# Statistical Learning Analysis of Physics-Informed Neural Networks
**Authors**: David Barajas-Solano
> David.Barajas-Solano@pnnl.gov
Abstract
We study the training and performance of physics-informed learning for initial and boundary value problems (IBVP) with physics-informed neural networks (PINNs) from a statistical learning perspective. Specifically, we restrict ourselves to parameterizations with hard initial and boundary condition constraints and reformulate the problem of estimating PINN parameters as a statistical learning problem. From this perspective, the physics penalty on the IBVP residuals can be better understood not as a regularizing term bus as an infinite source of indirect data, and the learning process as fitting the PINN distribution of residuals $p(y\mid x,t,w)q(x,t)$ to the true data-generating distribution $\delta(0)q(x,t)$ by minimizing the Kullback-Leibler divergence between the true and PINN distributions. Furthermore, this analysis show that physics-informed learning with PINNs is a singular learning problem, and we employ singular learning theory tools, namely the so-called Local Learning Coefficient [7] to analyze the estimates of PINN parameters obtained via stochastic optimization for a heat equation IBVP. Finally, we discuss implications of this analysis on the quantification of predictive uncertainty of PINNs and the extrapolation capacity of PINNs.
keywords: Physics-informed learning , physics-informed neural networks , statistical learning , singular learning theory
Pacific Northwest National Laboratory
1 Introduction
Physics-informed machine learning with physics-informed neural networks (PINNs) (see [6] for a comprehensive overview) has been widely advanced and successfully employed in recent years to solve approximate the solution of initial and boundary value problems (IBVPs) in computational mathematics. The central mechanism of physics-informed learning is model parameter identification by minimizing “physics penalty” terms penalizing the governing equation and initial and boundary condition residuals predicted by a given set of model parameters. While various significant longstanding issues of the use PINNs for physics-informed learning such as the spectral bias have been addressed, it remains active area of research, and our understanding of various features of the problem such as the loss landscape, and the extrapolation capacity of PINN models, remains incomplete.
It has been widely recognized that traditional statistical tools are inadequate for analyzing deep learning. Watanabe’s “singular learning theory” (SLT) (see [13] for a review) recognizes that this is due to the singular character of learning with deep learning models and aims to address these limitations. The development of efficient deep learning and Bayesian inference tools and frameworks [3, 15, 2, 1] has enabled the application of SLT to analyzing practical applications of deep learning, leading to various insights on the features of the loss landscape, the generalization capacity of the model, and the training process [7, 4, 10].
Given that physics-informed learning with PINNs is also singular due to its use of neural networks as the backbone architecture, we propose in this work to use SLT to analyze the training and performance of PINNs. The physics penalty in physics-informed learning precludes an out-of-the-box application of SLT, so we first reformulate the PINN learning problem as a statistical learning problem by restricting our attention to PINN parameterizations with hard initial and boundary condition constraints. Employing this formulation, it can be seen that the PINN learning problem is equivalent to the statistical learning problem of fitting the PINN distribution of governing equation residuals $p(y\mid x,t,w)q(x,t)$ to the true data-generating distribution $\delta(0)q(x,t)$ corresponding to zero residuals $y(x,t)\equiv 0$ for all $(x,t)∈\Omega×(0,T]$ , by minimizing the Kullback-Leibler divergence between the true and PINN distributions. We argue this implies that the physics penalty in physics-informed learning can be better understood as an infinite source of indirect data (with the residuals interpreted as indirect observations given that they are a function of the PINN prediction) as opposed to as a regularizing term as it is commonly interpreted in the literature. Furthermore, we employ the SLT’s so-called Local Learning Coefficient (LLC) to study the features of the PINN loss for a heat equation IBVP.
This manuscript is structured as follows: In section 2 we reformulate and analyze the physics-informed learning problem as a singular statistical learning problem. In section 3 we discuss the numerical approximation of the LLC and compute it for a heat equation IBVP. Finally, further implications of this analysis are discussed in section 4.
2 Statistical learning interpretation of physics-informed learning for IBVPs
Consider the well-posed boundary value problem
$$
\displaystyle\mathcal{L}(u(x,t),x,t) \displaystyle=0 \displaystyle\forall x\in\Omega, \displaystyle t\in(0,T], \displaystyle\mathcal{B}(u(x,t),x,t) \displaystyle=0 \displaystyle\forall x\in\partial\Omega, \displaystyle t\in[0,T], \displaystyle u(x,0) \displaystyle=u_{0}(x) \displaystyle\forall x\in\Omega, \tag{1}
$$
where the $\Omega$ denotes the IBVP’s spatial domain, $∂\Omega$ the domain’s boundary, $T$ the maximum simulation time, and $u(x,t)$ the IBVP’s solution. In the PINN framework, we approximate the solution $u(x,t)$ using an overparameterized deep learning-based model $u(x,t,w)$ , usually a neural network model $\text{NN}_{w}(x,t)$ , where $w∈ W$ denotes the model parameters and $W⊂\mathbb{R}^{d}$ denotes the parameter space. For simplicity, we assume that the model $u(x,t,w)$ is constructed in such a way that the initial and boundary conditions are identically satisfied for any model parameter, that is,
| | $\displaystyle\mathcal{B}(u(x,t,w),x,t)=0\quad∀ x∈∂\Omega,\ t∈(0,T],\ w∈ W,$ | |
| --- | --- | --- |
via hard-constraints parameterizations (e.g., [8, 9]). The model parameters are then identified by minimizing the squared residuals of the governing equation evaluated at $n$ “residual evaluation” points $\{x_{i},t_{i}\}^{n}_{i=1}$ , that is, by minimizing the loss
$$
L^{\text{PINN}}_{n}(w)\coloneqq\frac{1}{n}\sum^{n}_{i=1}\left[\mathcal{L}(u(x_{i},t_{i},w),x_{i},t_{i})\right]^{2}, \tag{4}
$$
In the physics-informed-only limit, the PINN framework aims to approximate the solution without using direct observations of the solution, that is, without using labeled data pairs of the form $(x_{i},t_{i},u(x_{i},t_{i}))$ , which leads many authors to refer to it as an “unsupervised” learning framework. An alternative view of the PINN learning problem is that it actually does use labeled data, but of the form $(x_{i},t_{i},\mathcal{L}(u(x_{i},t_{i}),x_{i},t_{i})\equiv 0)$ , as the PINN formulation encourages the residuals of the parameterized model, $\mathcal{L}(u(x,t,w),x,t)$ , to be as close to zero as possible. Crucially, “residual” data of this form is “infinite” in the sense that such pairs can be constructed for all $x∈\Omega$ , $t∈(0,T]$ . In fact, the loss 4 can be interpreted as a sample approximation of the volume-and-time-averaged loss $L^{\text{PINN}}\coloneqq∈t_{\Omega×(0,T]}\left[\mathcal{L}(u(x,t),x,t)\right]^{2}q(x,t)\,\mathrm{d}x\mathrm{d}t$ for a certain weighting function $q(x,t)$ .
We can then establish a parallel with statistical learning theory. Namely, we can introduce a density over the inputs, $q(x,t)$ , and the true data-generating distribution $q(x,t,y)=\delta(0)q(x,t)$ , where $y$ denotes the equation residuals. For a given error model $p(y\mid\mathcal{L}(u(x,t,w),x,t))$ , the PINN model induces a conditional distribution of residuals $p(y\mid x,w)$ . For example, for a Gaussian error model $p(y\mid\mathcal{L}(u(x,t,w),x,t))=\mathcal{N}(y\mid 0,\sigma^{2})$ , the conditional distribution of residuals is of the form
$$
p(y\mid x,t,w)\propto\exp\left\{-\frac{1}{2\sigma^{2}}\left[\mathcal{L}(u(x,t,w),x,t)\right]^{2}\right\}. \tag{5}
$$
The PINN framework can then be understood as aiming to fit the model $p(x,t,y\mid w)\coloneqq p(y\mid x,t,w)q(x,t)$ to the data-generating distribution by minimizing the sample negative log-likelihood $L_{n}(w)$ of the training dataset $\{(x_{i},t_{i}y_{i}\equiv 0)\}^{n}_{i=1}$ drawn i.i.d. from $q(x,t,y)$ , with
$$
L_{n}(w)\coloneqq-\frac{1}{n}\sum^{n}_{i=1}\log p(y_{i}\equiv 0\mid x_{i},t_{i},w). \tag{6}
$$
Associated to this negative log-likelihood we also have the population negative log-likelihood $L(w)\coloneqq-\mathbb{E}_{q(x,t,y)}\log p(y\mid x,t,w)$ . Note that for the Gaussian error model introduced above we have that $L^{\text{PINN}}_{n}(w)=2\sigma^{2}L_{n}(w)$ up to an additive constant, so that learning by minimizing the PINN loss 4 and the negative log-likelihood 6 is equivalent. Furthermore, also note that $L^{\text{PINN}}(w)=2\sigma^{2}L(w)$ up to an additive constant if the volume-and-time averaging function is taken to be the same as the data-generating distribution of the inputs, $q(x,t)$ . Therefore, it follows that PINN training is equivalent to minimizing the Kullback-Leibler divergence between the true and model distributions of the residuals.
This equivalent reformulation of the PINN learning problem allows us to employ the tools of statistical learning theory to analyze the training of PINNs. Specifically, we observe that, because of the neural network-based forward model $\text{NN}_{w}(x)$ is a singular model, the PINN model $p(y\mid x,t,w)$ is also singular, that is, the parameter-to-distribution map $w\mapsto p(x,t,y\mid w)$ is not one-to-one, and the Fisher information matrix $I(w)\coloneqq\mathbb{E}_{p(x,t,y\mid w)}∇∇_{w}\,\log p(x,t,y\mid w)$ is not everywhere positive definite in $W$ . As a consequence, the PINN loss 4 is not characterized by a discrete, measure-zero, possibly infinite set of local minima that can be locally approximated quadratically. Instead, the PINN loss landscape, just as is generally the case for deep learning loss landscapes, is characterized by “flat” minima, crucially even in the limit of the number of residual evaluation points $n→∞$ . This feature allows us to better understand the role of the residual data in physics-informed learning.
Namely, it has been widely observed that the solutions to the PINN problem $\operatorname*{arg\,min}_{w}L^{\text{PINN}}_{n}(w)$ for a given loss tolerance $\epsilon$ are not unique even for large number of residual evaluation points, and regardless of the scheme used to generate such points (uniformly distributed, low-discrepancy sequences such as latin hypercube sampling, random sampling, etc.), while on the other hand it has also been observed that increasing $n$ seems to “regularize” or “smoothen” the PINN loss landscape (e.g., [11]). Although one may interpret this observed smoothing as similar to the regularizing effect of explicit regularization (e.g., $\ell_{2}$ regularization or the use of a proper prior on the parameters), in fact increasing $n$ doesn’t sharpen the loss landscape into locally quadratic local minima as explicit regularization does. Therefore, residual data (and correspondingly, the physics penalty) in physics-informed learning should not be understood as a regularizer but simply as a source of indirect data.
As estimates of the solution to the PINN problem are not unique, it is necessary to characterize them through other means besides their loss value. In this work we employ the SLT-based LLC metric, which aims to characterize the “flatness” of the loss landscape of singular models. For completeness, we reproduce the definition of the LLC (introduced in [7]) in the sequel. Let $w^{\star}$ denote a local minima and $B(w^{\star})$ a sufficiently small neighborhood of $w^{\star}$ such that $L(\omega)≥ L(\omega^{\star})$ ; furthermore, let $B(w^{\star},\epsilon)\coloneqq\{w∈ B(w^{\star})\mid L(w)-L(w^{\star})<\epsilon\}$ denote the subset of the neighborhood of $w^{\star}$ with loss within a certain tolerance $\epsilon$ , and $V(\epsilon)$ denote the volume of such subset, that is,
$$
V(\epsilon)\coloneqq\operatorname{vol}(B(w^{\star},\epsilon))=\int_{B(w^{\star},\epsilon)}\mathrm{d}w.
$$
Then, the LLC is defined as the rational number $\lambda(w^{\star})$ such that the volume scales as $V(\epsilon)\propto\exp\{\lambda(w^{\star})\}$ . The LLC can be interpreted as a measure of flatness because it corresponds to the rate at which the volume of solutions within the tolerance $\epsilon$ decreases as $\epsilon→ 0$ , with smaller LLC values correspond to slower loss of volume.
As its name implies, the LLC is the local analog to the SLT (global) learning coefficient $\lambda$ , which figures in the asymptotic expansion of the free energy of a singular statistical model. Specifically $\lambda(w^{\star})\equiv\lambda$ if $w^{\star}$ is a global minimum and $B(w^{\star})$ is taken to be the entire parameter space $W$ . The free energy $F_{n}$ is defined as
$$
F_{n}\coloneqq-\log\int\exp\{-nL_{n}(w)\}\varphi(w)\,\mathrm{d}w,
$$
where $\varphi(w)$ is a prior on the model parameters. Then, according to the SLT, the free energy for a singular model has the asymptotic expansion [13]
$$
F_{n}=nL_{n}(w_{0})+\lambda\log n+O_{P}(\log\log n),\quad n\to\infty,
$$
where $w_{0}$ is the parameter vector that minimizes the Kullback-Leibler divergence between the true data-generating distribution and the estimated distribution $p(y\mid x,t,w)q(x,t)$ , whereas for a regular model it has the expansion
$$
F_{n}=nL_{n}(\hat{w})+\frac{d}{2}\log n+O_{P}(1),\quad n\to\infty, \tag{1}
$$
where $\hat{w}$ is the maximum likelihood estimator of the model parameters, and $d$ the number of model parameters. The learning coefficient can then be interpreted as measuring model complexity, as it is the singular model analog to a regular model’s model complexity $d/2$ . Therefore, by computing the LLC around different estimates to the PINN solution we can gain some insight into the PINN loss landscape.
3 Numerical estimation of the local learning coefficient
3.1 MCMC-based local learning coefficient estimator
While there may exist other approaches to estimating the LLC (namely, via Imai’s estimator [5] or Watanabe’s estimator [14] of the real log-canonical threshold), in this work we choose to employ the estimation framework introduced in [7], which we briefly describe as follows. Let $B_{\gamma}(w^{\star})$ denote a small ball of radius $\gamma$ centered around $w^{\star}$ . We introduce the local to $w^{\star}$ analog to the free energy and its asymptotic expansion:
$$
\begin{split}F_{n}(B_{\gamma}(w^{\star}))&=-\log\int_{B_{\gamma}(w^{\star})}\exp\{-nL_{n}(w)\}\varphi(w)\,\mathrm{d}w\\
&=nL_{n}(w^{\star})+\lambda(w^{\star})\log n+O_{P}(\log\log n).\end{split} \tag{7}
$$
We then substitute the hard restriction in the integral above to $B_{\gamma}(w^{\star})$ with a softer Gaussian weighting around $w^{\star}$ with covariance $\gamma^{-1}I_{d}$ , where $\gamma>0$ is a scale parameter. Let $\mathbb{E}_{w\mid w^{\star},\beta,\gamma}[·]$ denote the expectation with respect to the tempered distribution
$$
p(w\mid w^{\star},\beta,\gamma)\propto\exp\left\{-n\beta L_{n}(w)-\frac{\gamma}{2}\|w-w^{\star}\|^{2}_{2}\right\}
$$
with inverse temperature $\beta\equiv 1/\log n$ ; then, $\mathbb{E}_{w\mid w^{\star},\beta,\gamma}[nL_{n}(w)]$ is a good estimator of the partition function of this tempered distribution, which is, depending on the choice $\gamma$ , an approximation to the local free energy. Substituting this approximation into 7, we obtain the estimator
$$
\hat{\lambda}_{\gamma}(w^{\star})\coloneqq n\beta\left[\mathbb{E}_{w\mid w^{\star},\beta,\gamma}[L_{n}(w)]-L_{n}(w^{\star})\right]. \tag{8}
$$
In this work we compute this estimate by computing samples $w\mid w^{\star},\beta,\gamma$ via Markov chain Monte Carlo (MCMC) simulation. Specifically, we employ the No-U Turns Sampler (NUTS) algorithm [3]. Furthermore, to evaluate $L_{n}(w)$ we choose a fixed set of $n$ residual evaluation points chosen randomly from $q(x,t)$ , with $n$ large enough that we can employ the asymptotic expansion 7 and that eq. 8 is valid. Note that $\hat{\lambda}_{\gamma}(w^{\star})$ is strictly positive as $w^{\star}$ is a local minima of $L_{n}(w)$ , so that $\mathbb{E}_{w\mid w^{\star},\beta,\gamma}[L_{n}(w)]-L_{n}(w^{\star})≥ 0$ . In fact, negative estimates of the LLC imply that the MCMC chain was able to visit a significant volume of parameter space for which $L_{n}(w)<L_{n}(w^{\star})$ , which means that $w^{\star}$ is not an actual minima of $L_{n}(w)$ . Given that minimization of the PINN loss via stochastic optimization algorithms is not guaranteed to produce exact minima, in practice it may be possible to obtain negative estimates of the LLC.
3.2 Numerical experiments
To analyze the PINN loss landscape for a concrete case, we compute and analyze the LLC for the following physics-informed learning problem: We consider the heat equation IBVP
| | $\displaystyle∂_{t}u-∂_{xx}u$ | $\displaystyle=f(x,t)$ | $\displaystyle∀ x∈(0,2),\ t∈(0,2],$ | |
| --- | --- | --- | --- | --- |
where $f(x,t)$ is chosen so that the solution to the IBVP is
$$
u(x,t)=\exp^{-t}\sin(\pi x).
$$
To exactly enforce initial and boundary conditions, we employ the hard-constraints parameterization of the PINN model
$$
u(x,t,w)=\sin(\pi x)+tx(x-1)\text{NN}_{w}(x,t). \tag{9}
$$
The neural network model $\text{NN}_{w}$ is taken to be a multi-layer perceptron (MLP) with 2 hidden layers of $50$ units each. This model has a total number of parameters $d=20,601.$ This relatively small model ensures that the wall-clock time of computing multiple MCMC chains is reasonable. The model eq. 9 is implemented using Flax NNX library [2], and MCMC sampling is performed using the BlackJAX library [1].
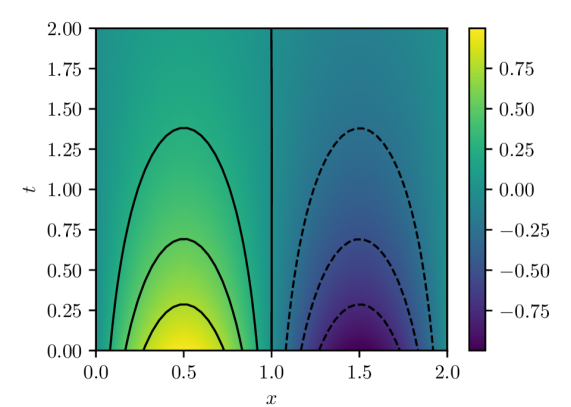
The parameters of the model are estimated by minimizing the PINN loss eq. 4 using the Adam stochastic optimization algorithm for $50,000$ iterations. At each iteration, a new batch of residual evaluation points is drawn from $q(x,t)$ and used to evaluate the PINN loss and its gradient. Note that these batches of residual evaluation points are distinct from the set used to evaluate $L_{n}(w)$ in eq. 8, which is fixed for a given experiment. Figure 1 shows the PINN solution $u(x,t,w^{\star})$ for $w^{\star}$ estimated using a batch size of $32$ and learning rate of $1× 10^{-4}$ . This solution has an associated PINN loss of $1.854× 10^{-5}$ , which, although small, shows that the true data-generating distribution $\delta(0)q(x,t)$ is not realizable within the model class 9 with the MLP architecture indicated above due to the finite capacity of the class.
<details>
<summary>x1.png Details</summary>
### Visual Description
## Heatmap: Spatio-temporal Data Visualization
### Overview
The image is a heatmap visualizing data across two dimensions, x and t, with a color gradient representing the data values. The heatmap is divided into two regions by a vertical line at x=1.0. The left region shows positive values, while the right region shows negative values. Contour lines are overlaid on the heatmap to highlight specific value levels.
### Components/Axes
* **X-axis:** Horizontal axis labeled "x", ranging from 0.0 to 2.0 in increments of 0.5.
* **Y-axis:** Vertical axis labeled "t", ranging from 0.0 to 2.0 in increments of 0.25.
* **Colorbar:** Located on the right side of the image, representing the data values. The colorbar ranges from -0.75 (dark purple) to 0.75 (yellow). Intermediate values are represented by shades of green and blue.
* **Contour Lines:** Solid black lines on the left side and dashed black lines on the right side, indicating specific data value levels.
### Detailed Analysis
* **Left Region (x: 0.0 to 1.0):**
* The heatmap shows a region of positive values, with the highest values (yellow) concentrated around x=0.5 and t=0.0.
* The values decrease as t increases, forming a bell-shaped distribution.
* Three solid black contour lines are present, indicating specific value levels. The innermost contour line is closest to the yellow region, and the outermost contour line is furthest.
* The approximate t values for the peaks of the three solid contour lines are 0.2, 0.7, and 1.3.
* **Right Region (x: 1.0 to 2.0):**
* The heatmap shows a region of negative values, with the lowest values (dark purple) concentrated around x=1.5 and t=0.0.
* The values increase (become less negative) as t increases, forming an inverted bell-shaped distribution.
* Three dashed black contour lines are present, indicating specific value levels. The innermost contour line is closest to the dark purple region, and the outermost contour line is furthest.
* The approximate t values for the peaks of the three dashed contour lines are 0.2, 0.7, and 1.3.
### Key Observations
* The data is symmetrical around x=1.0, with the left region showing positive values and the right region showing negative values.
* The magnitude of the values decreases as t increases in both regions.
* The contour lines provide a visual representation of the data distribution, highlighting the regions of highest and lowest values.
### Interpretation
The heatmap visualizes a spatio-temporal dataset where the values are positive on one side of a boundary (x=1.0) and negative on the other side. The data exhibits a symmetrical distribution, with the magnitude of the values decreasing over time (t). The contour lines help to identify the regions where the data values are most concentrated. The data could represent a physical phenomenon, such as the diffusion of a substance from two sources with opposite concentrations, or a mathematical function with similar properties.
</details>
(a) PINN solution $u(x,t,w^{\star})$
<details>
<summary>x2.png Details</summary>
### Visual Description
## Heatmap: Time vs. X with Contour Lines
### Overview
The image is a heatmap representing data over a two-dimensional space defined by the variables 'x' and 't'. The heatmap uses a color gradient to indicate the magnitude of the data, ranging from dark purple (lowest values) to bright yellow (highest values). Superimposed on the heatmap are black contour lines, both solid and dashed, indicating specific data levels.
### Components/Axes
* **X-axis:** Labeled 'x', ranges from 0.0 to 2.0 in increments of 0.5.
* **Y-axis:** Labeled 't', ranges from 0.0 to 2.0 in increments of 0.25.
* **Colorbar:** Located on the right side of the image, it represents the data values corresponding to the heatmap colors. The colorbar ranges from -0.002 (dark purple) to 0.001 (bright yellow), with 0.000 (green) in the middle.
* **Contour Lines:** Black lines overlaid on the heatmap. Solid lines represent positive values, while dashed lines represent negative values.
### Detailed Analysis
* **Heatmap Color Distribution:**
* The lower-left corner (x=0, t=0) is greenish, indicating values around -0.001.
* The upper-left corner (x=0, t=2) is dark purple, indicating values around -0.002.
* The center of the heatmap, around (x=1.5, t=1.5), is bright yellow, indicating values around 0.001.
* The lower-right corner (x=2, t=0) is greenish, indicating values around -0.001.
* **Contour Lines:**
* A dashed contour line is present in the upper-left corner, indicating a negative value region. It starts at approximately (x=0.25, t=2.0) and curves down to approximately (x=0.5, t=1.25).
* A solid contour line starts at approximately (x=0.75, t=0.0) and curves upwards, passing through approximately (x=1.0, t=0.75), then curves back down to approximately (x=1.75, t=2.0).
* Another solid contour line forms a closed loop around the yellow region, centered approximately at (x=1.5, t=1.5).
* **Specific Data Points (Approximations based on color):**
* (x=0.0, t=0.0): Value ≈ -0.001
* (x=2.0, t=2.0): Value ≈ -0.001
* (x=1.5, t=1.5): Value ≈ 0.001
* (x=0.0, t=2.0): Value ≈ -0.002
### Key Observations
* There is a clear positive peak in the center of the heatmap.
* The negative values are concentrated in the upper-left corner.
* The contour lines provide a visual representation of the data's gradient and highlight regions of constant value.
### Interpretation
The heatmap visualizes a two-dimensional function or field, where the color intensity represents the magnitude of the function at each point (x, t). The contour lines further delineate regions of equal value, providing a more detailed understanding of the function's behavior. The data suggests a localized maximum around (x=1.5, t=1.5) and a localized minimum in the upper-left corner. The relationship between x and t influences the data values, with certain combinations resulting in higher or lower magnitudes. The presence of both positive and negative values indicates that the function is not strictly positive or negative across the entire domain.
</details>
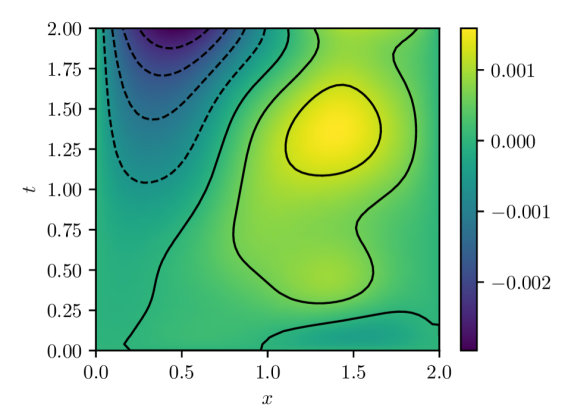
(b) Pointwise errors $u(x,t)-u(x,t,w^{\star})$
Figure 1: PINN solution and estimation error computed with batch size of $32$ and Adam learning rate of $1× 10^{-4}.$
LLC estimates are computed for batch sizes of $8$ , $16$ , and $32$ , and for Adam learning rates $\eta$ of $1× 10^{-3}$ and $1× 10^{-4}.$ Furthermore, for all experiments we compute the LLC estimate using a fixed set of $256$ residual evaluation points drawn randomly from $q(x,t).$ To compute the LLC, we employ the Gaussian likelihood of residuals (5). As such, the LLC estimate depends not only on the choice of $\gamma$ but also on the standard deviation $\sigma$ . For each experiment we generate 2 MCMC chains and each chain is initialized with an warmup adaptation interval.
To inform the choice of hyperparameters of the LLC estimator we make the following observations: First, both hyperparameters control the volume of the local neighborhood employed to estimate the LLC, in the case of $\gamma$ by expanding or contracting the prior centered around $w^{\star}$ , and in the case of $\sigma$ by increasing or decreasing the range of values $L_{n}(w)$ can take within the local neighborhood. Second, given that the true data-generating distribution is not realizable, the PINN loss will always be non-zero. Therefore, for the true zero residuals to be resolvable with non-trivial probability by the PINN model we must choose $\sigma$ larger than the average optimal PINN residuals (the squared root of the optimal PINN loss.)
Given that $\gamma=1$ controls the volume of the local neighborhood, we take the value of $\gamma=1$ For the architecture described above, we find that the optimal PINN loss is $O(10^{-5})$ , so that $\sigma$ must be chosen to be $3× 10^{-3}$ or larger. Given that $\sigma$ also controls the volume of the local neighborhood, we take the conservative value of $\sigma=1$ . It is not entirely obvious whether this conservative choice overestimates or underestimates the LLC. Numerical experiments not presented in this work showed that the small choice $\sigma=1× 10^{-2}$ led often to negative LLC estimates and therefore was rejected. LLC values estimated for $\sigma=1× 10^{-1}$ where consistently slightly larger than those for $\sigma=1$ , which indicates that the model class of PINN models that are more physics-respecting has slightly more complexity than a less physics-respecting class. Nevertheless, given that the difference in LLC estimates is small we proceeded with the conservative choice.
<details>
<summary>x3.png Details</summary>
### Visual Description
## Chart/Diagram Type: Combined Line Charts
### Overview
The image presents two line charts side-by-side. The left chart displays "Train and test loss" on a logarithmic scale against "Iteration". It compares the performance of two learning rates (η = 1 x 10^-4 and η = 1 x 10^-3) for both training and testing data. The right chart shows "Local learning coefficient" against "Iteration" for the same two learning rates.
### Components/Axes
**Left Chart:**
* **Y-axis:** "Train and test loss" (logarithmic scale). Axis markers are 10^0, 10^-2, 10^-4, and 10^-6.
* **X-axis:** "Iteration". Axis markers are 0, 10000, 20000, 30000, 40000, and 50000.
* **Legend (top-right):**
* Blue line: "η = 1 x 10^-4 train"
* Orange line: "η = 1 x 10^-4 test"
* Green line: "η = 1 x 10^-3 train"
* Red line: "η = 1 x 10^-3 test"
**Right Chart:**
* **Y-axis:** "Local learning coefficient". Axis markers are 7.0, 7.5, 8.0, 8.5, 9.0, 9.5, and 10.0.
* **X-axis:** "Iteration". Axis markers are 10000, 20000, 30000, 40000, and 50000.
* **Legend (bottom-left):**
* Blue line with 'x' markers: "η = 1 x 10^-4"
* Orange dashed line with 'x' markers: "η = 1 x 10^-3"
* Shaded regions around each line indicate uncertainty.
### Detailed Analysis
**Left Chart (Train and Test Loss):**
* **η = 1 x 10^-4 train (Blue):** Starts at approximately 2, decreases sharply until around iteration 10000, then decreases gradually until iteration 50000, reaching approximately 0.01.
* **η = 1 x 10^-4 test (Orange):** Starts at approximately 0.1, decreases until around iteration 10000, then fluctuates around 10^-4 until iteration 50000.
* **η = 1 x 10^-3 train (Green):** Starts at approximately 0.3, decreases sharply until around iteration 10000, then decreases gradually until iteration 50000, reaching approximately 0.005.
* **η = 1 x 10^-3 test (Red):** Starts at approximately 0.01, fluctuates around 10^-5 until iteration 50000.
**Right Chart (Local Learning Coefficient):**
* **η = 1 x 10^-4 (Blue):** Starts at approximately 7.0 at iteration 10000, increases to approximately 9.0 at iteration 20000, then remains relatively stable around 9.3 until iteration 50000.
* **η = 1 x 10^-3 (Orange):** Starts at approximately 9.3 at iteration 10000, increases slightly to approximately 9.6 at iteration 30000, then decreases slightly to approximately 9.4 at iteration 50000.
### Key Observations
* In the left chart, the training loss (blue and green lines) decreases more consistently than the test loss (orange and red lines).
* The test loss for η = 1 x 10^-3 (red line) fluctuates significantly more than the test loss for η = 1 x 10^-4 (orange line).
* In the right chart, the local learning coefficient for η = 1 x 10^-4 (blue line) increases sharply initially, while the local learning coefficient for η = 1 x 10^-3 (orange line) remains relatively stable.
* The shaded regions in the right chart indicate the uncertainty or variability in the local learning coefficient.
### Interpretation
The charts compare the performance of two different learning rates (η = 1 x 10^-4 and η = 1 x 10^-3) during the training of a model. The left chart shows that both learning rates result in decreasing training loss, but the test loss behaves differently. The higher learning rate (η = 1 x 10^-3) leads to more fluctuation in the test loss, suggesting potential overfitting. The right chart shows the local learning coefficient, which provides insight into how the model is adapting during training. The lower learning rate (η = 1 x 10^-4) initially requires a larger adjustment in the learning coefficient, while the higher learning rate (η = 1 x 10^-3) maintains a relatively stable learning coefficient. The shaded regions indicate the variability in the local learning coefficient, which could be due to factors such as the batch size or the specific data used during training.
</details>
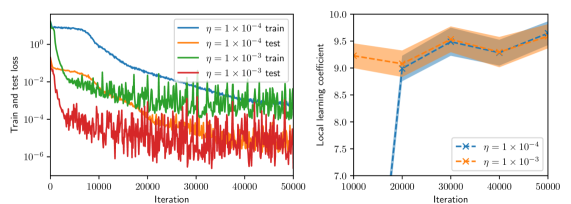
(a) Batch size $=8$
<details>
<summary>x4.png Details</summary>
### Visual Description
## Chart Type: Combined Line Charts
### Overview
The image presents two line charts side-by-side. The left chart displays the train and test loss as a function of iteration for two different learning rates (η = 1 x 10^-4 and η = 1 x 10^-3). The right chart shows the local learning coefficient as a function of iteration for the same two learning rates.
### Components/Axes
**Left Chart:**
* **Title:** Train and test loss
* **X-axis:** Iteration (ranging from 0 to 50000)
* **Y-axis:** Train and test loss (logarithmic scale from 10^-6 to 10^0)
* **Legend (top-right):**
* Blue line: η = 1 x 10^-4 train
* Orange line: η = 1 x 10^-4 test
* Green line: η = 1 x 10^-3 train
* Red line: η = 1 x 10^-3 test
**Right Chart:**
* **Title:** Local learning coefficient
* **X-axis:** Iteration (ranging from 10000 to 50000)
* **Y-axis:** Local learning coefficient (linear scale from 7.0 to 10.0)
* **Legend (bottom-right):**
* Blue dashed line with 'x' markers: η = 1 x 10^-4
* Orange dashed line with 'x' markers: η = 1 x 10^-3
* Shaded regions around each line indicate uncertainty or variance.
### Detailed Analysis
**Left Chart (Train and Test Loss):**
* **η = 1 x 10^-4 train (Blue):** Starts at approximately 1.5, decreases rapidly until around iteration 10000, then decreases more gradually to approximately 0.01 at iteration 50000.
* **η = 1 x 10^-4 test (Orange):** Starts at approximately 0.1, decreases to approximately 0.001 around iteration 20000, and then fluctuates around that value until iteration 50000.
* **η = 1 x 10^-3 train (Green):** Starts at approximately 0.1, decreases rapidly until around iteration 5000, then fluctuates between 0.0001 and 0.1 until iteration 50000.
* **η = 1 x 10^-3 test (Red):** Starts at approximately 0.001, fluctuates significantly between 0.000001 and 0.01 until iteration 50000.
**Right Chart (Local Learning Coefficient):**
* **η = 1 x 10^-4 (Blue, dashed with 'x'):** Starts at approximately 7.3 at iteration 10000, increases to approximately 9.3 at iteration 20000, and then remains relatively stable around 9.3 until iteration 50000.
* **η = 1 x 10^-3 (Orange, dashed with 'x'):** Starts at approximately 9.3 at iteration 10000, increases slightly to approximately 9.5 at iteration 20000, and then remains relatively stable around 9.5 until iteration 50000.
### Key Observations
* The training loss decreases for both learning rates, but the lower learning rate (1 x 10^-4) results in a smoother and more consistent decrease.
* The test loss for the lower learning rate also decreases and stabilizes, while the test loss for the higher learning rate fluctuates significantly, suggesting overfitting.
* The local learning coefficient is higher for the higher learning rate (1 x 10^-3) and remains relatively stable after the initial increase.
### Interpretation
The charts demonstrate the impact of different learning rates on the training and testing performance of a model. A lower learning rate (η = 1 x 10^-4) leads to more stable training and testing loss, suggesting better generalization. A higher learning rate (η = 1 x 10^-3) results in lower training loss, but significant fluctuations in the test loss, indicating overfitting. The local learning coefficient reflects these trends, with the higher learning rate exhibiting a higher coefficient. The shaded regions in the right chart likely represent the variance or standard deviation of the local learning coefficient across multiple runs or batches, providing an indication of the stability and reliability of the learning process.
</details>
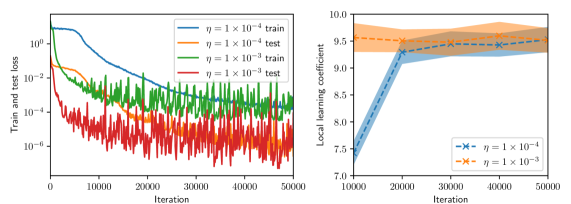
(b) Batch size $=16$
<details>
<summary>x5.png Details</summary>
### Visual Description
## Chart Type: Combined Line Charts
### Overview
The image contains two line charts side-by-side. The left chart displays "Train and test loss" on a logarithmic scale against "Iteration". It shows four data series representing training and testing loss for two different learning rates (η = 1 x 10^-4 and η = 1 x 10^-3). The right chart displays "Local learning coefficient" against "Iteration" for the same two learning rates, with shaded regions indicating variability.
### Components/Axes
**Left Chart:**
* **Y-axis:** "Train and test loss" (logarithmic scale). Axis markers: 10^-7, 10^-5, 10^-3, 10^-1, 10^1.
* **X-axis:** "Iteration". Axis markers: 0, 10000, 20000, 30000, 40000, 50000.
* **Legend (top-right):**
* Blue: "η = 1 x 10^-4 train"
* Orange: "η = 1 x 10^-4 test"
* Green: "η = 1 x 10^-3 train"
* Red: "η = 1 x 10^-3 test"
**Right Chart:**
* **Y-axis:** "Local learning coefficient". Axis markers: 7.0, 7.5, 8.0, 8.5, 9.0, 9.5, 10.0.
* **X-axis:** "Iteration". Axis markers: 10000, 20000, 30000, 40000, 50000.
* **Legend (bottom-left):**
* Blue with 'x' markers: "η = 1 x 10^-4"
* Orange with 'x' markers: "η = 1 x 10^-3"
### Detailed Analysis
**Left Chart (Train and test loss):**
* **η = 1 x 10^-4 train (Blue):** Starts at approximately 10^1, decreases rapidly until around iteration 10000, then decreases more gradually, reaching approximately 10^-3 at iteration 50000.
* **η = 1 x 10^-4 test (Orange):** Starts at approximately 10^-1, decreases rapidly until around iteration 10000, then remains relatively constant around 10^-5.
* **η = 1 x 10^-3 train (Green):** Starts at approximately 10^-1, decreases rapidly until around iteration 10000, then fluctuates around 10^-3.
* **η = 1 x 10^-3 test (Red):** Starts around 10^-4, fluctuates significantly between 10^-5 and 10^-7.
**Right Chart (Local learning coefficient):**
* **η = 1 x 10^-4 (Blue):** Starts at approximately 8.9 at iteration 10000, increases to approximately 9.7 at iteration 30000, then decreases slightly to approximately 9.4 at iteration 50000. The shaded region indicates a variability of approximately +/- 0.3.
* **η = 1 x 10^-3 (Orange):** Starts at approximately 9.4 at iteration 10000, increases slightly to approximately 9.4 at iteration 20000, then decreases slightly to approximately 9.4 at iteration 50000. The shaded region indicates a variability of approximately +/- 0.3.
### Key Observations
* In the left chart, the training loss decreases more consistently than the testing loss, especially for η = 1 x 10^-3.
* The testing loss for η = 1 x 10^-3 fluctuates significantly, suggesting potential overfitting.
* In the right chart, the local learning coefficient for η = 1 x 10^-3 is consistently higher than for η = 1 x 10^-4.
* The variability in the local learning coefficient is similar for both learning rates.
### Interpretation
The charts suggest that a learning rate of η = 1 x 10^-4 results in more stable training and testing loss compared to η = 1 x 10^-3. The higher learning rate (η = 1 x 10^-3) leads to significant fluctuations in the testing loss, indicating potential overfitting. The local learning coefficient is higher for η = 1 x 10^-3, which might contribute to the observed instability. The data suggests that a lower learning rate might be preferable for this particular model and dataset to achieve better generalization performance.
</details>
(c) Batch size $=32$
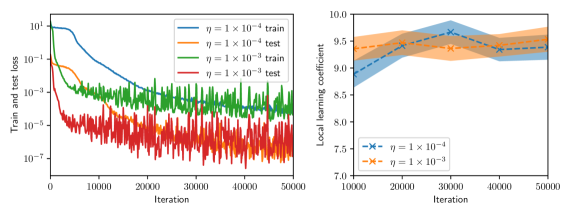
Figure 2: Training and test histories (left, shown every 100 iterations), and LLC estimates with 95% confidence intervals (right, shown every 1,000 iterations) for various values of batch size and Adam learning rate.
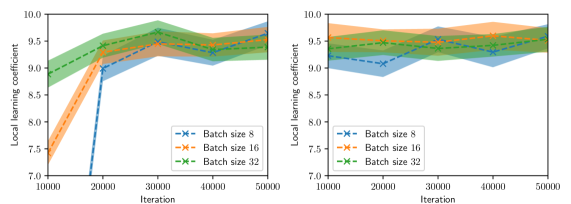
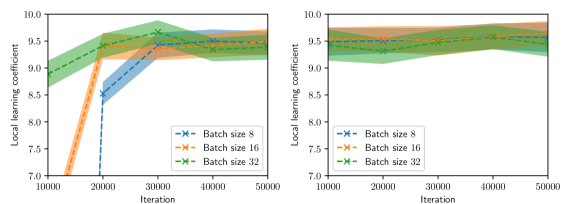
Figure 2 summarizes the results of the computational experiments. The left panels show the history of the training and test losses every $100$ iterations. Here, the test loss is the average of $(u(x,t,w)-u(x,t))^{2}$ evaluated over a $51× 51$ uniform grid of $(x,t)$ pairs. The right panels show the LLC estimates computed every $10,000$ iterations. Here, care must be taken when interpreting the smaller LLC estimates seen for earlier iterations. For early iterations, the parameter estimate $w_{t}$ is not a minima; therefore, the MCMC chains cover regions of parameter space for which $L_{n}(w)<L_{n}(w_{t})$ , leading to small and possibly negative values. Modulo this observation, it can be seen that across all experiments (batch size and learning rate values), the estimated LLC is $\hat{\lambda}(w^{\star})≈ 9.5$ . Additional experiments conducted by varying the pseudo-random number generator seed used to initialize $w$ also lead to the same estimate of the LLC (see fig. 3.)
This number is remarkable for two reasons: First, despite the estimate of the local minima $w^{\star}$ being vastly different across experiments, the LLC estimate is very consistent. Second, $\hat{\lambda}(w^{\star})$ is significantly smaller than the number of parameters $d=20,601$ of the PINN model. If the PINN model were regular, then it would have a much smaller $O(10)$ number of parameters. These results indicate that the PINN solution corresponds to a very flat region of parameter space, and different choices of initializations and stochastic optimization hyperparameters lead to the same region. As seen in figs. 2 and 3, the lower learning rate of $1× 10^{-4}$ , due to reduced stochastic optimization noise, takes a larger number of iterations to arrive at this region than the larger learning rate of $1× 10^{-3}.$
<details>
<summary>x6.png Details</summary>
### Visual Description
## Chart: Local Learning Coefficient vs. Iteration for Different Batch Sizes
### Overview
The image contains two line charts comparing the local learning coefficient against the iteration number for three different batch sizes (8, 16, and 32). The charts show the performance of each batch size over iterations, with shaded regions indicating variability or confidence intervals. The chart on the left shows the performance starting at iteration 10000, while the chart on the right shows the performance starting at iteration 0.
### Components/Axes
* **X-axis (Iteration):**
* Ranges from 10000 to 50000 in both charts.
* Markers at 10000, 20000, 30000, 40000, and 50000.
* **Y-axis (Local learning coefficient):**
* Ranges from 7.0 to 10.0 in both charts.
* Markers at 7.0, 7.5, 8.0, 8.5, 9.0, 9.5, and 10.0.
* **Legend (bottom):**
* Batch size 8 (blue line with 'x' markers)
* Batch size 16 (orange line with 'x' markers)
* Batch size 32 (green line with 'x' markers)
### Detailed Analysis
**Left Chart:**
* **Batch size 8 (blue):**
* Starts at approximately 7.0 at iteration 10000.
* Increases sharply to approximately 9.0 at iteration 20000.
* Fluctuates between 9.0 and 9.5 from iteration 20000 to 50000.
* **Batch size 16 (orange):**
* Starts at approximately 7.5 at iteration 10000.
* Increases to approximately 9.0 at iteration 20000.
* Reaches approximately 9.5 at iteration 30000.
* Fluctuates between 9.2 and 9.7 from iteration 30000 to 50000.
* **Batch size 32 (green):**
* Starts at approximately 9.0 at iteration 10000.
* Increases to approximately 9.5 at iteration 20000.
* Reaches approximately 9.7 at iteration 30000.
* Fluctuates between 9.3 and 9.7 from iteration 30000 to 50000.
**Right Chart:**
* **Batch size 8 (blue):**
* Starts at approximately 9.0 at iteration 10000.
* Fluctuates between 9.0 and 9.5 from iteration 10000 to 50000.
* **Batch size 16 (orange):**
* Starts at approximately 9.5 at iteration 10000.
* Fluctuates between 9.3 and 9.8 from iteration 10000 to 50000.
* **Batch size 32 (green):**
* Starts at approximately 9.5 at iteration 10000.
* Fluctuates between 9.3 and 9.7 from iteration 10000 to 50000.
### Key Observations
* In the left chart, batch sizes 16 and 32 start with lower local learning coefficients but quickly catch up to batch size 8.
* In the left chart, batch size 8 has a sharp increase in local learning coefficient between iterations 10000 and 20000.
* In the right chart, all batch sizes start with relatively high local learning coefficients.
* The shaded regions around each line indicate the variability or confidence interval for each batch size.
* The right chart shows the performance starting at iteration 0, while the left chart shows the performance starting at iteration 10000.
### Interpretation
The charts compare the performance of different batch sizes in terms of local learning coefficient over iterations. The left chart suggests that smaller batch sizes (8 and 16) initially have lower local learning coefficients but improve significantly over the first 20000 iterations. The right chart shows that when starting from iteration 0, all batch sizes have relatively high local learning coefficients. This suggests that the initial iterations are crucial for the smaller batch sizes to catch up in performance. The shaded regions indicate the variability in performance, which is relatively consistent across all batch sizes. Overall, the choice of batch size can impact the initial learning rate, but all batch sizes converge to similar performance levels after a certain number of iterations.
</details>
(a) First initialization
<details>
<summary>x7.png Details</summary>
### Visual Description
## Line Chart: Local Learning Coefficient vs. Iteration for Different Batch Sizes
### Overview
The image contains two line charts comparing the local learning coefficient against the iteration number for different batch sizes (8, 16, and 32). The charts show the performance of each batch size over a range of iterations, with shaded regions indicating variability. The left chart shows the initial learning phase, while the right chart shows a more stable, later phase.
### Components/Axes
* **Y-axis (Local learning coefficient):** Ranges from 7.0 to 10.0 in both charts, with tick marks at 7.0, 7.5, 8.0, 8.5, 9.0, 9.5, and 10.0.
* **X-axis (Iteration):** Ranges from 10000 to 50000 in both charts, with tick marks at 10000, 20000, 30000, 40000, and 50000.
* **Legend (bottom-left of the left chart, bottom-left of the right chart):**
* Blue line with 'x' markers: Batch size 8
* Orange line with 'x' markers: Batch size 16
* Green line with 'x' markers: Batch size 32
### Detailed Analysis
**Left Chart:**
* **Batch size 8 (Blue):** The line starts at approximately 7.0 at 10000 iterations and rises sharply to approximately 8.5 at 20000 iterations. It then continues to rise, reaching approximately 9.3 at 30000 iterations, and stabilizes around 9.5 at 40000 and 50000 iterations.
* **Batch size 16 (Orange):** The line starts at approximately 7.0 at 10000 iterations and rises sharply to approximately 9.3 at 20000 iterations. It then stabilizes around 9.4 at 30000, 40000 and 50000 iterations.
* **Batch size 32 (Green):** The line starts at approximately 8.8 at 10000 iterations and rises to approximately 9.6 at 30000 iterations. It then stabilizes around 9.5 at 40000 and 50000 iterations.
**Right Chart:**
* **Batch size 8 (Blue):** The line starts at approximately 9.3 at 10000 iterations and remains relatively stable around 9.5 at 20000, 30000, 40000 and 50000 iterations.
* **Batch size 16 (Orange):** The line starts at approximately 9.5 at 10000 iterations and remains relatively stable around 9.6 at 20000, 30000, 40000 and 50000 iterations.
* **Batch size 32 (Green):** The line starts at approximately 9.3 at 10000 iterations and remains relatively stable around 9.5 at 20000, 30000, 40000 and 50000 iterations.
### Key Observations
* In the left chart, batch sizes 8 and 16 show a significant initial increase in the local learning coefficient, while batch size 32 starts with a higher initial value.
* In the right chart, all batch sizes have stabilized, showing similar performance with minor fluctuations.
* The shaded regions around each line indicate the variability or uncertainty in the local learning coefficient for each batch size.
### Interpretation
The charts suggest that different batch sizes have varying initial learning curves. Batch sizes 8 and 16 require more iterations to reach a stable local learning coefficient compared to batch size 32. However, after a certain number of iterations (as shown in the right chart), all batch sizes converge to a similar performance level. The shaded regions indicate the robustness of each batch size, with narrower regions suggesting more consistent performance. The choice of batch size may depend on the trade-off between initial learning speed and overall stability.
</details>
(b) Second initialization
Figure 3: LLC estimates with 95% confidence intervals shown every 1,000 iterations, computed for various values of batch size and Adam learning rate $\eta$ and for two intializations. Left: $\eta=1× 10^{-4}$ . Right: $\eta=1× 10^{-3}$ .
4 Discussion
A significant limitation of the proposed reformulation of the physics-informed learning problem is that it’s limited to PINN models with hard initial and boundary condition constraints. In practice, physics-informed learning is often carried without such hard constraints by penalizing not only the residual of the governing equation, $y_{\text{GE}}(x,t)\coloneqq\mathcal{L}(u(x,t,w),x,t)$ but also the residual of the initial conditions, $y_{\text{IC}}(x,w)\coloneqq u(x,0,w)$ , and boundary conditions, $y_{\text{BC}}\coloneqq\mathcal{B}(u(x,t,w),x,t)$ . These residuals can also be regarded as observables of the PINN model. As such, a possible approach to reformulate the physics-informed learning problem with multiple residuals is to introduce a single vector observable $y\coloneqq[y_{\text{GE}},y_{\text{IC}},y_{\text{BC}}]$ and consider the statistical learning problem problem with model $p(y\mid x,t,w)q(x,t).$ In such a formulation, the number of samples $n$ would be the same for all residuals, which goes against the common practice of using different numbers of residual evaluation points for each term of the PINN loss. Nevertheless, we note that as indicated in section 2, PINN learning with randomized batches approximates minimizing the population loss $L(w)\equiv\lim_{n→∞}L_{n}(w)$ , which establishes a possible equivalency between the physics-informed learning problem with multiple residuals and the reformulation described above. Furthermore, special care must be taken when considering the weighting coefficients commonly used in physics-informed learning to weight the different residuals in the PINN loss. These problems will be considered in future work.
The presented statistical learning analysis of the physics-informed learning problem has interesting implications on the quantification of uncertainty in PINN predictions. Bayesian uncertainty quantification (UQ) in PINN predictions is often carried out by choosing a prior on the model parameters and a fixed set of residual evaluation points and computing samples from the posterior distribution of PINN parameters (see, e.g. [17, 16].) As noted in [17], MCMC sampling of the PINN loss is highly challenging computationally due to the singular nature of PINN models, but such an exhaustive exploration of parameter space may not be necessary because the solution candidates may differ significantly in parameter space but correspond to only a limited range of loss values and thus a limited range of possible predictions. In other words, this analysis justifies taking a function-space view of Bayesian UQ for physics-informed learning due to the singular nature of the PINN models involved.
<details>
<summary>x8.png Details</summary>
### Visual Description
## Heatmap: Spatio-temporal Data Visualization
### Overview
The image is a heatmap visualizing data across two dimensions, x and t. The color intensity represents the magnitude of the data, ranging from dark purple (low) to bright yellow (high). Black contour lines indicate regions of equal value. A red dashed line is present at t=2.0.
### Components/Axes
* **X-axis:** Labeled "x", ranges from 0.0 to 2.0 in increments of 0.5.
* **Y-axis:** Labeled "t", ranges from 0.0 to 4.0 in increments of 0.5.
* **Colorbar:** Located on the right side, ranging from 0.000 to 0.175. The color gradient goes from dark purple (0.000) to yellow (0.175), with intermediate values of 0.025, 0.050, 0.075, 0.100, 0.125, and 0.150.
* **Contour Lines:** Black lines indicating regions of equal value.
* **Red Dashed Line:** Horizontal line at t = 2.0.
### Detailed Analysis
* **Background Color:** The background color is dark purple, indicating low values across most of the x-t plane.
* **High-Value Region:** A region of high values (yellow) is located in the upper-center of the plot, approximately between x = 1.0 and x = 1.5, and t = 3.5 and t = 4.0.
* **Contour Lines:** The contour lines are concentrated around the high-value region, indicating a gradient of decreasing values as you move away from the center. The contours are roughly elliptical.
* **Red Dashed Line:** The red dashed line at t=2.0 appears to be a reference line.
### Key Observations
* The highest data values are concentrated in the upper-center region of the plot.
* The data values are generally low across the rest of the plot.
* The contour lines suggest a smooth, continuous distribution of values.
### Interpretation
The heatmap visualizes a spatio-temporal distribution where the highest values are localized in a specific region (around x=1.25, t=3.75). The red dashed line at t=2.0 might represent a threshold or a point of interest in the temporal dimension. The data suggests a phenomenon that is strongest in the upper-center region and diminishes as you move away from it in both the x and t dimensions. The nature of the phenomenon cannot be determined without additional context.
</details>
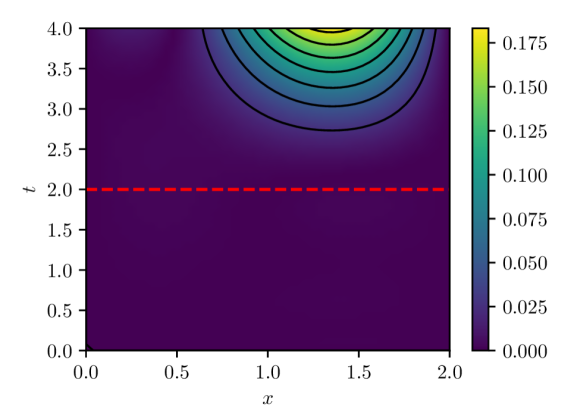
(a) $|u(x,t)-u(x,t,w^{0})|$
<details>
<summary>x9.png Details</summary>
### Visual Description
## Heatmap: Concentration Over Time and Space
### Overview
The image is a heatmap displaying the concentration of a substance over time (t) and space (x). The x-axis ranges from 0.0 to 2.0, and the t-axis ranges from 0.0 to 4.0. The color gradient represents the concentration, ranging from dark purple (0.00) to bright yellow (0.30). Black contour lines indicate specific concentration levels. A red dashed line is present at t=2.0.
### Components/Axes
* **X-axis:** Represents spatial dimension 'x', ranging from 0.0 to 2.0 in increments of 0.5.
* **Y-axis:** Represents time 't', ranging from 0.0 to 4.0 in increments of 0.5.
* **Colorbar:** Located on the right side, indicating concentration levels. The color gradient ranges from dark purple (0.00) to bright yellow (0.30), with intermediate values marked at 0.05, 0.10, 0.15, 0.20, and 0.25.
* **Contour Lines:** Black lines indicating specific concentration levels.
* **Red Dashed Line:** A horizontal red dashed line at t=2.0.
### Detailed Analysis
* **Concentration Distribution:** The concentration is highest in the upper-center region of the plot (around x=1.0 and t=4.0), indicated by the yellow color. The concentration decreases towards the bottom of the plot (lower t values) and towards the edges (x=0.0 and x=2.0).
* **Contour Lines:** The contour lines are denser in the upper region, indicating a steeper gradient of concentration change.
* **Red Dashed Line (t=2.0):** Below this line, the concentration is consistently low (dark purple). Above this line, the concentration increases, reaching its maximum near t=4.0.
* **Specific Values (Approximate):**
* At x=1.0 and t=4.0, the concentration is approximately 0.30 (yellow).
* At x=0.0 and t=0.0, the concentration is approximately 0.00 (dark purple).
* At x=2.0 and t=0.0, the concentration is approximately 0.00 (dark purple).
* **Trend:** The concentration increases with time (t) for a given spatial location (x), especially above t=2.0. The concentration also varies with spatial location (x), with the highest concentration near the center (x=1.0).
### Key Observations
* The concentration is significantly lower below t=2.0.
* The highest concentration is observed near the center of the spatial domain (x=1.0) and at the highest time point (t=4.0).
* The concentration gradient is steeper in the upper region of the plot.
### Interpretation
The heatmap illustrates the spatiotemporal distribution of a substance's concentration. The data suggests that the substance is introduced or generated after t=2.0, with the highest concentration accumulating near the center of the spatial domain over time. The red dashed line at t=2.0 serves as a clear demarcation point, separating regions of low and increasing concentration. The contour lines provide a visual representation of the concentration gradient, indicating how rapidly the concentration changes across different locations and times. The data could represent a diffusion process, a chemical reaction, or any other phenomenon where concentration varies with both space and time.
</details>
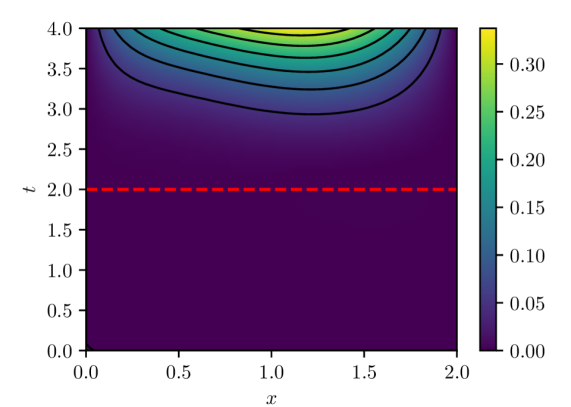
(b) $|u(x,t)-u(x,t,w^{1})|$
Figure 4: PINN pointwise absolute estimation errors for two different parameter vectors, $w^{0}$ and $w^{1}$ , obtained via different initializations of the PINN model parameters. Both parameter estimates were computed using batch size of 32 and Adam learning rate of $1× 10^{-4}$ over $100,000$ iterations. Despite both cases exhibiting result in similar loss $L_{n}(w)$ values ( $O(10^{-5}$ ) and similar LLCs $\hat{\lambda}(w^{0})≈\hat{\lambda}(w^{1})≈ 9.5$ , they exhibit noticeably different extrapolation behavior.
Finally, another implication of the presented analysis is related to the extrapolatory capacity of PINNs. It is understood that PINNs, particularly time-dependent PINNs, exhibit limited extrapolation capacity for times outside the training window $(0,T]$ (see, e.g., [12].) The presented analysis indicates that this is not a feature of the IBVP being solved, or the number and arrangement of residual evaluation points in space and time, or the optimization algorithm, but rather because physics-informed learning estimates model parameters by effectively minimizing $L(w)\propto\mathbb{E}_{q(x,t)}[\mathcal{L}(u(x,t,w),x,t)]^{2}$ , that is, the expected loss over the input-generating distribution $q(x,t)$ with support given by the spatio-temporal training windows. While PINN solution candidates along the flat minima are all within a limited range of loss values, there is no guarantee that this would hold for a different loss $L^{\prime}(w)\propto\mathbb{E}_{q^{\prime}(x,t)}[\mathcal{L}(u(x,t,w),x,t)]^{2}$ because the structure of the flat minima is tightly tied to the definition of the original loss, which is tightly tied to the input-generating distribution and thus to the spatio-temporal training window. In fact, it can be expected that two $w$ s with losses $L(w)$ within tolerance will lead to vastly different extrapolation losses $L^{\prime}(w)$ (e.g., see fig. 4), consistent with extrapolation results reported in the literature.
5 Conclusions
We have presented in this work a reformulation of physics-informed learning for IBVPs as a statistical learning problem valid for hard-constraints parameterizations. Employing this reformulation we argue that the physics penalty in physics-informed learning is not a regularizer as it is often interpreted (see, e.g. [6]) but as an infinite source of data, and that physics-informed learning with randomized batches of residual evaluation points is equivalent to minimizing $L(w)\propto\mathbb{E}_{q(x,t)}[\mathcal{L}(u(x,t,w),x,t)]^{2}$ . Furthermore, we show that the PINN loss landscape is not characterized by discrete, measure-zero local minima even in the limit $n→∞$ of the number of residual evaluation points but by very flat local minima corresponding to a low-complexity (relative to the number of model parameters) model class. We characterize the flatness of local minima of the PINN loss landscape by computing the LLC for a heat equation IBVP and various initializations and choices of learning hyperparameters, and find that across all initializations and hyperparameters studied the estimated LLC is very similar, indicating that the neighborhoods around the different PINN parameter estimates obtained across the experiments exhibit similar flatness properties. Finally, we discuss the implications of the proposed analysis on the quantification of the uncertainty in PINN predictions, and the extrapolation capacity of PINN models.
Acknowledgments
This material is based upon work supported by the U.S. Department of Energy (DOE), Office of Science, Office of Advanced Scientific Computing Research. Pacific Northwest National Laboratory is operated by Battelle for the DOE under Contract DE-AC05-76RL01830.
References
- [1] A. Cabezas, A. Corenflos, J. Lao, and R. Louf (2024) BlackJAX: composable Bayesian inference in JAX. External Links: 2402.10797 Cited by: §1, §3.2.
- [2] Flax: a neural network library and ecosystem for JAX External Links: Link Cited by: §1, §3.2.
- [3] M. D. Homan and A. Gelman (2014-01) The no-u-turn sampler: adaptively setting path lengths in hamiltonian monte carlo. J. Mach. Learn. Res. 15 (1), pp. 1593–1623. External Links: ISSN 1532-4435 Cited by: §1, §3.1.
- [4] J. Hoogland, G. Wang, M. Farrugia-Roberts, L. Carroll, S. Wei, and D. Murfet (2025) Loss landscape degeneracy and stagewise development in transformers. Transactions on Machine Learning Research. Note: External Links: ISSN 2835-8856, Link Cited by: §1.
- [5] T. Imai (2019-06) Estimating Real Log Canonical Thresholds. arXiv e-prints, pp. arXiv:1906.01341. External Links: Document, 1906.01341 Cited by: §3.1.
- [6] G. E. Karniadakis, I. G. Kevrekidis, L. Lu, P. Perdikaris, S. Wang, and L. Yang (2021) Physics-informed machine learning. Nature Reviews Physics 3 (6), pp. 422–440. Cited by: §1, §5.
- [7] E. Lau, Z. Furman, G. Wang, D. Murfet, and S. Wei (2025-03–05 May) The local learning coefficient: a singularity-aware complexity measure. In Proceedings of The 28th International Conference on Artificial Intelligence and Statistics, Y. Li, S. Mandt, S. Agrawal, and E. Khan (Eds.), Proceedings of Machine Learning Research, Vol. 258, pp. 244–252. External Links: Link Cited by: §1, §2, §3.1, Statistical Learning Analysis of Physics-Informed Neural Networks.
- [8] X. Li, J. Deng, J. Wu, S. Zhang, W. Li, and Y. Wang (2024) Physical informed neural networks with soft and hard boundary constraints for solving advection-diffusion equations using fourier expansions. Computers & Mathematics with Applications 159, pp. 60–75. External Links: ISSN 0898-1221, Document, Link Cited by: §2.
- [9] L. Lu, R. Pestourie, W. Yao, Z. Wang, F. Verdugo, and S. G. Johnson (2021) Physics-informed neural networks with hard constraints for inverse design. SIAM Journal on Scientific Computing 43 (6), pp. B1105–B1132. External Links: Document, Link, https://doi.org/10.1137/21M1397908 Cited by: §2.
- [10] A. Muhamed and V. Smith (2025) The geometry of forgetting: analyzing machine unlearning through local learning coefficients. In ICML 2025 Workshop on Reliable and Responsible Foundation Models, External Links: Link Cited by: §1.
- [11] A. M. Tartakovsky and Y. Zong (2024) Physics-informed machine learning method with space-time karhunen-loève expansions for forward and inverse partial differential equations. Journal of Computational Physics 499, pp. 112723. External Links: ISSN 0021-9991, Document, Link Cited by: §2.
- [12] Y. Wang, Y. Yao, and Z. Gao (2025) An extrapolation-driven network architecture for physics-informed deep learning. Neural Networks 183, pp. 106998. External Links: ISSN 0893-6080, Document, Link Cited by: §4.
- [13] S. Watanabe (2009) Algebraic geometry and statistical learning theory. Vol. 25, Cambridge University Press. Cited by: §1, §2.
- [14] S. Watanabe (2013-03) A widely applicable bayesian information criterion. 14 (1), pp. 867–897. External Links: ISSN 1532-4435 Cited by: §3.1.
- [15] M. Welling and Y. W. Teh (2011) Bayesian learning via stochastic gradient langevin dynamics. In Proceedings of the 28th International Conference on International Conference on Machine Learning, ICML’11, Madison, WI, USA, pp. 681–688. External Links: ISBN 9781450306195 Cited by: §1.
- [16] L. Yang, X. Meng, and G. E. Karniadakis (2021) B-pinns: bayesian physics-informed neural networks for forward and inverse pde problems with noisy data. Journal of Computational Physics 425, pp. 109913. External Links: ISSN 0021-9991, Document, Link Cited by: §4.
- [17] Y. Zong, D. A. Barajas-Solano, and A. M. Tartakovsky (2025) Randomized Physics-informed Neural Networks for Bayesian Data Assimilation. Comput. Methods Appl. Mech. Eng. 436, pp. 117670. External Links: Document Cited by: §4.