> - α⋅x - β⋅v + γ⋅p<sub>4</sub>⋅x<sup>2</sup> + γ⋅p<sub>6</sub>⋅v<sup>2</sup> + γ⋅p<sub>7</sub>⋅x⋅v<sup>2</sup> + γ⋅p<sub>10</sub>
### Key Observations
* Both algorithms converge to a very low NMSE (around 10<sup>-24</sup>).
* LLMSR initially converges faster than PIT-PO.
* PIT-PO exhibits a period of stagnation between 500 and 1500 iterations.
* Both algorithms show a sudden drop in NMSE around 1500 iterations, suggesting a significant improvement in the solution at that point.
* The equations provided are complex and likely represent the core calculations within each algorithm.
### Interpretation
The chart demonstrates the effectiveness of both LLMSR and PIT-PO in minimizing the NMSE. The logarithmic scale highlights the substantial reduction in error achieved by both methods. The initial faster convergence of LLMSR suggests it might be more efficient for the first few iterations. However, both algorithms ultimately reach a similar level of accuracy. The stagnation observed in PIT-PO between 500 and 1500 iterations could indicate a challenging region in the optimization landscape, or a need for parameter tuning. The sudden drop in NMSE around 1500 iterations for both algorithms could be due to a change in the optimization strategy, or the algorithm overcoming a local minimum. The provided equations offer insight into the mathematical foundations of each algorithm, but a deeper understanding would require further analysis of the parameters (p<sub>1</sub>, p<sub>2</sub>, etc.) and their roles in the calculations. The chart suggests that both algorithms are viable options for minimizing the NMSE, but their performance characteristics differ, and the choice between them might depend on the specific application and computational constraints.
</details>
(b) LLM-SR.
Figure 10. Iterative trajectories of PiT-PO and LLM-SR on Oscillator1 with Llama-3.1-8B-Instruct. The NMSE curves (log scale) report the best-so-far value over iterations. Annotated equation snapshots illustrate how the discovered structure evolves along the search trajectory. The green-shaded terms are correct, whereas the red-shaded terms are erroneous.
Figure 10 visualizes how symbolic structures evolve along the iterative search. For PiT-PO (Figure 10(a)), the trajectory reveals a clear process of progressively excluding spurious terms. Under the theorem-guided mathematical constraints and the token-aware policy update, tokens marked as redundant are assigned token-level penalties; as this penalty signal repeatedly accumulates across many sampled equations during optimization, such spurious components become progressively less preferred and are unlikely to reappear in later generations. As a result, the search does not merely fit numerically; instead, it repeatedly transitions to cleaner structures, and each step-wise NMSE drop aligns with the removal of nuisance terms and a move toward a more parsimonious functional form. This “error-term elimination” behavior effectively shrinks the search space and substantially strengthens the exploration capability.
In contrast, LLM-SR (Figure 10(b)) exhibits much weaker structural correction. Since its guidance is primarily prompt-level rather than parameter-level, terms that appear early—even if structurally incorrect—tend to persist and continue influencing subsequent proposals. Consequently, the search frequently enters a plateau where NMSE stops improving meaningfully: the method keeps revisiting or retaining earlier spurious components instead of systematically pruning them, indicating a stagnation regime with limited ability to reach the correct structure.
## Appendix D Datasets
### D.1. LLM-SR Suite
We use the LLM-SR Suite, which consists of four standard scientific equation discovery tasks spanning physics, biology, and materials science. The suite includes two nonlinear oscillator systems (Oscillation 1/2), an E. coli growth model (E. coli Growth), and a temperature-dependent stress–strain dataset (Stress–Strain). The first three tasks are dynamical systems and are simulated over a fixed time horizon, while the last task is a static constitutive relation evaluated on experimental measurements.
#### D.1.1. Oscillation 1 (Nonlinear Oscillator)
Nonlinear oscillators with dissipative and non-polynomial couplings provide a demanding test for recovering interacting terms from trajectories. We define the state as $x(t)$ and $v(t)=\dot{x}(t)$ , and simulate the following system:
| | $\displaystyle\dot{x}$ | $\displaystyle=v,$ | |
| --- | --- | --- | --- |
We set $F=0.8$ , $\alpha=0.5$ , $\beta=0.2$ , $\gamma=0.5$ , and $\omega=1.0$ , with initial conditions $x(0)=0.5$ , $v(0)=0.5$ , and simulate over $t\in[0,50]$ .
#### D.1.2. Oscillation 2 (Nonlinear Oscillator)
The second oscillator introduces explicit time forcing and an exponential nonlinearity, leading to a different coupling pattern between $x$ and $v$ :
| | $\displaystyle\dot{x}$ | $\displaystyle=v,$ | |
| --- | --- | --- | --- |
We use $F=0.3$ , $\alpha=0.5$ , $\beta=1.0$ , $\delta=5.0$ , $\gamma=0.5$ , and $\omega=1.0$ . The initial conditions and simulation window are the same as Oscillation 1: $x(0)=0.5$ , $v(0)=0.5$ , $t\in[0,50]$ .
#### D.1.3. E. coli Growth (Bacterial Growth)
This task models the growth rate of E. coli as a product of multiplicative factors capturing distinct physiological effects from population size, nutrient availability, temperature, and acidity:
$$
\frac{dB}{dt}=f_{B}(B)\cdot f_{S}(S)\cdot f_{T}(T)\cdot f_{\mathrm{pH}}(\mathrm{pH}),
$$
where $B$ is bacterial density, $S$ is nutrient concentration, $T$ is temperature, and $\mathrm{pH}$ denotes acidity. In our benchmark, the explicit functional form is:
This specification combines saturation kinetics, sharp temperature modulation, and a structured pH response, yielding a challenging multivariate nonlinear discovery setting.
#### D.1.4. Stress–Strain (Material Stress Behavior)
To assess performance on experimental measurements, we consider tensile stress–strain data for Aluminum 6061-T651 collected at multiple temperatures (from room temperature up to $300^{\circ}\text{C}$ ). While no single exact governing equation is provided, a commonly used phenomenological approximation for temperature-dependent hardening is:
$$
\sigma=\left(A+B\varepsilon^{n}\right)\left(1-\left(\frac{T-T_{r}}{T_{m}-T_{r}}\right)^{m}\right).
$$
where $\sigma$ is stress, $\varepsilon$ is strain, $T$ is temperature, $T_{r}$ is a reference temperature, and $T_{m}$ is the melting temperature. The coefficients $A$ , $B$ , $n$ , and $m$ are fitted from data for the given alloy.
### D.2. LLM-SRBench
We additionally report results on LLM-SRBench, a benchmark specifically constructed to evaluate data-driven scientific equation discovery with LLMs while reducing the risk of trivial memorization of canonical textbook formulas. It contains 239 problems spanning four scientific domains (chemistry, biology, physics, and material science), and each task is packaged with (i) a short scientific context describing variables and the target quantity and (ii) tabular numerical samples used for discovery.
Dataset composition
LLM-SRBench is organized into two complementary subsets.
(1) LSR-Transform. This subset converts well-known physics equations into less-common but analytically equivalent forms by changing the prediction target. Concretely, starting from the original equation, one input variable is chosen as a pivot and promoted to the new target; the equation is then symbolically solved (e.g., via SymPy) for that pivot variable, yielding an alternative representation of the same underlying physical law. Only transformations that admit an analytic solution are retained, and the original datapoints are filtered to satisfy the valid domain of the transformed expression (e.g., avoiding invalid denominators or square-root domains). Finally, a new natural-language problem statement is generated to match the transformed input–output specification. This pipeline produces 111 transformed problems.
(2) LSR-Synth. This subset is designed to test discovery beyond memorized templates by composing equations from both known scientific terms and synthetic (novel yet plausible) terms. Candidate known/synthetic terms are proposed with an LLM given the domain context, combined into full equations, and then filtered through multiple checks: numerical solvability (e.g., via standard ODE solvers for dynamical systems), novelty assessment in context (using an LLM-based novelty evaluator), and expert plausibility validation. After passing these filters, datapoints are generated for each approved equation, yielding 128 synthetic problems across the same four domains.
Symbolic Accuracy (SA)
Besides numeric fit metrics (e.g., $\mathrm{NMSE}$ and $\mathrm{Acc}_{all}(\tau)$ ), LLM-SRBench emphasizes symbolic correctness. Symbolic Accuracy (SA) is computed using an LLM-based equivalence judge (GPT-4o): given a predicted hypothesis and the ground-truth expression, the evaluator first normalizes both by removing extraneous text (e.g., comments) and replacing explicit numeric constants with placeholder parameters, then decides whether there exist parameter values that make the hypothesis mathematically equivalent to the ground truth. SA is the fraction (percentage) of problems judged equivalent under this protocol. The benchmark authors validated this evaluation by comparing against human judgments on 130 sampled problems and reported 94.6% agreement between GPT-4o and human evaluators.
## Appendix E Turbulence Task Setup: Periodic Hill Flow
### E.1. Dataset
We consider the periodic hill flow configuration with streamwise periodic boundary conditions and no-slip walls.
In this work, the features selected for the symbolic regression process are computed from the mean strain-rate ( $S$ ) and rotation-rate ( $R$ ) tensors. Following established practice, these tensors are normalized using local turbulence scales prior to invariant calculation to ensure well-conditioned inputs:
$$
\displaystyle S \displaystyle=\frac{1}{2}\,\frac{1}{\beta^{*}\omega}\Bigl[\nabla\boldsymbol{u}+(\nabla\boldsymbol{u})^{\mathsf{T}}\Bigr], \displaystyle R \displaystyle=\frac{1}{2}\,\frac{1}{\beta^{*}\omega}\Bigl[\nabla\boldsymbol{u}-(\nabla\boldsymbol{u})^{\mathsf{T}}\Bigr], \tag{13}
$$
where $(\cdot)^{\mathsf{T}}$ denotes matrix transposition. The invariants are then computed as $I_{1}=\mathrm{Tr}(S^{2})$ and $I_{2}=\mathrm{Tr}(R^{2})$ .
The dataset provides (i) two invariant features $(I_{1},I_{2})$ (stored in the Lambda columns), (ii) tensor bases $\{T^{m}\}_{m=1}^{3}$ (stored in the first three $3\times 3$ tensors in the Tensor columns), and (iii) the target anisotropy tensor field $b$ (and its nonlinear correction form used for training, denoted by $b^{\perp}$ ), together with auxiliary fields such as turbulent kinetic energy $k$ when needed by the evaluator.
#### E.1.1. Data Collection
DNS source.
The DNS data are obtained from the publicly available NASA turbulence modeling resource (Turbulence Modeling Resource / TMBWG) for periodic hills of parameterized geometries. https://tmbwg.github.io/turbmodels/Other_DNS_Data/parameterized_periodic_hills.html
RANS baseline generation (for non-DNS methods).
For all non-DNS methods, we compute baseline solutions using OpenFOAM with the same initial and boundary conditions as the DNS. Specifically, we employ the incompressible steady-state solver simpleFoam and apply the turbulence models considered in this paper (all belonging to RANS and its SST variants). The solver outputs, at each sampling point and converged iteration/time-step, the required physical quantities for subsequent post-processing.
DNS–RANS grid alignment and target construction.
To construct training targets consistent with the RANS discretization, we interpolate the DNS fields onto the RANS grid points. The interpolated DNS quantities are then post-processed in the CFD workflow (i.e., as offline preprocessing prior to training) to compute the Reynolds stress tensor, which is packaged into the dataset files as the supervision signal for training.
Feature & basis construction.
Following Pope’s tensor-basis expansion hypothesis, the Reynolds stress decomposition involves (i) tensor bases and (ii) scalar coefficient functions of invariant features. Since our final turbulence closure is embedded as a correction to the baseline kOmegaSST model, we use the physical fields produced by the baseline kOmegaSST computation for post-processing to obtain the input features and the tensor bases. These are stored in separate files (features and bases), while the learning target is defined to match the Reynolds-stress-related quantity described in Section E.2.
#### E.1.2. Pre-processing
We normalize the two invariant inputs prior to feeding them into the learned symbolic expressions:
$$
\tilde{I}=\tanh\!\left(\frac{I}{2.0}\right),\qquad\tilde{I}_{1}=\tanh\!\left(\frac{I_{1}}{2.0}\right),\quad\tilde{I}_{2}=\tanh\!\left(\frac{I_{2}}{2.0}\right), \tag{15}
$$
which maps the raw values into approximately $[-1,1]$ and improves numerical stability during parameter fitting.
### E.2. Casting Turbulence Closure as a Symbolic Regression Problem
Predicting scalar coefficients instead of tensor entries.
Rather than directly searching over tensor-valued symbolic forms, we let the LLM predict three scalar coefficient functions $(G^{1},G^{2},G^{3})$ , each parameterized as a symbolic expression of only two variables $(I_{1},I_{2})$ . The evaluator then reconstructs the predicted tensor via the (fixed) tensor bases:
$$
\hat{b}\;=\;G^{1}(I_{1},I_{2})\,T^{1}\;+\;G^{2}(I_{1},I_{2})\,T^{2}\;+\;G^{3}(I_{1},I_{2})\,T^{3}. \tag{16}
$$
This design compresses the LLM input variable dimension to two, significantly reducing the symbolic search space.
Scoring by MSE in the evaluator.
Given a candidate hypothesis, we compute the fitting score by the mean squared error between the target tensor and the reconstructed tensor. In the final evaluator, we additionally scale both prediction and target by the sample-wise $k$ before error aggregation:
$$
\hat{\tau}_{ij}(x_{n})=k(x_{n})\,\hat{b}_{ij}(x_{n}),\qquad\tau_{ij}(x_{n})=k(x_{n})\,b_{ij}(x_{n}), \tag{17}
$$
We compute the MSE only over a selected subset of tensor components. This is because the Reynolds-stress (and anisotropy) tensor is symmetric, so only six components are independent, and in the present periodic hill flow setting two off-plane shear components are typically several orders of magnitude smaller and are commonly neglected in practical turbulence modeling. Following this standard practice, we evaluate the error only on
$$
\Omega=\{(0,0),(0,1),(1,1),(2,2)\}, \tag{18}
$$
$$
\qquad\mathrm{MSE}=\frac{1}{|\Omega|}\sum_{(i,j)\in\Omega}\frac{1}{N}\sum_{n=1}^{N}\Bigl(\tau_{ij}(x_{n})-\hat{\tau}_{ij}(x_{n})\Bigr)^{2}. \tag{19}
$$
The resulting MSE is used as the numerical fitting score for the symbolic regression loop.
Prompt and fairness across methods.
We provide the same problem specification text (task description, variable definitions, and operator constraints) to both LLM-SR and PiT-PO to ensure a fair comparison. An example prompt is shown in Figure 11 – 13. In our current experiments, we also explicitly restrict the hypothesis space by asking the LLM to avoid trigonometric functions.
<details>
<summary>x15.png Details</summary>

### Visual Description
\n
## Text Document: Symbolic Regression for Periodic Hill Turbulence Modeling
### Overview
This document outlines the task of symbolic regression for modeling periodic hill turbulence. It details the problem setup, evaluation rules, and the tensor bases used in the regression process. The document primarily consists of text with mathematical formulations and descriptions of the underlying physical phenomena.
### Components/Axes
The document is structured as a series of paragraphs describing the task, the flow context, evaluation rules, and tensor bases. There are no explicit axes or charts. The document uses mathematical notation to define the hill profile and the anisotropy tensor.
### Detailed Analysis or Content Details
**Task:** Symbolic regression for periodic hill turbulence modeling. Learn three scalar functions G1(I1, I2), G2(I1, I2), G3(I1, I2).
**Flow case context (for description only):**
* Bottom wall profile:
* y(x) = a(1 + cos(πx / L)) for |x| ≤ L
* y(x) = 0 for |x| > L
* where *a* is the hill height (characteristic length h), and α = L/h controls hill steepness (training case: α = 0.8).
* Streamwise (x) direction uses periodic boundary conditions; top and bottom walls use no-slip boundary conditions.
* Reynolds number: Re\_h = U\_b h / ν = 5600.
**Evaluation rule:**
Given tensor bases T1, T2, T3 (each 3x3), the predicted anisotropy is:
b\_hat = G1 \* T1 + G2 \* T2 + G3 \* T3
The evaluator computes MSE between b\_hat and the target b.
**Tensor bases:** T1, T2, T3 (provided by the evaluator, do NOT compute them). For each sample n, the evaluator provides three 3x3 tensor bases (symmetric, traceless) constructed from the non-dimensionalized mean strain-rate tensor S and rotation tensor R:
* **T1 (linear strain basis):**
* T1 = S
* Interpretation: the Boussinesq/linear eddy-viscosity direction. It dominates in simple shear flows and acts as the baseline anisotropy response aligned with the mean strain.
* **T2 (strain-rotation coupling basis):**
* T2 = S @ R - R @ S
* Interpretation: captures the interaction between mean strain and mean rotation (curvature/turning, strong vortices). This term is crucial in separated flows, reattachment, and swirling/shear-layer regions where linear eddy-viscosity assumptions tend to fail.
* **T3 (quadratic strain nonlinearity basis):**
* T3 = S @ S - (1/3) \* tr(S @ S) \* I
* Interpretation: introduces nonlinear strain effects and normal-stress anisotropy beyond linear models. It helps represent differences among normal stress components and improves predictions in strongly anisotropic regions.
The predicted anisotropy is assembled as:
b\_hat = G1(I1, I2) \* T1 + G2(I1, I2) \* T2 + G3(I1, I2) \* T3
### Key Observations
The document focuses on defining the mathematical framework for modeling turbulence over periodic hills. The key components are the hill profile, the Reynolds number, the tensor bases (T1, T2, T3), and the scalar functions G1, G2, and G3 that need to be learned through symbolic regression. The tensor bases represent different aspects of the anisotropy, ranging from linear strain effects to nonlinear strain and strain-rotation coupling.
### Interpretation
This document describes a problem in computational fluid dynamics (CFD) where the goal is to model the anisotropy of Reynolds stresses in turbulent flow over periodic hills. The use of symbolic regression suggests that the researchers are attempting to discover analytical expressions for the scalar functions G1, G2, and G3 that best fit the observed data. The tensor bases T1, T2, and T3 provide a structured way to represent the anisotropy, and their interpretations highlight the physical mechanisms that contribute to it. The document emphasizes the limitations of linear eddy-viscosity models and the need for more sophisticated approaches that can capture nonlinear effects and strain-rotation coupling. The Reynolds number of 5600 indicates a relatively high Reynolds number flow, where turbulence is fully developed. The hill steepness parameter α = 0.8 suggests a moderately steep hill, which is likely to induce significant flow separation and turbulence. The overall goal is to develop a more accurate and physically meaningful model for turbulence over periodic hills, which can be used for predicting the flow behavior and optimizing the design of engineering systems.
</details>
Figure 11. Problem Specification.
<details>
<summary>x16.png Details</summary>

### Visual Description
\n
## Code Block: Python Function Definitions
### Overview
The image contains a block of Python code defining functions related to evaluating data and minimizing a loss function, likely within a machine learning or scientific computing context. The code appears to be focused on reconstructing an anisotropy tensor based on input tensors and scalar coefficients.
### Components/Axes
The code consists of function definitions with comments explaining the purpose of each section. There are no axes or charts in this image. The code uses the `numpy` library (indicated by `np.mean` and `np.ones`).
### Detailed Analysis or Content Details
The code defines two functions: `evaluate` and `loss`.
**`evaluate(data, equation)` function:**
- Takes `data` and `equation` as input.
- Extracts `I1`, `I2` from `data['inputs']`.
- Extracts `T1`, `T2`, `T3` from `data['tensors']`.
- Extracts `b_true` from `data['outputs']`.
**`loss(params)` function:**
- Takes `params` as input.
- Predicts scalar coefficients `G = [G1, G2, G3]` using the `equation` function, which returns a shape (N, 3) based on the `params`. `G_pred` is the result of this prediction.
- Reconstructs the predicted anisotropy tensor `b_hat` using the formula: `b_hat = G1*T1 + G2*T2 + G3*T3`. The code expands this using `G_pred` and indexing to calculate each component.
- Computes the Mean Squared Error (MSE) between the predicted tensor `b_hat` and the target tensor `b_true`.
- Returns the MSE.
**Initialization and Minimization:**
- Initializes parameters with `np.ones(30)`, creating a numpy array of 30 ones.
- Uses the `minimize` function (likely from `scipy.optimize`) to minimize the `loss` function, starting with the `initial_params` and using the BFGS method.
- Returns the function value (`result.fun`) after minimization.
### Key Observations
- The code is well-commented, explaining the purpose of each step.
- The `equation` function is passed as an argument to `evaluate`, suggesting a flexible framework where different equations can be used.
- The loss function is based on the Mean Squared Error, a common metric for regression problems.
- The use of `BFGS` suggests an optimization problem with a smooth objective function.
- The code assumes the existence of `data` dictionary with keys 'inputs', 'tensors', and 'outputs'.
### Interpretation
The code implements a process for estimating parameters that reconstruct an anisotropy tensor from input tensors. The `loss` function quantifies the difference between the reconstructed tensor and a target tensor, and the `minimize` function finds the parameters that minimize this difference. This suggests a model fitting or parameter estimation task, potentially in materials science, image processing, or other fields where anisotropy is important. The use of numpy and scipy indicates a numerical computation focus. The code is designed to be modular, with the equation being passed as a parameter, allowing for different models to be tested.
</details>
Figure 12. Evaluation and Optimization.
<details>
<summary>x17.png Details</summary>

### Visual Description
\n
## Code Block: Python Function Definition
### Overview
The image presents a Python code block defining a function named `equation`. This function takes three NumPy arrays as input: `I1`, `I2`, and `params`, and returns a NumPy array. The function's purpose is to predict three scalar coefficients (G1, G2, G3) based on tensor bases.
### Components/Axes
The code block consists of:
- A function definition line: `def equation(I1: np.ndarray, I2: np.ndarray, params: np.ndarray) -> np.ndarray:`
- A docstring explaining the function's purpose, arguments, and return value.
- Three blocks of code, each calculating one of the coefficients (G1, G2, G3).
- A return statement that stacks the calculated coefficients into a single NumPy array.
### Detailed Analysis or Content Details
The function `equation` is defined as follows:
```python
def equation(I1: np.ndarray, I2: np.ndarray, params: np.ndarray) -> np.ndarray:
"""
Predict three scalar coefficients G1, G2, G3 for tensor bases.
Args:
I1, I2: numpy arrays of shape (N), invariants.
params: numpy array of free constants.
Returns:
numpy array of shape (N, 3), columns are G1, G2, G3.
"""
# G1 block: params[0:10]
g1 = (
params[0] * I1 +
params[1] * I2 +
params[2]
)
# G2 block: params[10:20]
g2 = (
params[10] * I1 +
params[11] * I2 +
params[12]
)
# G3 block: params[20:30]
g3 = (
params[20] * I1 +
params[21] * I2 +
params[22]
)
return np.stack((g1, g2, g3), axis=1)
```
- **Input Arguments:**
- `I1`: A NumPy array of shape (N).
- `I2`: A NumPy array of shape (N).
- `params`: A NumPy array containing free constants.
- **Coefficient Calculation:**
- `g1` is calculated using `params[0]`, `params[1]`, and `params[2]`.
- `g2` is calculated using `params[10]`, `params[11]`, and `params[12]`.
- `g3` is calculated using `params[20]`, `params[21]`, and `params[22]`.
- **Return Value:**
- A NumPy array of shape (N, 3), where each column represents one of the coefficients (G1, G2, G3).
### Key Observations
- The function uses a linear combination of `I1` and `I2` to calculate each coefficient.
- The `params` array is partitioned into three blocks: `params[0:10]` for G1, `params[10:20]` for G2, and `params[20:30]` for G3.
- The function utilizes NumPy's `np.stack` function to combine the calculated coefficients into a single array.
### Interpretation
The code implements a linear regression-like model to predict three coefficients (G1, G2, G3) based on two input arrays (I1, I2) and a set of parameters. The parameters are organized into blocks, suggesting that each block corresponds to a specific coefficient. The function's structure suggests that it could be used as part of a larger system for tensor decomposition or model fitting. The use of NumPy arrays indicates that the function is designed for efficient numerical computation. The docstring provides clear documentation of the function's purpose, arguments, and return value, which is good programming practice.
</details>
Figure 13. Equation Program Example.
### E.3. Back to Turbulence Modeling
From discovered expressions to a deployable closure
After obtaining an explicit expression for the Reynolds-stress-related term from symbolic regression, we rewrite the discovered formula into an OpenFOAM-readable form.
Embedding into OpenFOAM
We embed the resulting closure into the existing kOmegaSST implementation by replacing the routine that evaluates the Reynolds stress (or its modeled contribution) with the discovered expression, thereby forming an optimized turbulence model within the RANS framework.
Verification against DNS
We then run RANS simulations with the modified model under the same configuration as the baseline and compare the resulting flow statistics against the DNS reference data. This completes the end-to-end turbulence modeling and validation pipeline considered in this work.
### E.4. Domain-Specific Constraints for Turbulence Modeling
To encode expert knowledge as inductive biases, we augment the reward with a domain-specific penalty term:
$$
P_{\text{domain}}\;=\;\sum_{j=1}^{4}w_{j}\,P_{\text{domain}}^{(j)}, \tag{20}
$$
where each $P_{\text{domain}}^{(j)}\geq 0$ quantifies violations of the $j$ -th turbulence constraint.
(1) Realizability.
A physically admissible Reynolds stress tensor must be symmetric and positive semi-definite (PSD), which implies that all eigenvalues are nonnegative. Non-realizable predictions are penalized by
$$
P_{\text{domain}}^{(1)}\;=\;\frac{1}{N}\sum_{n=1}^{N}\max\Bigl(0,\,-\lambda_{\min}(\tau(x_{n}))\Bigr), \tag{21}
$$
where $\lambda_{\min}(\cdot)$ denotes the smallest eigenvalue. This PSD requirement is a standard realizability condition for Reynolds stresses (Pope, 2000).
(2) Boundary-condition consistency.
At a no-slip wall, the Reynolds stress tensor must decay to zero. Accordingly, non-vanishing stresses in a near-wall band $\mathcal{W}=\{x:y(x)<y_{0}\}$ are penalized as
$$
P_{\text{domain}}^{(2)}\;=\;\frac{1}{|\mathcal{W}|}\sum_{x_{n}\in\mathcal{W}}\|\tau(x_{n})\|_{F}, \tag{22}
$$
where $y(x)$ is the wall distance field. In practice, $y(x)$ can be computed and exported using standard CFD post-processing tools (OpenFOAM wall-distance utilities such as wallDist and checkMesh-writeAllFields) (Monkewitz, 2021).
(3) Asymptotic scaling in the viscous sublayer.
Near-wall Taylor expansions under the no-slip constraint imply that $u^{\prime}=O(y)$ and, for incompressible flow, $v^{\prime}=O(y^{2})$ . Consequently, the Reynolds shear stress $-\overline{u^{\prime}v^{\prime}}$ exhibits cubic leading-order scaling, $O(y^{3})$ , in the viscous sublayer (Tennekes and Lumley, 1972). This constraint is enforced by evaluating the slope of $\log|\tau_{xy}^{+}|$ with respect to $\log(y^{+})$ over the viscous sublayer range $\mathcal{V}=\{x:y^{+}(x)\in[y_{1}^{+},y_{2}^{+}]\}$ :
$$
p\;=\;\mathrm{slope}\bigl(\log|\tau_{xy}^{+}|\;\text{vs}\;\log(y^{+})\bigr),\qquad P_{\text{domain}}^{(3)}\;=\;|p-3|. \tag{23}
$$
Here $y^{+}=y\,u_{\tau}/\nu$ is the wall unit, with friction velocity $u_{\tau}=\sqrt{\tau_{w}/\rho}$ . In CFD, $y^{+}$ and $\tau_{w}$ can be obtained through standard post-processing utilities (OpenFOAM yPlus and wallShearStress).
(4) Energy consistency.
Energy consistency is enforced by constraining the turbulent kinetic energy production implied by the predicted stress:
$$
P_{k}(x)\;=\;-\tau_{ij}(x)\,\frac{\partial U_{i}}{\partial x_{j}}(x), \tag{24}
$$
and penalizing mismatches with the reference production $P_{k}^{\text{ref}}$ (computed from DNS-aligned fields) via
$$
P_{\text{domain}}^{(4)}\;=\;\mathrm{MSE}\bigl(P_{k},\,P_{k}^{\text{ref}}\bigr). \tag{25}
$$
The definition in Eq. (24) is the standard production term in the TKE budget (Pope, 2000).
Efficient evaluation via elite-CFD refinement (multi-fidelity constraint scheduling).
Constraints (2)–(4) require additional CFD-derived fields (wall distance $y$ , wall shear stress $\tau_{w}$ and the associated $y^{+}$ , and mean velocity gradients $\nabla U$ ), which can be expensive to obtain at each candidate-evaluation step. To keep the symbolic regression loop efficient, we adopt a two-stage evaluation schedule.
Stage-A (inexpensive screening, applied to all candidates) evaluates the data-fitting score (MSE) together with realizability $P_{\text{domain}}^{(1)}$ using only the predicted stress tensor.
Stage-B (expensive physics checks, triggered by an improving best MSE) is activated when a candidate in the current batch achieves a new best MSE value relative to all previously evaluated candidates. In this case, the candidate is treated as an elite solution, and CFD-based post-processing is performed to obtain the required auxiliary fields, compute $P_{\text{domain}}^{(2)}$ – $P_{\text{domain}}^{(4)}$ , and conduct an additional online fine-tuning step using the full domain-specific penalty. All non-elite candidates are trained only with the realizability constraint, whereas elite candidates receive the full turbulence constraint suite after CFD evaluation. This design enables rigorous physics enforcement without incurring prohibitive CFD costs during exploration.